Team:Oxford/codon optimisation
From 2014.igem.org
(Difference between revisions)
Olivervince (Talk | contribs) |
Olivervince (Talk | contribs) |
||
| Line 434: | Line 434: | ||
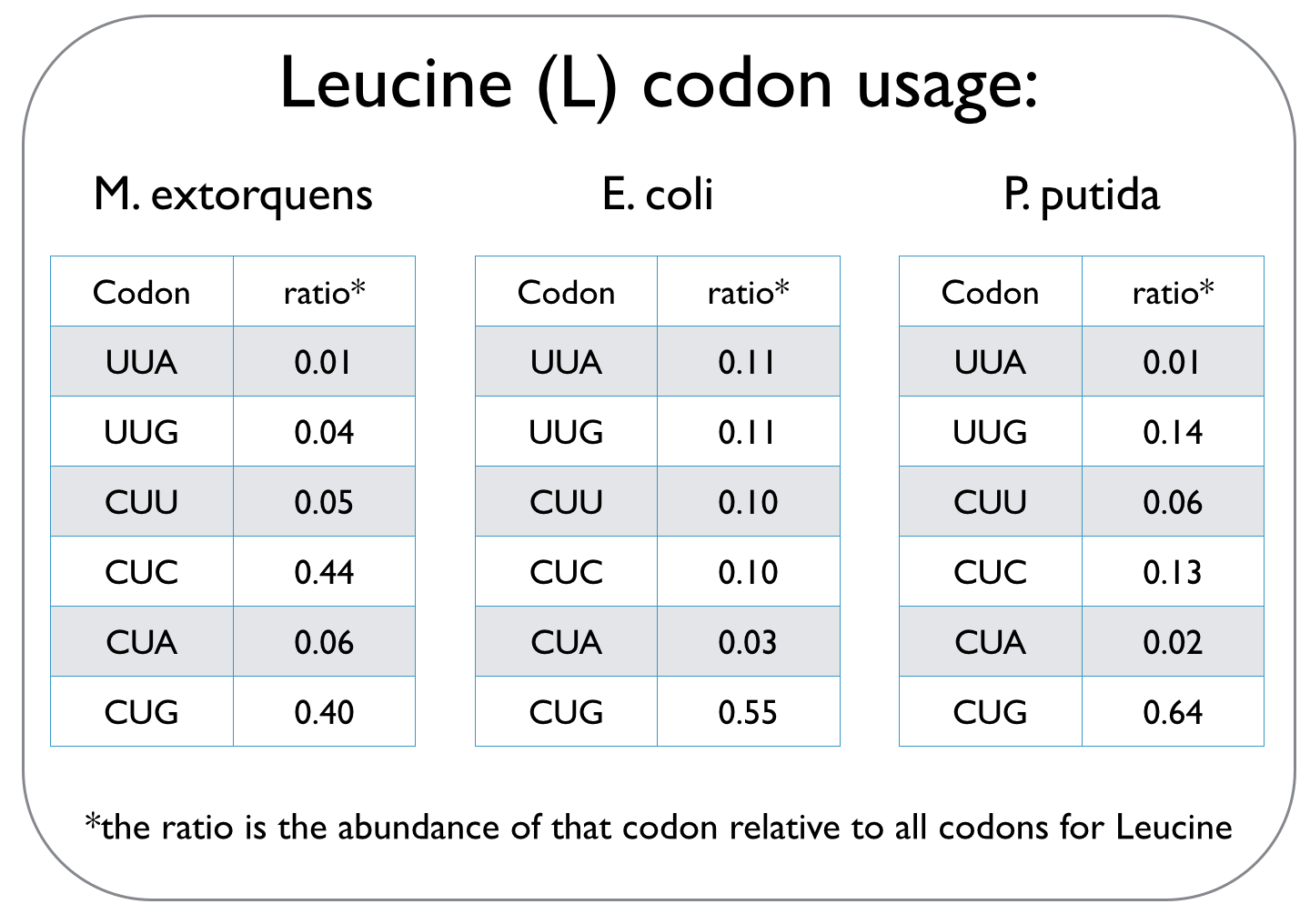
Cells use codons, which are composed of three consecutive nucleotides, to read the genetic code and translate it into proteins. This genetic code can not only vary between species, but it is also degenerate, meaning that several codons may specify a particular amino acid. Different species have unique codon usage biases, implying that they prefer particular codons over other codons that define the same amino acid. An excerpt of a codon usage frequency table is given for the three strains we worked with to highlight different codon biases. | Cells use codons, which are composed of three consecutive nucleotides, to read the genetic code and translate it into proteins. This genetic code can not only vary between species, but it is also degenerate, meaning that several codons may specify a particular amino acid. Different species have unique codon usage biases, implying that they prefer particular codons over other codons that define the same amino acid. An excerpt of a codon usage frequency table is given for the three strains we worked with to highlight different codon biases. | ||
<br><br> | <br><br> | ||
| - | <img src="https://static.igem.org/mediawiki/2014/0/0d/Oxford_codon_optimisation.png" style="float:left;position:relative; width:80%;margin-left:10%;" /> | + | <img src="https://static.igem.org/mediawiki/2014/0/0d/Oxford_codon_optimisation.png" style="float:left;position:relative; width:80%;margin-left:10%;margin-right:10%" /> |
<br><br> | <br><br> | ||
Since we are expressing the dcmA enzyme from M. extorquens in E. coli and P. putida, we used an online tool to optimise the sequence of dcmA so that it can be translated by our host strains. To ensure that dcmA is not over-produced and therefore a strain on the metabolic capacity of the cell, codon usage was only optimised to ~70%. | Since we are expressing the dcmA enzyme from M. extorquens in E. coli and P. putida, we used an online tool to optimise the sequence of dcmA so that it can be translated by our host strains. To ensure that dcmA is not over-produced and therefore a strain on the metabolic capacity of the cell, codon usage was only optimised to ~70%. | ||
Revision as of 21:53, 20 September 2014
#list li { list-style-image: url("https://static.igem.org/mediawiki/2014/6/6f/OxigemTick.png"); } }



























 "
"