Team:Paris Saclay/Notebook/September/8
From 2014.igem.org
(→PCR with bacteria) |
m (→PCR with bacteria) |
||
| (20 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| + | {{Team:Paris_Saclay/notebook_header}} | ||
| + | =Monday 8th September= | ||
| + | |||
==Lab Work== | ==Lab Work== | ||
| + | ''by Mélanie'' | ||
| - | === | + | ===Lemon Scent=== |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
====PCR with bacteria==== | ====PCR with bacteria==== | ||
From the petri dish ([https://2014.igem.org/Team:Paris_Saclay/Notebook/September/5#Transformation made here], I select some clone to do a PCR : | From the petri dish ([https://2014.igem.org/Team:Paris_Saclay/Notebook/September/5#Transformation made here], I select some clone to do a PCR : | ||
| Line 12: | Line 11: | ||
clones 4-5 for PS | clones 4-5 for PS | ||
clones 6 for LS | clones 6 for LS | ||
| + | |||
| + | {| class="wikitable centre" width="50%" | ||
| + | |+ | ||
| + | |- | ||
| + | ! scope=col | component | ||
| + | ! scope=col | volume | ||
| + | |- | ||
| + | |H<sub>2</sub>O | ||
| + | |41.5μl | ||
| + | |- | ||
| + | |buffer | ||
| + | |5μl | ||
| + | |- | ||
| + | |dNTPs | ||
| + | |1μl | ||
| + | |- | ||
| + | |Primer 1 | ||
| + | |1μl | ||
| + | |- | ||
| + | |Primer 2 | ||
| + | |1μl | ||
| + | |- | ||
| + | |bacteria | ||
| + | |(about 1µl = 1 colony) | ||
| + | |- | ||
| + | |Dream taq | ||
| + | |0.5μl | ||
| + | |} | ||
| + | |||
| + | Primer used: | ||
| + | |||
| + | For cloning in Topo vector = Pu Pr (universal primer) | ||
| + | |||
| + | For pPS2 = iPS 66/67 | ||
{| class="wikitable centre" width="80%" | {| class="wikitable centre" width="80%" | ||
| Line 55: | Line 88: | ||
| 1 | | 1 | ||
|} | |} | ||
| + | |||
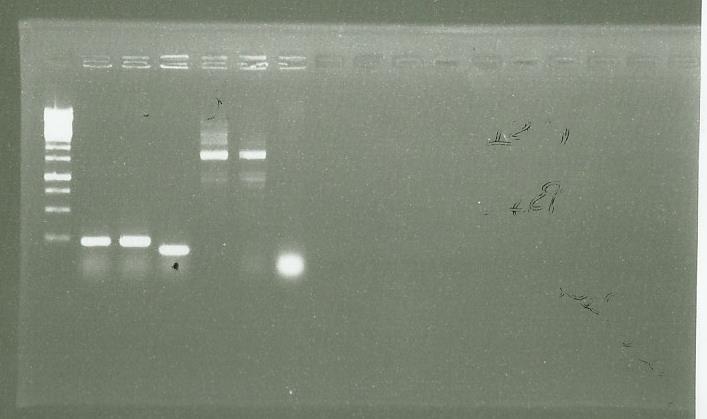
| + | [[File:0809 PCR clones.jpg|400px|center]] | ||
| + | |||
| + | We identify that we have success with to clone of PS. (well 5-6) | ||
| + | |||
| + | ====Liquide culture of clones==== | ||
| + | The PS clone that reveal to be ok are cultivate in LB + AMP médium | ||
| + | |||
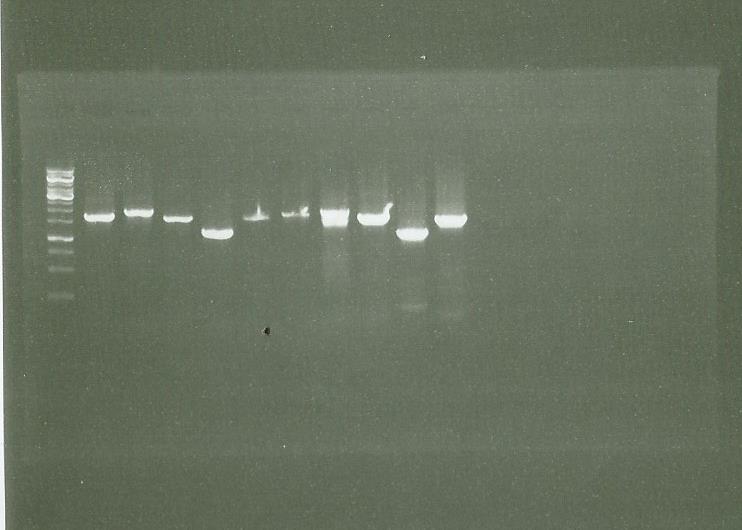
| + | ====Checking PCR==== | ||
| + | Electrophoresis of the PCR made [https://2014.igem.org/Team:Paris_Saclay/Notebook/September/5#PCR_LS_PS_GS_Cad Friday] | ||
| + | |||
| + | LS PS GS CAD (and chromo) (2 enzymes = vent taq and dream taq) | ||
| + | |||
| + | [[File:0809 PCR LSPSGSCADCRHOMO vent dream.jpg|400px|center]] | ||
| + | |||
| + | ====Cloning==== | ||
| + | With this PCR, I do a cloning in a Topo Cloning vector (Zero Blunt TOPO PCR Cloning Kit) (LS GS and CAD) | ||
| + | |||
| + | {| class="wikitable centre" width="50%" | ||
| + | |+ | ||
| + | |- | ||
| + | ! scope=col | component | ||
| + | ! scope=col | volume | ||
| + | |- | ||
| + | |PCR product | ||
| + | |1μl | ||
| + | |- | ||
| + | |Salt | ||
| + | |1μl | ||
| + | |- | ||
| + | |Vector | ||
| + | |1 μl | ||
| + | |- | ||
| + | |H<sub>2</sub>O | ||
| + | |3μl | ||
| + | |} | ||
| + | |||
| + | |||
| + | Between 5 and 30 min at room temperature | ||
| + | |||
| + | ==== Transformation==== | ||
| + | |||
| + | Add 50µL of bacteria to the clonage previously done | ||
| + | |||
| + | 30min on ice | ||
| + | 45 sec at 42°C | ||
| + | |||
| + | add 1ml of LB | ||
| + | |||
| + | 1hour at 37° | ||
| + | |||
| + | Spread on a petri dish | ||
| + | |||
| + | ===Construction of the fusion protein=== | ||
| + | We check the sequence of the fusion protein and identify a clone with our chromoprotein = W6 | ||
| + | |||
| + | ==Photo of the Day== | ||
| + | [[File:Paris Saclay 8_september.jpg|150px|center]] | ||
| + | |||
| + | [http://ifris.org/membre/morgan-meyer/ Morgan Meyer] , one of those few we have interviewed during this month of september. | ||
| + | |||
| + | To learn more about the ethic aspect of our project, just click [https://2014.igem.org/Team:Paris_Saclay/Ethics here] | ||
| + | |||
| + | |||
| + | {{Team:Paris_Saclay/notebook_footer}} | ||
Latest revision as of 12:55, 14 October 2014
Contents |
Monday 8th September
Lab Work
by Mélanie
Lemon Scent
PCR with bacteria
From the petri dish (made here, I select some clone to do a PCR : clones 1-2-3 for cad clones 4-5 for PS clones 6 for LS
| component | volume |
|---|---|
| H2O | 41.5μl |
| buffer | 5μl |
| dNTPs | 1μl |
| Primer 1 | 1μl |
| Primer 2 | 1μl |
| bacteria | (about 1µl = 1 colony) |
| Dream taq | 0.5μl |
Primer used:
For cloning in Topo vector = Pu Pr (universal primer)
For pPS2 = iPS 66/67
| Cycle step | Temperature | Time | Cycle |
|---|---|---|---|
|
Bacteria lysis |
95°C |
5 min |
1 |
| Denaturation | 94°C | 30 s | 25 |
| Annealing | 50°C | 25 s | 25 |
| Extension | 72°C | 1 min | 25 |
| Final extension | 72°C | 10 min | 1 |
| Final extension | 8°C | hold | 1 |
We identify that we have success with to clone of PS. (well 5-6)
Liquide culture of clones
The PS clone that reveal to be ok are cultivate in LB + AMP médium
Checking PCR
Electrophoresis of the PCR made Friday
LS PS GS CAD (and chromo) (2 enzymes = vent taq and dream taq)
Cloning
With this PCR, I do a cloning in a Topo Cloning vector (Zero Blunt TOPO PCR Cloning Kit) (LS GS and CAD)
| component | volume |
|---|---|
| PCR product | 1μl |
| Salt | 1μl |
| Vector | 1 μl |
| H2O | 3μl |
Between 5 and 30 min at room temperature
Transformation
Add 50µL of bacteria to the clonage previously done
30min on ice 45 sec at 42°C
add 1ml of LB
1hour at 37°
Spread on a petri dish
Construction of the fusion protein
We check the sequence of the fusion protein and identify a clone with our chromoprotein = W6
Photo of the Day
Morgan Meyer , one of those few we have interviewed during this month of september.
To learn more about the ethic aspect of our project, just click here
 "
"