Achievements
Characterization of GFP-REACh1 and GFP-REACh 2 together with an IPTG inducible TEV protease
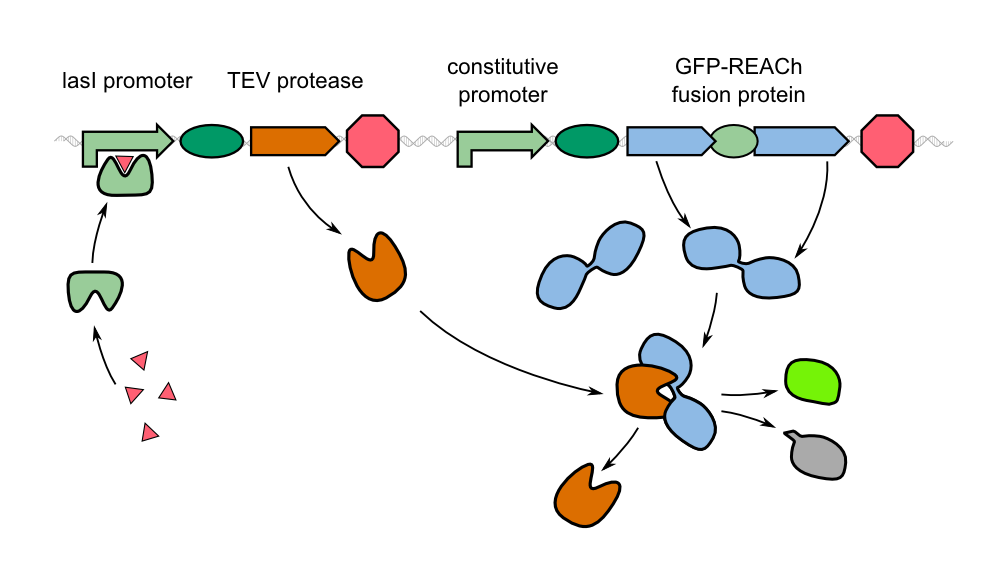
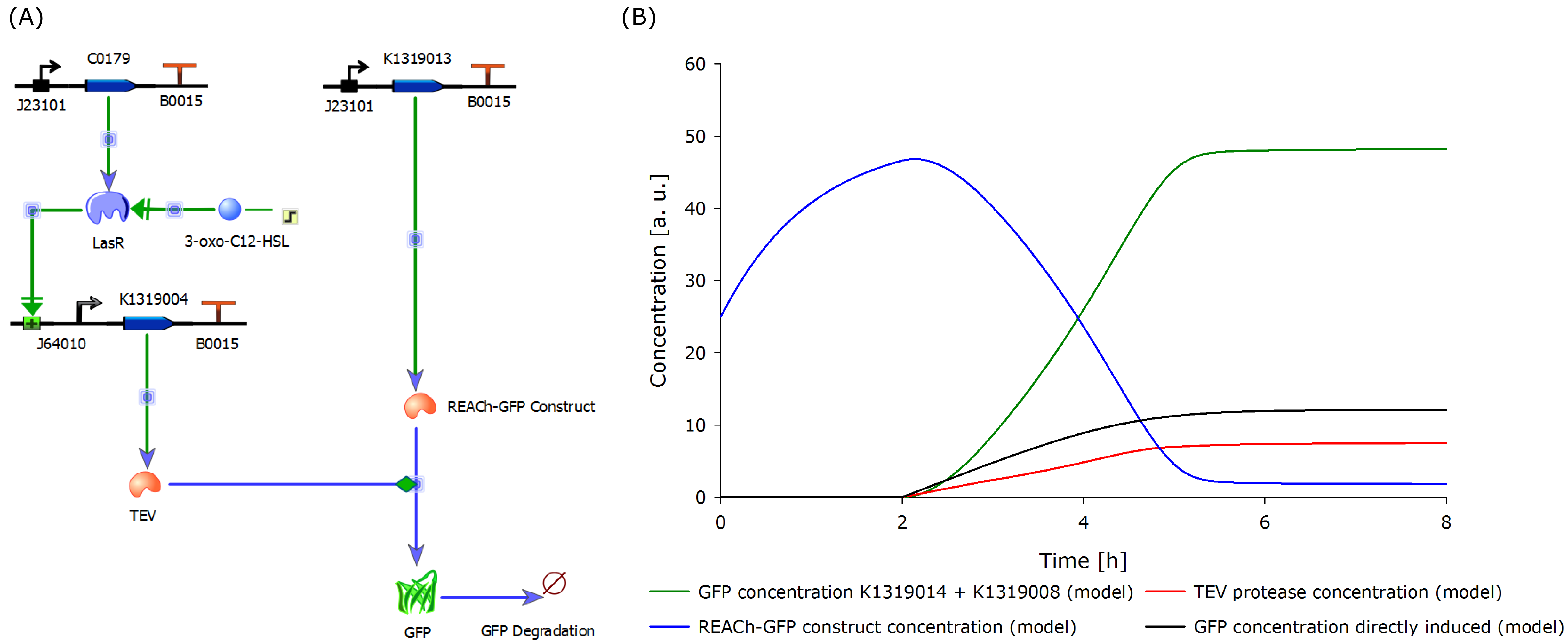
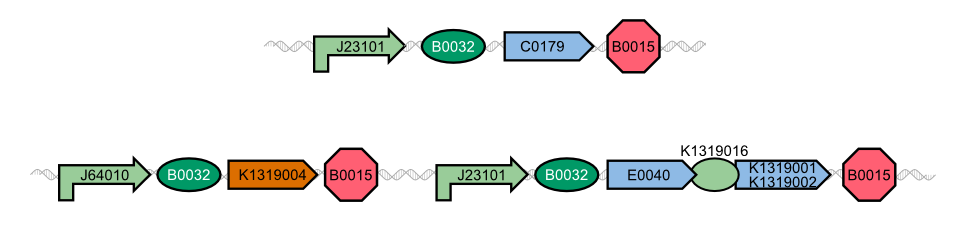
The characterization of the TEV protease and the REACh 1 and REACh 2 dark quencher proteins was performed by introducing both simultaneously into E. coli. The resulting double plasmid cells therefore contained [http://parts.igem.org/Part:BBa_K1319013 K1319013] (GFP-REACh 1 fusion protein) and [http://parts.igem.org/Part:BBa_K1319008 K1319008] (IPTG inducible TEV protease) or [http://parts.igem.org/Part:BBa_K1319014 K1319014] (GFP-REACh 2 fusion protein) and K1319008 respectively. K1319013 and K1319014 were situated on the pSB3K3 plasmid backbone and K1319008 on the pSB1C3 backbone, two standard plasmids with different oris allowing their simultaneous use in one cell.
As a positive control [http://parts.igem.org/Part:BBa_I20260 I20260] was used. I20260 contains the same promoter ([http://parts.igem.org/Part:BBa_J23101 J23101]] and RBS ([http://parts.igem.org/Part:BBa_B0032 B0032]) as well as the same fluorescence protein (GFP, [http://parts.igem.org/Part:BBa_E0040 E0040]) and is located on the same plasmid backbone (pSB3K3). Therefore it is expected that when all fusion proteins are successfully cut by the TEV protease, the fluorescence level of the double plasmid constructs reaches the same level as the positive control of I20260. As a negative control we used [http://parts.igem.org/Part:BBa_B0015 B0015], a coding sequence of a Terminator which shouldn't show any sign of fluorescence.
To better evaluate the fluorescence we adjusted the fluorescence reading with the observed OD to achieve a fluorecence reading independent of the amount of cells present but rather representative of the amount of fluorescence per cell. With this the fluorescence of a constitutive expression, no expression and both of the REACh + TEV constructs induced and not induced are observed. All measurements were done in a biological triplicate.
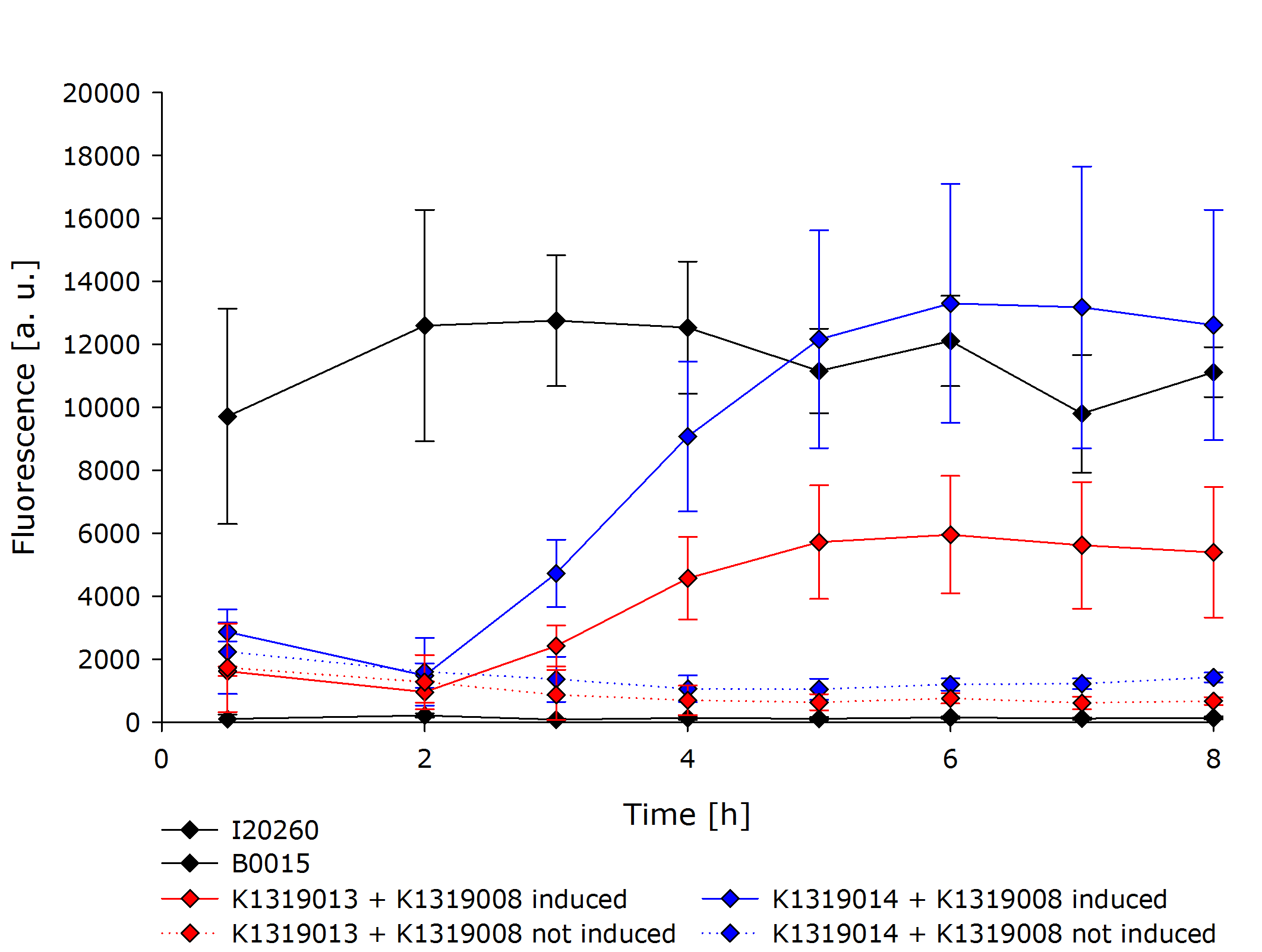
The negative control B0015 does not exhibit any significant fluuorescence as expected. The positive control I20260 shows a steady increase in fluorescence for the first X hours. After that the fluorescence stays constant due to the end of the exponential growth phase and the cells becoming stationary and not producing any more GFP. The production is also indpendent od the induction with IPTG as it is expected.
Both double plasmid constructs K1319013 + K1319008 and K1319014 + K1319008 don't exhibit a strong fluorescence before induction with IPTG. In the non induced constructs the fluorescence stays low and only increases slightly over time. It is severely weaker than the fluorescence reached by the induced constructs or the positive control but also higher then the negative control. This shows that the promoter system used is not completely shut down without induction but significantly weaker compared with the induced constructs. It is also a partly due to an imperfect dark quenching of GFP by the REACh 1 and REACh 2 proteins.
The induced double plasmid constructs exhibit a fast rise in fluorescence after induction up to an increase of over 10 fold compared to the non induced constructs. K1319013 + K1319008 reaches the the same level of fluorescence as I20260 indicating a complete cutting of the fusion proteins by the TEV protease. K1319014 + K1319008 doesn't reach the same level of fluorescence but the nearly 10 fold increase in fluorescence is a clear indicator for the TEV protease cutting the fusionprotein prodiced by K1319014. the same level of fluorescence as a the positive control is not achieved probably due to generally lower expression level of K1319014 in the cells.
Summary
The double plasmid systems of K1319013 + K1319008 as well as K1319014 + K1319008 clearly demonstrate the Quenching ability of the REACh 1 and REACh 2 proteins as well as the funcionality of the TEV protease. Both REACH 1 and REACH 2 show a significant quenching ability of GFP shown in the difference of fluorescence between the positive control I20260 and the non induced double plasmid systems. This is also comfirmed by the resulting fluorescence after induction showing that the TEV protease is successfully able to cut the fusion proteins as well as the proper expression of both fusion proteins. Combined this characterization shows a validation of the functionality of the REACh 1 protein ([http://parts.igem.org/Part:BBa_K1319001 K1319001]), the REACh 2 protein ([http://parts.igem.org/Part:BBa_K1319002 K1319002]) and the TEV protease ([http://parts.igem.org/Part:BBa_K1319004 K1319004]).
Comparing the kinetic of the GFP-REACh fusion proteins with a standard lacI inducible GFP epression
To assess the kinetic of the fusion proteins K1319013 (GFP-REACh 1) and K1319014 (GFP-REACh 2) the double plasmid systems of K1319013 + K1319008 and K1319014 + K1319008 were compared to a standard expression of GFP under the control of a lacI promoter in [http://parts.igem.org/Part:BBa_K731520 K731520] made by the iGEM Team TRENTO in 2012 to evaluate the kinetic prediction of an a faster fluorescence response with our construct compared to a normal expression.
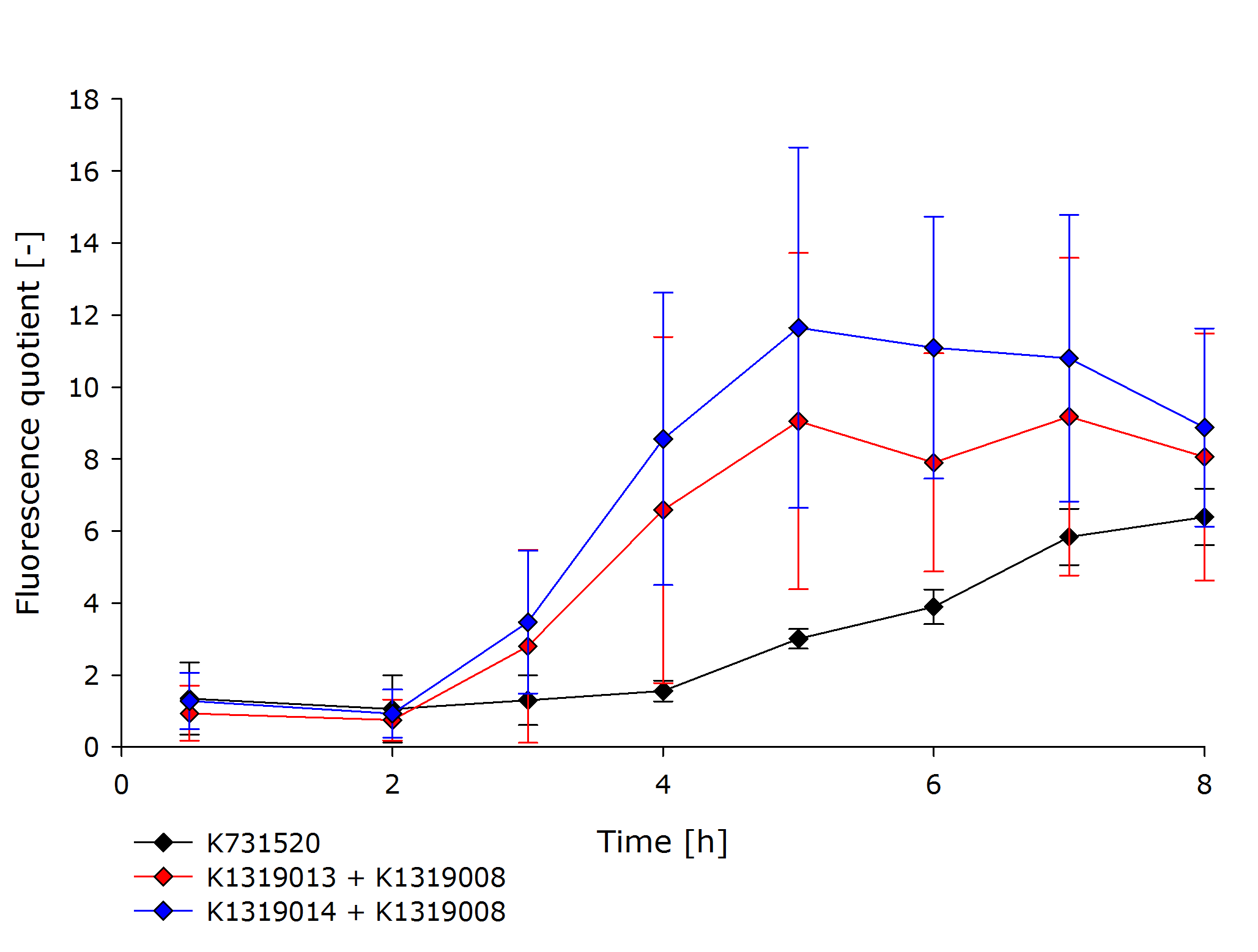
Therefore K731520 and the same double plasmid construct that was described earlier (K1319013 + K1319008 and K1319014 + K1319008) were cultivated in E. coli BL21(DE3) and fluorescence and OD were measured. Once again the fluorescence was adjusted for the OD to show a relative fluorescence on a cell per cell basis. Also it was especially looked at the difference between the induced and not induced state. This difference (fluorescence quotient) gives a better indicator for a system which is used as a sensor because the difference between an on and off state is more important for a clear and unmistakable signal compared to the overall fluorescence. Hence the OD adjusted fluorescence quotient for both double plasmid constructs (K1319013 + K1319008 and K1319014 + K1319008) and K731520 was obtained and plotted in the following graphic.
The graphic clearly shows the faster kinetic of the cut GFP-REACh fusion protein compared to a standard GFP expression. Both fluorescence signals of the double plasmid constructs achieve a higher difference in fluorescence signal netween induced and non induced state as well as at a faster rate. This proves the earlier made hypothesis of the kinetic of the GFP-REACh fusion protein combined with a TEV protease.
summary
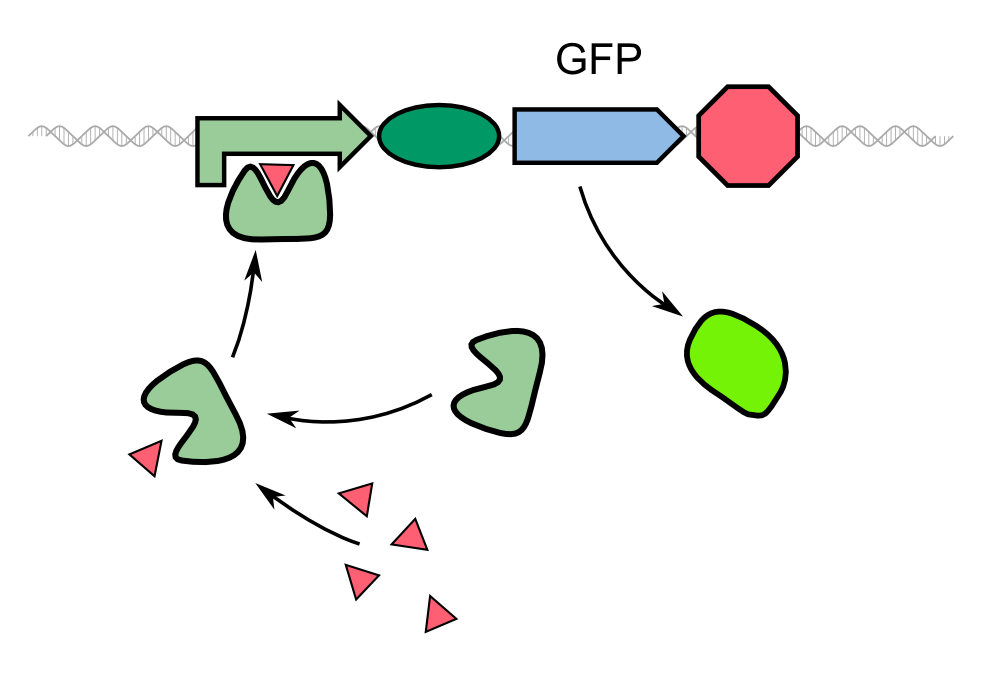
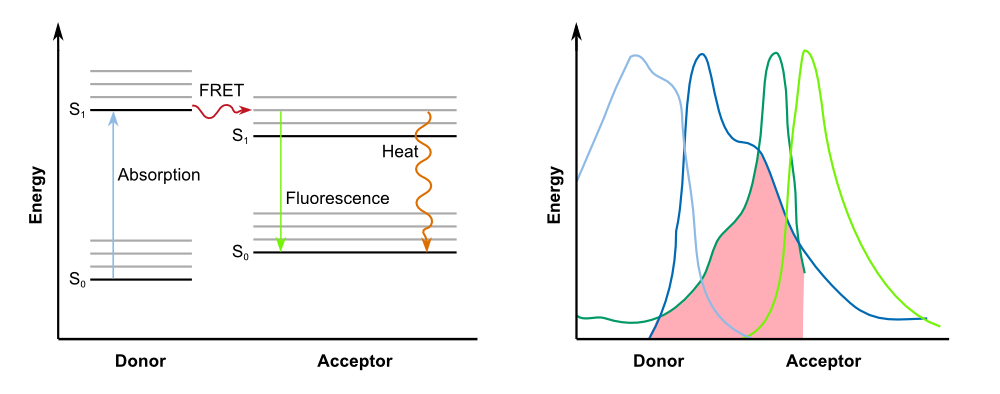
The kinectic of the fusion protein combined with the TEV protease exhibits the exact characteristics as predicted earlier. The response is clearly faster than normal expression by accumulating a reservoir of fusion proteins who are not fluorescing due to the dark quencher attaches to it. This reservoir is then activated by the induction of the TEV protease which results in the cutting of the fusion protein, releasing GFP from the dark quencher and disturbing the FRET mechanims between it and GFP. This results in the observed faster fluorescence reaction due to multiplicating effect by the TEV protease in which every one TEV protease can account for many fluorescence proteins being activated.
Characterizing the GFP-REACh constructs in sensor chips
lorem ipsum...
|
 "
"