Team:Paris Saclay/Project/Lemon Scent
From 2014.igem.org
(Created page with "{{Team:Paris_Saclay/default_header}} =Lemon Scent= ==Sub title== Lorem ipsum dolor sit amet, consectetur adipiscing elit. In vestibulum ac lacus luctus semper. Etiam feugiat faci...") |
(→Lemon Scent) |
||
| Line 1: | Line 1: | ||
{{Team:Paris_Saclay/default_header}} | {{Team:Paris_Saclay/default_header}} | ||
=Lemon Scent= | =Lemon Scent= | ||
| - | == | + | ==Summary== |
| - | + | In the part of the project, we want to express the main components of the lemon scent (limonene, β-pinene and citral) in the odor-free ''E. coli'' we constructed. For this, we plan to engineer a plasmid constructed by the team of Taek Soon Lee, bearing a complete mevalonate pathway, a geranyldiphosphate synthase and a (S)-limonene synthase, to express (i) a β-pinene synthase (BBa_K517003), (ii) a geraniol synthase (GES) and a cinnamyl alcohol dehydrogenase (CAD) to produce β-pinene and citral respectively. We also would like to characterize the limonene synthase biobrick constructed by the Edinburgh team (BBa_K722100) and verify whether it is functional by replaced the intial (S)-limonene synthase by the biobrick. | |
| + | Finally we construct two of our plasmid, the β-pinene synthase and the geraniol synthase plasmid. We couldn’t obtain more during this summer. | ||
| - | + | ==Introduction== | |
| + | The lemon scent is due to many volatile compounds, but the three main components are the limonene, pinene and citral. All these molecules belong the terpene family of metabolites. | ||
| + | |||
| + | ===Limonene=== | ||
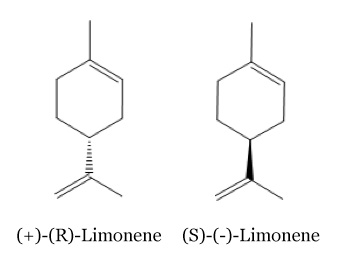
| + | There are two isomeric forms of the limonene molecules: | ||
| + | The smell of these two isomers is a little bit different, the (R)-Limonene have a near orange smell than the (S)-Limonene have a strongly lemon scent. [2] | ||
| + | |||
| + | [[File:Paris Saclay project-odor-limonene.jpg|center]] | ||
| + | |||
| + | ===Pinene=== | ||
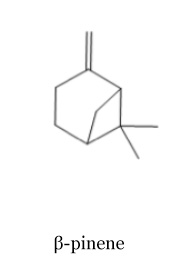
| + | There are also two isomeric forms of pinene: alpha and beta pinene. In the lemon oil, 1 β-pinene represent between 10-17% of the total compounds [3]. In contrast, there is no trace of α-pinene. | ||
| + | |||
| + | [[File:Paris Saclay project-odor-pinene.jpg|center]] | ||
| + | |||
| + | ===Citral=== | ||
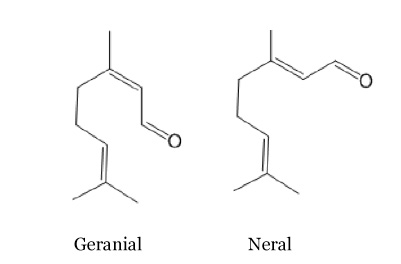
| + | It’s the principal compound of the lemon grass, lemon smell and it is present in many other plants as orange, verbena... Here again there are two stereoisomeric forms, geranial and neral. The trans isomer, the geranial or citral A, has a strong lemon scent. On the contrary, the cis isomer named neral or citral B has a weaker lemon scent. There compounds are often used in the perfume industry to reproduce the smell of lemon. | ||
| + | |||
| + | [[File:Paris Saclay project-odor-citral.jpg|center]] | ||
| + | |||
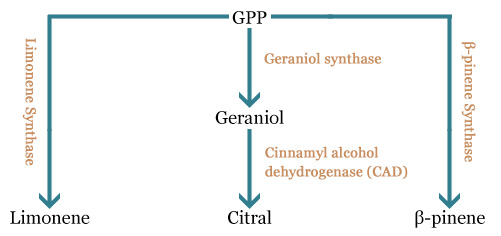
| + | All these compounds derive from a common precursor, the geranylpyrophosphate (GPP). This precursor is converted in one or two steps in the molecules we are interested in obtaining. | ||
| + | |||
| + | [[File:Paris Saclay project-odor-gpp.jpg|center]] | ||
| + | |||
| + | The desoxyxylulose 5-phosphate (DXP) parthway is the common pathway used by prokaryote cells to produce GPP. | ||
| + | E.coli synthesizes GPP with the DXP pathway but in low quantity [4]. | ||
| + | However, another pathway, the mevalonate pathway (MEV) exists in eukaryote and some prokaryotes. Moreover, plants have the two pathways to produce GPP in higher quantity. | ||
| + | To copy the plants' strategy, we chose to add the MEV pathway to our engineering E. coli in order to produce higher quantity of GPP and thereby have a stronger lemon smell. | ||
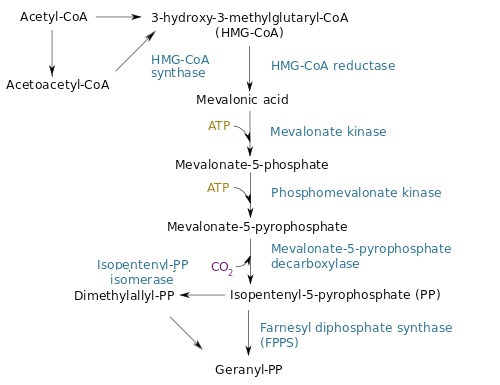
| + | The figure below illustrates the MEV pathway: | ||
| + | |||
| + | [[File:Paris Saclay project-odor-mev-pathway.jpg|center]] | ||
| + | |||
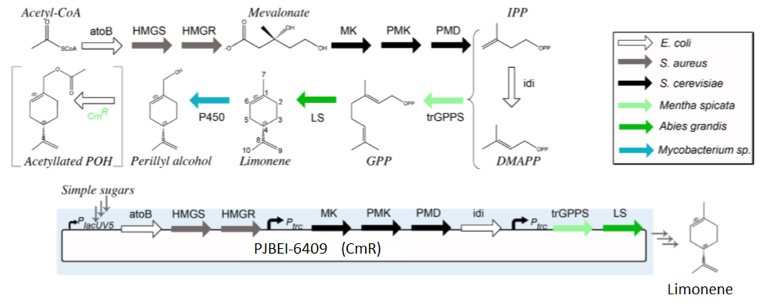
| + | To tackle the problem of the low GPP bioavailability in E. coli, the Taek Soon Lee’s research group designed and constructed a plasmid with a complete mevalonate pathway biosynthetic pathway for the production of GPP [3]. They then expressed the ''Abies grandis'' (S)-limonene synthase (LS) gene in the same plasmid, named pJBEI6409. With this construction, they were able to produce 400 mg/L of (S)-limonene. | ||
| + | |||
| + | [[File:Paris Saclay project-odor-limonene-synthase.jpg|center]] | ||
| + | |||
| + | We decided to make use of their construction for the production of limonene, β-pinene and citral in E. coli. We thank the team of Taek Soon Lee for the gift of the pJBEI6409 plasmid. | ||
| + | |||
| + | ==Principle of our Constructions== | ||
| + | For our project, we need to replace the (S)-limonene synthase gene by the (R)-limonene synthase gene, the β-pinene synthase gene and the geraniol synthase (GES) and cinnamyl alcohol dehydrogenase (CAD) genes. | ||
| + | |||
| + | However, pJBEI6409 was constructed using a 3A assembly method, meaning that there are no restriction sites flanking the LS gene. | ||
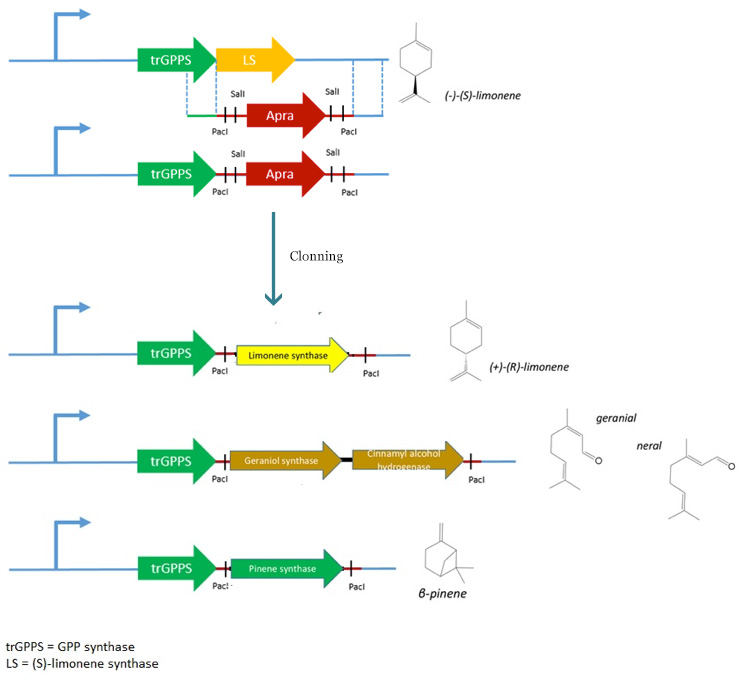
| + | We therefore decided to replace the (S) Limonene Synthase gene by an apramycin resistance cassette flanked by ''PacI'' and ''SalI'' restriction sites by double homologous recombination. The resulting plasmid (pPS1) will then be used to clone the various genes (limonene synthase – β-pinene synthase – geraniol synthase and cinnamyl alcohol dehydrogenase). | ||
| + | |||
| + | [[File:Paris Saclay project-odor-recap.jpg|center]] | ||
| + | |||
| + | ===Construction of pPS1=== | ||
| + | ''Replacement of the LS gene by an apramycin cassette'' | ||
| + | |||
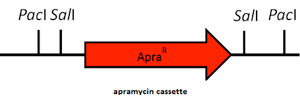
| + | The first step of this sub-project is to replace the (S)-limonene synthase gene by an apramycin-resistant cassette. This cassette is flanked by ''PacI'' and ''SalI'' restriction sites to facilitate the cloning of the different genes. | ||
| + | |||
| + | [[File:Paris Saclay project-odor-apramycine-k7.jpg|center]] | ||
| + | |||
| + | The (S)-LS starting gene is replaced by the apramycin cassette by PCR targeting. | ||
| + | # PCR amplification of the apramycin cassette with oligonucleotides iPS70 and iPS71 that contain the restriction sites PacI and SalI. The ends of these oligonucleotides are homologous with the sequence that constitutes LS. | ||
| + | # Transformation of the E. coli DY330 strain [5] (hyperrecombinant strain) by the pJBEI6409 plasmid. | ||
| + | # Transformation of the E. coli DY330/pJBEI6409 with the PCR product after induction of the recombination genes. | ||
| + | # Verification of the clones by ''BglII'' digestion. | ||
| + | [https://2014.igem.org/Team:Paris_Saclay/Notebook/August/11 We call this plasmid pPS1.] | ||
| + | |||
| + | ===Construction pPS2=== | ||
| + | ''Replacement of the apramycin cassette by the (R)- LS gene, '''''Bba_K722100''''' | ||
| + | |||
| + | This step consists in replacing the apramycin resistance gene cassette introduced in pJBEI6409 by the (R)-LS gene. | ||
| + | # PCR amplification of '''Bba_K722100''' with oligonucleotides iPS66 and iPS67 that contain the restriction sites ''PacI'' and ''SalI''. | ||
| + | # TA-cloning of the PCR product in the Pcr 4 Blunt-Topo vector (LS Topo plasmid). | ||
| + | # [https://2014.igem.org/Team:Paris_Saclay/Notebook/September/9 Verification] of the clones LS-TOPO by PCR with the oligonucleotides iPS66 and iPS67 and sequencing. | ||
| + | # Digestion of the LS-TOPO plasmid by ''PacI'' and by ''MscI'' to eliminate the topo vector. | ||
| + | # Digestion of pPS1 by ''PacI''. | ||
| + | # Ligation of the LS gene into pPS1/''PacI''. | ||
| + | # Transformation of E. coli DH5α with the ligation product. | ||
| + | # Verification of the clones by PCR with the oligonucleotides iPS72 and iPS96 | ||
| + | Here we failed to obtain an insert in the right sens. | ||
| + | |||
| + | ===Construction of pPS3=== | ||
| + | ''Replacement of the apramycin cassette by the β pinene gene, '''BBa_K517003''''' | ||
| + | |||
| + | This step consists in replacing the apramycin resistance gene cassette introduced in pJBEI6409 by the (R)-LS gene. | ||
| + | # PCR amplification of '''Bba_K722100''' with oligonucleotides iPS68bis and iPS69 that contain the restriction sites ''PacI'' and ''SalI''. | ||
| + | # [https://2014.igem.org/Team:Paris_Saclay/Notebook/September/12 TA-cloning] of the PCR product in the Pcr 4 Blunt-Topo vector (PS Topo plasmid). | ||
| + | # Verification of the clones LS-TOPO by PCR with the oligonucleotides iPS68bis and iPS69 and sequencing. | ||
| + | # Digestion of the LS-TOPO plasmid by PacI and by ''PstI'' to eliminate the topo vector. | ||
| + | # Digestion of pPS1 by ''PacI''. | ||
| + | # Ligation of the LS gene into pPS1/''PacI''. | ||
| + | # Transformation of E. coli DH5α with the ligation product. | ||
| + | # Verification of the clones by PCR with the oligonucleotides iPS72 and iPS93. | ||
| + | |||
| + | ===Construction of pPS4=== | ||
| + | ''Replacement of the apramycin cassette by the geraniol synthase gene, '''BBa_K517003''''' | ||
| + | |||
| + | This step consists in replacing the apramycin resistance gene cassette introduced in pJBEI6409 by geraniol synthase gene (from plasmid pCola (Strasbourg team)). | ||
| + | # PCR amplification of '''Bba_K722100''' with oligonucleotides iPS81bis and iPS82 that contain the restriction sites ''PacI'' and ''SalI''. | ||
| + | # TA-cloning of the PCR product in the Pcr 4 Blunt-Topo vector (Topo plasmid). | ||
| + | # [https://2014.igem.org/Team:Paris_Saclay/Notebook/September/9 Verification] of the clones LS-TOPO by PCR with the oligonucleotides i iPS81bis and iPS82 and sequencing. | ||
| + | # Digestion of the LS-TOPO plasmid by ''PacI'' and ''PstI'' to eliminate the topo vector. | ||
| + | # Digestion of pPS1 by ''PacI''. | ||
| + | # Ligation of the LS gene into pPS1/''PacI''. | ||
| + | # Transformation of E. coli DH5α with the ligation product. | ||
| + | # Verification of the clones by PCR with the oligonucleotides iPS72 and iPS96 | ||
| + | |||
| + | ===Construction of pPS5=== | ||
| + | ''Replacement of the apramycin cassette by the geraniol synthase gene, '''BBa_K517003''''' | ||
| + | |||
| + | This step consists in replacing the apramycin resistance gene cassette introduced in pJBEI6409 by geraniol synthase gene (from plasmid pCola (Strasbourg team)). | ||
| + | # PCR amplification of Cad (synthetic gene with oligonucleotides iPS79bis and iPS80 that contain the restriction sites ''PacI'' and ''SalI'' (like apramycin cassette). | ||
| + | # TA-cloning of the PCR product in the PCR 4 Blunt-Topo vector (Topo plasmid). | ||
| + | # [https://2014.igem.org/Team:Paris_Saclay/Notebook/September/9 Verification] of the clones LS-TOPO by PCR with the oligonucleotides i iPS81bis and iPS82 and sequencing. | ||
| + | # Digestion of the LS-TOPO plasmid by ''SalI'' and ''PvuII'' to eliminate the topo vector. | ||
| + | # Digestion of pPS4 by ''SalI''. | ||
| + | # Ligation of the LS gene into pPS1/''PacI''. | ||
| + | # Transformation of E. coli DH5α with the ligation product | ||
| + | # Verification of the clones by PCR with the oligonucleotide | ||
| + | |||
| + | ==Réferences== | ||
| + | * [1] | ||
| + | * [2] http://www.societechimiquedefrance.fr/produit-du-jour/limonene-et-monoterpenes.html | ||
| + | * [3] http://booksofdante.wordpress.com/tag/beta-pinene/ | ||
| + | * [4] Jorge Alonso-Gutierrez and al - Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production – metabolique engineering 2013 | ||
| + | * [5] Daiguan Yu and al - An efficient recombination system for chromosome engineering in Escherichia coli – 2000) | ||
| - | |||
{{Team:Paris_Saclay/default_footer}} | {{Team:Paris_Saclay/default_footer}} | ||
Revision as of 19:12, 1 October 2014
Contents |
Lemon Scent
Summary
In the part of the project, we want to express the main components of the lemon scent (limonene, β-pinene and citral) in the odor-free E. coli we constructed. For this, we plan to engineer a plasmid constructed by the team of Taek Soon Lee, bearing a complete mevalonate pathway, a geranyldiphosphate synthase and a (S)-limonene synthase, to express (i) a β-pinene synthase (BBa_K517003), (ii) a geraniol synthase (GES) and a cinnamyl alcohol dehydrogenase (CAD) to produce β-pinene and citral respectively. We also would like to characterize the limonene synthase biobrick constructed by the Edinburgh team (BBa_K722100) and verify whether it is functional by replaced the intial (S)-limonene synthase by the biobrick. Finally we construct two of our plasmid, the β-pinene synthase and the geraniol synthase plasmid. We couldn’t obtain more during this summer.
Introduction
The lemon scent is due to many volatile compounds, but the three main components are the limonene, pinene and citral. All these molecules belong the terpene family of metabolites.
Limonene
There are two isomeric forms of the limonene molecules: The smell of these two isomers is a little bit different, the (R)-Limonene have a near orange smell than the (S)-Limonene have a strongly lemon scent. [2]
Pinene
There are also two isomeric forms of pinene: alpha and beta pinene. In the lemon oil, 1 β-pinene represent between 10-17% of the total compounds [3]. In contrast, there is no trace of α-pinene.
Citral
It’s the principal compound of the lemon grass, lemon smell and it is present in many other plants as orange, verbena... Here again there are two stereoisomeric forms, geranial and neral. The trans isomer, the geranial or citral A, has a strong lemon scent. On the contrary, the cis isomer named neral or citral B has a weaker lemon scent. There compounds are often used in the perfume industry to reproduce the smell of lemon.
All these compounds derive from a common precursor, the geranylpyrophosphate (GPP). This precursor is converted in one or two steps in the molecules we are interested in obtaining.
The desoxyxylulose 5-phosphate (DXP) parthway is the common pathway used by prokaryote cells to produce GPP. E.coli synthesizes GPP with the DXP pathway but in low quantity [4]. However, another pathway, the mevalonate pathway (MEV) exists in eukaryote and some prokaryotes. Moreover, plants have the two pathways to produce GPP in higher quantity. To copy the plants' strategy, we chose to add the MEV pathway to our engineering E. coli in order to produce higher quantity of GPP and thereby have a stronger lemon smell. The figure below illustrates the MEV pathway:
To tackle the problem of the low GPP bioavailability in E. coli, the Taek Soon Lee’s research group designed and constructed a plasmid with a complete mevalonate pathway biosynthetic pathway for the production of GPP [3]. They then expressed the Abies grandis (S)-limonene synthase (LS) gene in the same plasmid, named pJBEI6409. With this construction, they were able to produce 400 mg/L of (S)-limonene.
We decided to make use of their construction for the production of limonene, β-pinene and citral in E. coli. We thank the team of Taek Soon Lee for the gift of the pJBEI6409 plasmid.
Principle of our Constructions
For our project, we need to replace the (S)-limonene synthase gene by the (R)-limonene synthase gene, the β-pinene synthase gene and the geraniol synthase (GES) and cinnamyl alcohol dehydrogenase (CAD) genes.
However, pJBEI6409 was constructed using a 3A assembly method, meaning that there are no restriction sites flanking the LS gene. We therefore decided to replace the (S) Limonene Synthase gene by an apramycin resistance cassette flanked by PacI and SalI restriction sites by double homologous recombination. The resulting plasmid (pPS1) will then be used to clone the various genes (limonene synthase – β-pinene synthase – geraniol synthase and cinnamyl alcohol dehydrogenase).
Construction of pPS1
Replacement of the LS gene by an apramycin cassette
The first step of this sub-project is to replace the (S)-limonene synthase gene by an apramycin-resistant cassette. This cassette is flanked by PacI and SalI restriction sites to facilitate the cloning of the different genes.
The (S)-LS starting gene is replaced by the apramycin cassette by PCR targeting.
- PCR amplification of the apramycin cassette with oligonucleotides iPS70 and iPS71 that contain the restriction sites PacI and SalI. The ends of these oligonucleotides are homologous with the sequence that constitutes LS.
- Transformation of the E. coli DY330 strain [5] (hyperrecombinant strain) by the pJBEI6409 plasmid.
- Transformation of the E. coli DY330/pJBEI6409 with the PCR product after induction of the recombination genes.
- Verification of the clones by BglII digestion.
Construction pPS2
Replacement of the apramycin cassette by the (R)- LS gene, Bba_K722100
This step consists in replacing the apramycin resistance gene cassette introduced in pJBEI6409 by the (R)-LS gene.
- PCR amplification of Bba_K722100 with oligonucleotides iPS66 and iPS67 that contain the restriction sites PacI and SalI.
- TA-cloning of the PCR product in the Pcr 4 Blunt-Topo vector (LS Topo plasmid).
- Verification of the clones LS-TOPO by PCR with the oligonucleotides iPS66 and iPS67 and sequencing.
- Digestion of the LS-TOPO plasmid by PacI and by MscI to eliminate the topo vector.
- Digestion of pPS1 by PacI.
- Ligation of the LS gene into pPS1/PacI.
- Transformation of E. coli DH5α with the ligation product.
- Verification of the clones by PCR with the oligonucleotides iPS72 and iPS96
Here we failed to obtain an insert in the right sens.
Construction of pPS3
Replacement of the apramycin cassette by the β pinene gene, BBa_K517003
This step consists in replacing the apramycin resistance gene cassette introduced in pJBEI6409 by the (R)-LS gene.
- PCR amplification of Bba_K722100 with oligonucleotides iPS68bis and iPS69 that contain the restriction sites PacI and SalI.
- TA-cloning of the PCR product in the Pcr 4 Blunt-Topo vector (PS Topo plasmid).
- Verification of the clones LS-TOPO by PCR with the oligonucleotides iPS68bis and iPS69 and sequencing.
- Digestion of the LS-TOPO plasmid by PacI and by PstI to eliminate the topo vector.
- Digestion of pPS1 by PacI.
- Ligation of the LS gene into pPS1/PacI.
- Transformation of E. coli DH5α with the ligation product.
- Verification of the clones by PCR with the oligonucleotides iPS72 and iPS93.
Construction of pPS4
Replacement of the apramycin cassette by the geraniol synthase gene, BBa_K517003
This step consists in replacing the apramycin resistance gene cassette introduced in pJBEI6409 by geraniol synthase gene (from plasmid pCola (Strasbourg team)).
- PCR amplification of Bba_K722100 with oligonucleotides iPS81bis and iPS82 that contain the restriction sites PacI and SalI.
- TA-cloning of the PCR product in the Pcr 4 Blunt-Topo vector (Topo plasmid).
- Verification of the clones LS-TOPO by PCR with the oligonucleotides i iPS81bis and iPS82 and sequencing.
- Digestion of the LS-TOPO plasmid by PacI and PstI to eliminate the topo vector.
- Digestion of pPS1 by PacI.
- Ligation of the LS gene into pPS1/PacI.
- Transformation of E. coli DH5α with the ligation product.
- Verification of the clones by PCR with the oligonucleotides iPS72 and iPS96
Construction of pPS5
Replacement of the apramycin cassette by the geraniol synthase gene, BBa_K517003
This step consists in replacing the apramycin resistance gene cassette introduced in pJBEI6409 by geraniol synthase gene (from plasmid pCola (Strasbourg team)).
- PCR amplification of Cad (synthetic gene with oligonucleotides iPS79bis and iPS80 that contain the restriction sites PacI and SalI (like apramycin cassette).
- TA-cloning of the PCR product in the PCR 4 Blunt-Topo vector (Topo plasmid).
- Verification of the clones LS-TOPO by PCR with the oligonucleotides i iPS81bis and iPS82 and sequencing.
- Digestion of the LS-TOPO plasmid by SalI and PvuII to eliminate the topo vector.
- Digestion of pPS4 by SalI.
- Ligation of the LS gene into pPS1/PacI.
- Transformation of E. coli DH5α with the ligation product
- Verification of the clones by PCR with the oligonucleotide
Réferences
- [1]
- [2] http://www.societechimiquedefrance.fr/produit-du-jour/limonene-et-monoterpenes.html
- [3] http://booksofdante.wordpress.com/tag/beta-pinene/
- [4] Jorge Alonso-Gutierrez and al - Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production – metabolique engineering 2013
- [5] Daiguan Yu and al - An efficient recombination system for chromosome engineering in Escherichia coli – 2000)
 "
"