Team:Heidelberg/Modeling/Enzyme Modeling
From 2014.igem.org
– Modeling of lysozyme activity with product inhibition
ABSTRACT
The theory of enhancing heatstability by circularization suggests, that the more the ends are constrained, the better stabilization works. Based on this theoretical assumption we identified patterns to have the ability to customly build rigid linkers, that fit between the ends of the protein. Based on over 1000 experimental degradation curves from 9 different constructs with biological and technical replicates of lambda lysozyme, we wanted to gain quantitative insight into the effect of different linkers into the functionality after heatshock. Therefore, a comprehensive approach based on quantitative dynamic modeling (ODEs) was made to determine the relevant information. We started with easy fitting of the data and continued by testing different enzyme kinetics models. We evaluated and identified the most relevant and robust parameters by profile likelihood analysis. Research suggested that a Michaelis-Menten model with substrate inhibition was appropriate for our data. Our final findings show that circularization of lysozyme can have a considerable influence on enzyme thermostability and that suboptimal linker design can decrease thermostability. These results were extraordinarily important for the calibration of CRAUT.
Contents |
Introduction
Enzyme kinetics is a widely studied field in biology [5]. From the derived kinetic parameters one can make many different predictions about the function of a certain enzyme. A commonly used approach for the determination of the enzyme kinetic parameters, is the measurement of the reaction rate in time-dependent manner and with varying substrate concentrations. As this approach would be too laborious to apply in a high throughput manner, we instead decided to record the degradation curves for each lysozyme.
Lysozyme as model enzyme
Lysozyme of the $\lambda$-phage suits well as model for kinetic enzyme studies as it is a well characterized protein. Able to degredade the procaryotic cell wall composed of peptidoglycans. As already stated we anticipated that the lysozyme of the $\lambda$ bacteriophage could reasonably fulfill the requirements for our linker screen.
As described in the Linker screening project description, we try to infer the loss of activity of $\lambda$-lysozyme due to heatshock, by observing the kinetic behavior on the degradation of the peptidoglycane outer layer of M. lysodeikticus. This dynamic process, which ultimately leads to a change of turbidity, is very complex and has been widely discussed for more than 40 years now. On the other hand the activity of lysozyme is highly sensitive to outer conditions like salt concentrations in the media [-1] and the lysozyme concentration itself [0].
We have not only observed the non-enzymatic activity maximum of lysozyme described by Düring et al. [1] but also many observed effects can be explained by applying theory of product inhibition to the kinetics [2]. On the other hand lysozymes unfolding behavior from 37°C seems to be dominated by a rapid collapse when it is denaturated [3].
Michaelis Menten kinetics and Competitive Enzyme Kinetics
Michaelis Menten theory describes the catalytical behaviour of enzymes in simple reactions [4]. It's basic reactions are assumed as \[ E + S \, \overset{k_f}{\underset{k_r} \rightleftharpoons} \, ES \, \overset{k_\mathrm{cat}} {\longrightarrow} \, E + P \] , with E the enzyme, S substrate, ES the enzyme-substrate complex and P the reaction product. $k_r$, $k_f$ and $k_\mathrm{cat}$ are catalytical constants. This means part of the enzyme is always bound in an enzyme substrate complex. This kinetic behavior can be simplified in the basic differential equation: \[\frac{d\left[S\right]}{dt} = \frac{- V _{max} \left[S\right]}{K_m + \left[S\right]} \]. $V_{max}$ is the maximum reaction velocity, obtained from $V_{max} = k_{cat} * E$ and $K_m$ being the michaelis-menten constant
Competitive product inhibition has the effect [5], that part of the Enzyme is also bound in the enzyme-product complex EP. This leads to an apparent increase of $K_m$ as: $K^\text{app}_m=K_m(1+[I]/K_i)$ Thus the differential equation changes as: \[\frac{d\left[S\right]}{dt} = \frac{- V _{max} \left[S\right]}{K_m \left( 1 + \frac{S_0 - S}{k_i} \right) + \left[S\right]} \] where $S_0$ means the substrate concentration at start of the reaction and $k_i$ an inhibitory constant.
Notice that many methods for parameter estimation in these types of models have been developed [6] [7].
Methods
Data
Using the Lysozyme Assay assays we have obtained over 1000 degradation curves for different lysozyme variants. In total, we got more than 100 000 data points from 12 assays performed on 96 well plates. From each well we obtained the degradation curves of M. lysodeiktikus by lysozyme, measured by turbidimetry change at 600 nm. We tested 8 different constructs of circular lysozyme and as reference also linear lysozyme. For all but two constructs, not only technical replicates on one plate were made, but also biological replicates from different growths. On each plate we subjected the lysozymes a heat-shock for one minute at different temperatures. This led to minimally 4 different curves per biological replicate per temperature and per lysozyme.
Each degradation curve consisted in a measurement of the initial substrate concentration withoud lysozyme added, then there is a gap about 2 minutes, varying because of the sequence in that the plate-reader was measuring the wells. After that the degradation was measured every 100 seconds for 100 minutes. The first gap is due to the pipetting step, when adding the enzyme to the substrate and mixing the wells.
Notice, that in regards to conditions used for the measurements, particular care was taken for the following aspects: The reactions always took place at the same temperatures. Also another crucial part was the time after adding the enzyme to the substrate: This was minimized as much as possible and we tried to keep it constant. We always made the dilutions in buffer from the same stock, in order to keep salt concentrations fixed.
OD to concentration calibration
There was performed a measurement for calibrating the $OD_{600}$ to substrate concentration. We have seen that until a substrate concentration of 0.66 mg/ml in the 300 µl wells the behaviour is linear with an offset due to the protein mix and the well plate. We have concentration differences resulting in an $OD_{600}$ difference of: $\delta \mathit{OD} = ((1.160 \pm 0.004 \frac {\mathrm{ml}} {\mathrm{mg}}) * \delta \mathrm{concentration})$. With this result one can easily calculate the concentration differences in each assay. Also the $OD_{600}$ of a well, where all the substrate was completely degraded needed to be measured. We found out, that the influence of the added protein mix on the $OD_{600}$ could be neglected.
Assumptions and data-based considerations
The time between when lysozyme was added to the substrate and the first measurement in the platereader was measured and assumed that it nearly took the same time for each measurement with normally distributed errors. Also, the platereader took about 1s for measuring one well. This delay was also taken into account.
PLE analysis
Often when fitting large models to the data there one has the problem that parameters are connected functionally. The method of Profile likelihood estimation (PLE) enables to reveal of such dependencies. By evaluating the profile likelihood unidentifiable parameters can be grouped into structurally unidentifiable and practically unidentifiable parameters. [8] [9] A parameter is structurally unidentifiable when it is in a functional dependence of one or more other parameters from the model. It is only practically unidentifiable if the experimental data is not sufficient to identify the parameter. This can be easily distinguished from the profile likelihood. By applying PLE analysis and identifying structurally unidentifiable parameters, one is able to reduce the complexity of a given model. In our analysis we relied on d2d Framework, operating on Matlab and providing PLE analysis in an easy to use and fast manner.
Final model
For our model of the degradation we decided to apply product inhibited Michaelis Menten kinetics. A more detailed description on the model development can be found here. As all our data was measured in $OD_{600}$ so at first the substrate concentration had to be calculated. Therefore we include an offset turbidity value, that is due to the turbidity of an empty well and included the OD to substrate calibration. Also the initial substrate concentration was inserted. $V_{Max}$, $K_M$, $K_I$ were the three enzymatical parameters that were fitted. Furthermore the error was fitted automatically too. For temperatures higher than 37.0 °C $V_{Max}$ was replaced by a ratio, called the activity of a temperature. Representing how much activity is left, compared to the activity of 37°C. It was defined by: $V^{lysozyme}_{Max, T} = act^{lysozyme}_T * V^{lysozyme}_{Max, 37.0}$. Mathematically this just meant exchanging one parameter by another for enhanced readability. On the other hand we assumed $K_M$ and $K_I$ to stay the same for different temperatures, but to vary between different lysozyme types. We decided to always fit the data of one plate on its own, because we observed variation in functional behavior between the measurements from the different days. In table 1 it is shown which parameters are fixed for which part of the model.
| span of a parameter | $K_M$ | $K_I$ | $V_{Max}$ | $k_{decay}$ | OD offset | init_Sub | Error |
|---|---|---|---|---|---|---|---|
| Lysozymes | |||||||
| All lysozymes on the same plate | x | ||||||
| Same biological replicates of lysozyme on the same plate | x | x | |||||
| Same biological replicates of lysozyme on the same plate and the same temperature | x | ||||||
| Plate | |||||||
| The same plate | x | ||||||
| All plates | x | x | |||||
Different models tested
During the development of our model, we have tested and compared different models. We tried many models describing the data of all the assays at once. These resulted often in calculations going on for hours. Mainly they were all variations of the final model, always based on product inhibited Michaelis Menten theory. In all the models modeling all the assays, $V_{max}$ was split up into $k_{cat} * E$ where k_{cat} would be the same over different biological replicates and different plates, but E could vary.
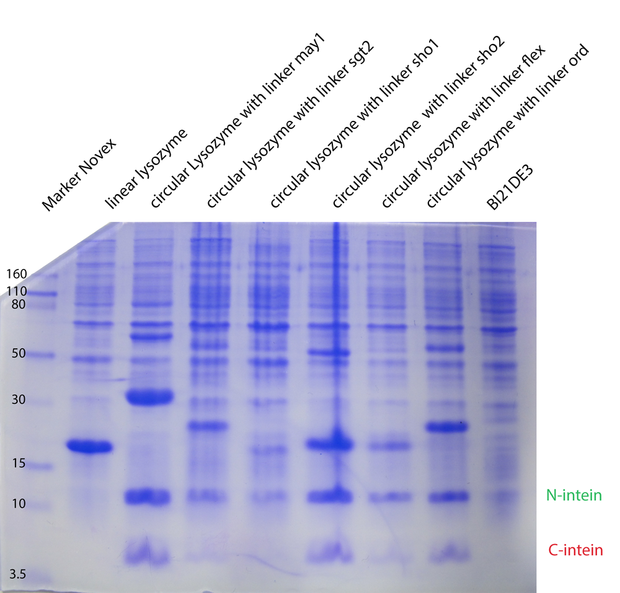
In the second model we have fixed $k_{cat}$ arbitrarily to 1 for all the different enzymes. In the third model we have tried $K_M, K_{cat}, K_I$ fixed for the different temperatures, varying for the different types of lysozymes. In the next model (4) $K_M, K_{cat}, K_I$ were fitted separately for each temperature and each enzyme type. Substantially different was model 5, where we have inserted ratios for the enzyme concentrations. These ratios were obtained from coomassie gels (Fig. 1). Unfortunately no calibration could be made, so we could not introduce concentrations, but just ratios from the different types. For all the models on the whole dataset, the enzyme concentration was fixed between biological replicates.
Model 6 was built to model the kinetics of one single plate. In contrast to the final model, here the kinetic parameters $K_{cat}, K_I$ were fitted for each temperature separately.
Results
To analyze the effect of circularization on the thermostability of the lysozyme variants, the heat shock dependent reaction rate parameters $v_{max}$ for all lysozyme variants had to be identified. For this purpose we analyzed the observed substrate degradation dynamics for the different lysozyme variants by ODE modeling. As detailed in the introduction, the enzymatic reaction mechanism of the lambdaphage lysozyme can be described by Michaelis-Menten kinetics with product inhibition. Furthermore, experiments on pH-dependent lysozyme degradation have shown that lysozyme exists in two distinct states when challenged with pH changes: the normal, functional state and a denatured, nonfunctional state [3]. We hypothesized that lysozyme deformation under heat shock conditions could be described by a similar shift from a functional conformation to a distinct, denatured state. Consequently, enzymatic activity after heat shock was assumed to be exerted by only one, homogeneous, population of functional lysozymes, differing in size depending on heat shock intensity. Because the structure of the active enzyme species was assumed to be identical independent of the applied heat shock, the kinetic parameters of the enzymatic reactions could be assumed to be independent of heat shock intensity. Therefore, based on this model of enzyme denuration, enzymatic activity after heat shock could be assumed to be only dependent on the remaining fraction of functional lysozymes.
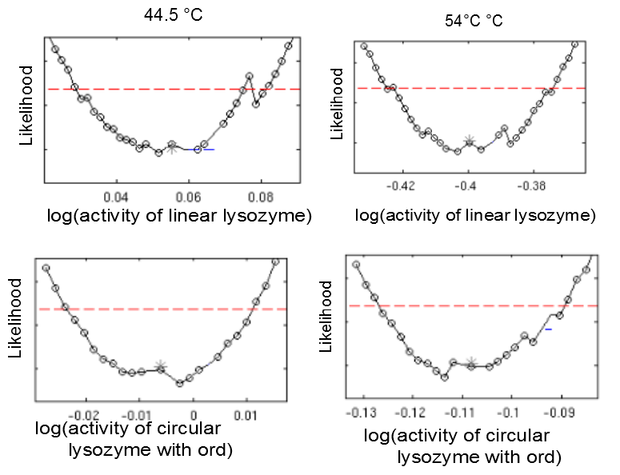
This model was fitted to all available data, using simultaneous multi-model fitting where appropriate. The model could emulated the substrate degradation dynamics for all lysozyme variants (Fig 2). Profile likelihood-based identifiability analysis was employed to verify practical identifiability of the relevant kinetic parameters. While the kinetic parameters representing enzyme affinity for the substrate and the inhibitors could not be identified in the model, the maximal reaction rate $v_{max}$ where identifiable in all cases (Fig 3). The complete result of the profile likelihood analysis can be found here.
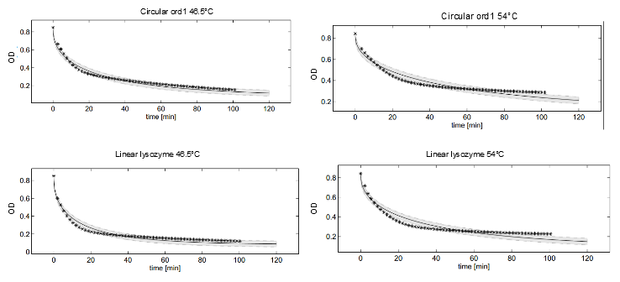
Dynamics of peptidoglycan degradation by the lambdaphage lysozyme can be emulated by a simple model assuming Michaelis-Menten kinetics with competitive product inhibition. The model was implemented with the assumption that lambdaphage lysozyme exists in two distinct states – functional or deformed - after heat shock within the considered range of intensities (citation). Following this assumption, kinetic parameters of the enzymatic reaction can be assumed to be independent of heat shock intensity. Thus, model complexity is considerably reduced, as explained in detail in the text. Exemplary measurements of peptidoglycan degradation by the linear lysozyme (a) and by a circularized lysozyme with the ord1 linker (b) are shown together with model fits. Substrate degradation is shown for basal enzyme activity after 1 min incubation at 44.5 °C and for diminished activity after 10 min incubation at 54 °C.
To compare thermostability of the different lysozyme variants, we analyzed the relationship between heat shock intensity and loss of enzymatic activity. As a measure for enzymatic activity, we used the normalized maximal reaction rate (the ratio of the enzymatic activity after heat shock and the basal enzymatic activity after incubation at 37 °C). Heat-shock dependent loss of enzymatic activity differed considerably between the different lysozyme variants (Fig 4). For a direct comparison of lysozyme variant thermostability we sought a robust statistic characterizing heat-shock resistance. This statistic should incorporate the threshold heat-shock intensity upon which significant loss of activity occurs as well as the steepness of the heat-shock intensity dependent loss of activity. We decided to focus on the heat-shock intensity window where most of the enzymatic activity was lost (45 °C to 57 °C).
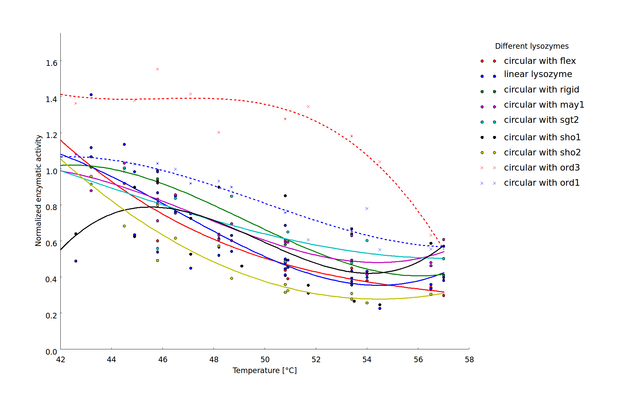
Heat-shock dependent enzyme activity for the linear lysozyme and 8 circularized lysozyme variants. Enzymatic activity is described here as the normalized maximal reaction rates, computed as the ratio of the maximal reaction rate after heat shock at the respective temperature and the maximal reaction rate after incubation at 37 °C. Two biological replicates were available for 7 of the 9 lysozyme variants and the $v_{max}$ values computed for each replicate are plotted separately. Temperature dependent decrease of the enzyme activity was fitted by splines to provide a better visualization of the relationship of heat shock intensity and enzyme deformation.
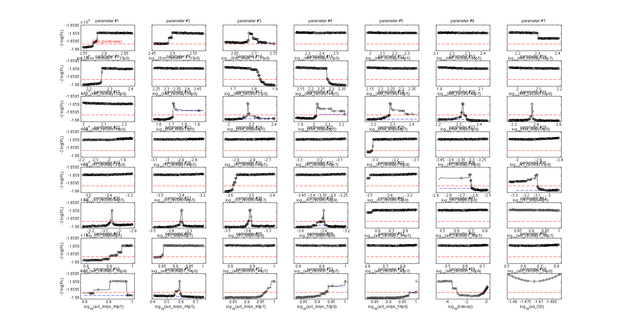
Introduction of heat shock dependent reaction rates does not significantly improve the model fit. It was tested whether the model fit could be improved by assuming that heat shock induced enzyme deformation occurs gradually and not in distinct stages. In this case, the kinetic parameters of the enzymatic activity are dependent on the heat shock intensity.
Finally, we tested whether the mechanistic assumption of a distinct transition between a single active and inactive state upon heat shock had affected the quality of the model fit. The alternative hypothesis concerning the mechanism of enzyme deformation would allow for continuous changes of the lysozyme structure in response to heat shock intensity. Thus, a gradual shift towards more deconformed structures would be expected for higher heat shock intensities. This would result in different kinetic parameters for the same lysozyme species under differing heat shock treatment. To test the effect of implementing this alternative deconformation mode in the model, model fitting was repeated with independent kinetic parameters for different heat shock intensities. Manual inspection of the fitting results did not show a better fit to the data. However, freeing the kinetic parameters resulted in a loss of parameter identifability (Fig. 5). Therefore, the increased number of kinetic parameters was considered to negatively affect the usability of the model and the original, parameter-reduced, model structure was retained for analysis.
Discussion
Using dynamic ODE modeling, we could extract the heat-shock dependent maximal reaction rates of different lysozyme variants from simple substrate degradation measurements. The $v_{max}$ parameters were identifiable in spite of the complex reaction mechanism of the lysozyme. This allowed us to compute a normalized enzymatic activity for all lysozyme variants after a variety of different heat shock challenges. By comparing these enzymatic activities, thermostability of the different lysozymes variants could be directly compared.
Our findings show that circularization of the lysozyme can have a considerable influence on enzyme thermostability. Similar findings have been reported for a variety of other proteins (sources). Here, we extend previous findings by demonstrating that the effect of circularization strongly depends on the chosen linker structure. Suboptimal linker design can decrease thermostability. The most evident example in the findings presented here is the sho2 linker which was chosen for testing as an example for linkers too short to bridge the natural distance between the C- and N-terminus of the lysozyme. In silico guided design of optimized linker sequences on the other hand can indeed result in increased thermostability, as demonstrated by the ord1 and ord3 linkers. These linkers where chosen as examples for linkers with a very low likelihood of crossing the active center of the enzyme. The implications of this analysis for the linker design are discussed in more detail in the documentation of the linker design software here.
References
[-1] Mörsky, P. Turbidimetric determination of lysozyme with Micrococcus lysodeikticus cells: reexamination of reaction conditions. Analytical biochemistry 128, 77-85 (1983).
[0] Friedberg, I. & Avigad G. High lysozyme concentration and lysis of Micrococcus lysodeikticus, Biochim. Biophys. Acta, 127 (1966) 532-535
[1] Düring, K., Porsch, P., Mahn, A., Brinkmann, O. & Gieffers, W. The non-enzymatic microbicidal activity of lysozymes. FEBS Letters 449, 93-100 (1999).
[2] Colobert, L. & Dirheimer G. Action du lysozyme sur un substrat glycopeptidique isolé du micrococcus lysodeiktikus. B1OCHIMICA ET BIOPHYSICA ACTA, 54, 455-468 (1961)
[3] Di Paolo, A., Balbeur, D., De Pauw, E., Redfield, C. & Matagne, A. Rapid collapse into a molten globule is followed by simple two-state kinetics in the folding of lysozyme from bacteriophage λ. Biochemistry 49, 8646-8657 (2010).
[4] Hommes, F. A. "The integrated Michaelis-Menten equation." Archives of biochemistry and biophysics 96.1 (1962): 28-31.
[5] Purich, Daniel L. Contemporary Enzyme Kinetics and Mechanism: Reliable Lab Solutions. Academic Press, 2009.
[6] Liao, Fei, et al. "The comparison of the estimation of enzyme kinetic parameters by fitting reaction curve to the integrated Michaelis–Menten rate equations of different predictor variables." Journal of biochemical and biophysical methods 62.1 (2005): 13-24.
[7] Goudar, Chetan T., Jagadeesh R. Sonnad, and Ronald G. Duggleby. "Parameter estimation using a direct solution of the integrated Michaelis-Menten equation." Biochimica et Biophysica Acta (BBA)-Protein Structure and Molecular Enzymology 1429.2 (1999): 377-383.
[8] Raue, A. et al. Lessons Learned from Quantitative Dynamical Modeling in Systems Biology. PLoS ONE 8, (2013).
[9] Raue, a et al. Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics 25, 19239 (2009).
 "
"