Team:ETH Zurich/modeling/whole
From 2014.igem.org
Whole cell model
Model
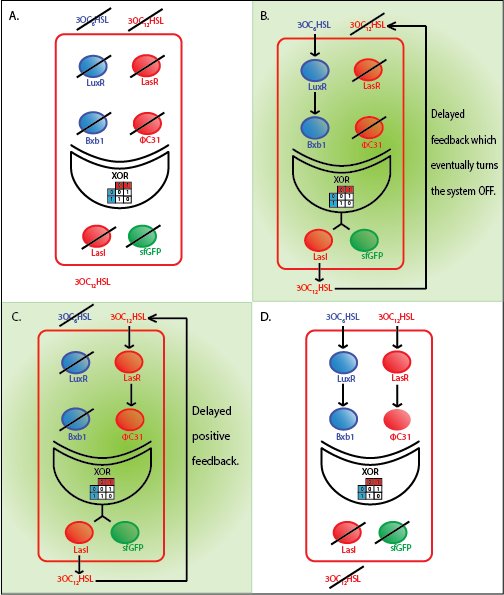
The whole cell model is the combination of the Quorum sensing, Integrase and XOR modules. The model shows the behaviour of the a single cell in response to incoming signals. The model enables us to understand the effect of leakiness, cross-talk and their combinations on the whole system. All the simulations here are described for a red cell .i.e. a cell producing LasI and GFP when it is ON.
Chemical Species
| Name | Description |
|---|---|
| LuxAHL | 30C6-HSL is an acyl homoserine lactone which mainly binds to LuxR. |
| LuxR | Constitutively expressed regulator protein that can bind LuxAHL and stimulate transcription of Bxb1. |
| RLux | LuxR and LuxAHL complex which can dimerize. |
| DRLux | Dimerized form of RLux. |
| mRNABxb1 | mRNA of the Bxb1 integrase being transcribed by the Lux promoter. |
| Bxb1 | Serine integrase that can fold into two conformations - Bxb1a and Bxb1b. We chose to use a common connotation for both conformations - Bxb1. |
| DBxb1 | Dimerized form of Bxb1. We chose to use a common connotation for both homodimers, DBxb1a and DBxb1b. |
| LasAHL | 30C12-HSL is an acyl homoserine lactone which mainly binds to LasR. |
| LasR | Constitutively expressed regulator protein that can bind LasAHL and stimulate transcription of ΦC31. |
| RLas | LasR and LasAHL complex which can dimerize. |
| DRLas | Dimerized form of RLas. |
| mRNAΦC31 | mRNA of the ΦC31 integrase being transcribed by the Lux promoter. |
| ΦC31 | Serine integrase that can fold into two conformations - ΦC31a and ΦC31b. We chose to use a common connotation for both conformations - ΦC31. |
| DΦC31 | Dimerized form of ΦC31. We chose to use a common connotation for both homodimers, DΦC31a and DΦC31b. |
| SIBxb1 | Inactive DNA binding site for Bxb1. No dimer is bound to this site. |
| SABxb1 | Active DNA binding site for Bxb1. A dimer is bound to this site. |
| SFBxb1 | Flipped DNA binding site for Bxb1. The site generated after recombination by integrase. |
| SIΦC31 | Inactive DNA binding site for ΦC31. No dimer is bound to this site. |
| SAΦC31 | Active DNA binding site for ΦC31. A dimer is bound to this site. |
| SFΦC31 | Flipped DNA binding site for ΦC31. The site generated after recombination by integrase. |
| Ton,i | The number of terminators which are blocking the transcription of GFP and LuxI/LasI initially. |
| ToffBxb1 | The number of terminators turned off by recombination due to Bxb1. Favours the transcription of GFP and LuxI. |
| ToffΦC31 | The number of terminators turned off by recombination due to ΦC31. Favours the transcription of GFP and LuxI. |
| Ton,f | The number of terminators blocking the transcription of GFP and LuxI/LasI after recombination by Bxb1 and ΦC31. No transcription. |
| mRNAGFP | mRNA for Green fluorescent protein which is produced when the cells are ON. |
| GFP | Green fluorescent protein which is produced when the cells are ON. |
| mRNALuxI | mRNA for LuxI which is produced when the cells are ON. |
| LuxI | Enzyme catalysing the production of LuxAHL from SAM and ACP. |
| mRNALasI | mRNA for LasI which is produced when the cell are ON. |
| LasI | Enzyme catalysing the production of LasAHL from SAM and ACP. |
Reactions
$$ \begin{align} \emptyset&\rightarrow LuxR \\ LuxAHL+LuxR & \leftrightarrow RLux\\ RLux+RLux &\leftrightarrow DRLux\\ DRLux+P_{luxOFF} & \leftrightarrow P_{luxON}\\ P_{luxON}&\rightarrow P_{luxON}+mRNA_{Bxb1}\\ mRNA_{Bxb1}&\rightarrow Bxb1\\ Bxb1 + Bxb1 &\leftrightarrow DBxb1 \\ DBxb1 + SI_{Bxb1} & \leftrightarrow SA_{Bxb1}\\ LuxAHL &\rightarrow \emptyset\\ LuxR &\rightarrow \emptyset\\ RLux &\rightarrow \emptyset\\ DRLux &\rightarrow \emptyset\\ mRNA_{Bxb1} &\rightarrow \emptyset\\ Bxb1 &\rightarrow \emptyset\\ DBxb1 &\rightarrow \emptyset\\ \\ \emptyset&\rightarrow LasR \\ LasAHL+LasR & \leftrightarrow RLas \\ RLas+RLas & \leftrightarrow DRLas\\ DRLas+P_{LasOFF} & \leftrightarrow P_{LasON}\\ P_{LasON}&\rightarrow P_{LasON}+mRNA_{\phi C31}\\ mRNA_{\phi C31}&\rightarrow \phi C31\\ \phi C31 + \phi C31 &\leftrightarrow D\phi C 31 \\ D\phi C 31 + SI_{\phi C31} & \leftrightarrow SA_{\phi C31}\\ LasAHL &\rightarrow \emptyset\\ LasR &\rightarrow \emptyset\\ RLas &\rightarrow \emptyset\\ DRLas &\rightarrow \emptyset\\ mRNA_{\phi C31} &\rightarrow \emptyset\\ \phi C31 &\rightarrow \emptyset\\ D\phi C31 &\rightarrow \emptyset\\ \\ SA_{Bxb1}+SA_{Bxb1}+T_{on,i}& \rightarrow T_{offBxb1}+ SF_{Bxb1}+SF_{Bxb1}\\ T_{offBxb1} &\rightarrow T_{offBxb1} + mRNA_{GFP} + mRNA_{LasI} \\ SA_{\phi C31}+SA_{\phi C31}+T_{on,i}& \rightarrow T_{off\phi C31}+SF_{\phi C31}+SF_{\phi C31}\\ T_{off\phi C31} &\rightarrow T_{off\phi C31} + mRNA_{GFP} + mRNA_{LasI} \\ SA_{Bxb1}+SA_{Bxb1}+T_{off,\phi C31}& \rightarrow T_{on,f}+SF_{Bxb1}+SF_{Bxb1}\\ SA_{\phi C31}+SA_{\phi C31}+T_{offBxb1}& \rightarrow T_{on,f}+SF_{\phi C31}+SF_{\phi C31}\\ mRNA_{GFP} &\rightarrow GFP\\ mRNA_{LasI} &\rightarrow LasI\\ mRNA_{GFP} &\rightarrow \emptyset\\ mRNA_{LasI}&\rightarrow \emptyset\\ GFP &\rightarrow \emptyset\\ LasI&\rightarrow \emptyset\\ \\ S_{LasI}+LasI & \rightarrow LasAHL\\ \end{align}$$
Equations
The following equations were used for the whole cell model. The production of Bxb1 and ΦC31 is reduced to a hill function of RLux an RLas respectively. For details, check Quorum sensing module.
The equation is for a system which produces LasI and GFP as outputs. The LasI catalyses the production of LasAHL from SAM and ACP. Since, we do not know the amount of ACP or SAM present in the cell, we assume the rate of production of LasAHL to depend only on LasI concentration.
$$\begin{align*}
\frac{d[LuxAHL]}{dt} &= -d_{LuxAHL}[LuxAHL]\\
\frac{d[LuxR]}{dt} &= \alpha_{LuxR} -k_{RLux}[LuxAHL][LuxR] + k_{-RLux}[RLux] - d_{LuxR}[LuxR] \\
\frac{d[RLux]}{dt} &= k_{RLux}[LuxAHL][LuxR] - k_{-RLux}[RLux] - d_{RLux} [RLux] \\
\frac{d[mRNA_{Bxb1}]}{dt} &= L_{P_{Lux}} + \frac{K_{mRNA_{Bxb1}}[RLux]^2}{K_{mLux}^2 + [RLux]^2 }- d_{mRNA_{Bxb1}} [mRNA_{Bxb1}]\\
\frac{d[Bxb1]}{dt} &= k_{Bxb1} [mRNA_{Bxb1}] -2 k_{DBxb1}[Bxb1]^2+ 2 k_{-DBxb1}[DBxb1] - d_{Bxb1}[Bxb1]\\
\frac{d[DBxb1]}{dt}&= k_{DBxb1}[Bxb1]^2-k_{-DBxb1}[DBxb1]-k_{SABxb1}[DBxb1][SI_{Bxb1}]+k_{-SABxb1}[SA_{Bxb1}]-d_{DBxb1}[DBxb1]\\
\frac{d[SI_{Bxb1}]}{dt}&=-k_{SABxb1}[DBxb1][SI_{Bxb1}]+k_{-SABxb1}[SA_{Bxb1}]\\
\frac{d[SA_{Bxb1}]}{dt}&=k_{SABxb1}[DBxb1][SI_{Bxb1}]-k_{-SABxb1}[SA_{Bxb1}] - 2 k_{ToffBxb1} [SA_{Bxb1}]^2 [T_{on,i}] - 2 k_{-Toff\phi C31} [SA_{Bxb1}]^2 [T_{off\phi C31}]\\
\frac{d[SF_{Bxb1}]}{dt}&= 2 k_{ToffBxb1} [SA_{Bxb1}]^2 [T_{on,i}] + 2 k_{-Toff\phi C31} [SA_{Bxb1}]^2 [T_{off\phi C31}]\\
\\
\frac{d[LasAHL]}{dt} &= -d_{LasAHL}[LasAHL] + DF * k_{LasAHL} [LasI]\\
\frac{d[LasR]}{dt} &= \alpha_{LasR} -k_{RLas}[LasAHL][LasR] + k_{-RLas}[RLas] - d_{LasR}[LasR] \\
\frac{d[RLas]}{dt} &= k_{RLas}[LasAHL][LasR] - k_{-RLas}[RLas] - d_{RLas} [RLas] \\
\frac{d[mRNA_{\phi c31}]}{dt} &= L_{P_{Las}} + \frac{K_{mRNA_{\phi C31}}[RLas]^2}{K_{mLas}^2 + [RLas]^2 } - d_{mRNA_{\phi c31}} [mRNA_{\phi c31}]\\
\frac{d[\phi c31]}{dt} &= k_{\phi c31} [mRNA_{\phi c31}] -2 k_{D\phi c31}[\phi c31]^2+ 2 k_{-D\phi c31}[D\phi c31] - d_{\phi c31}[\phi c31]\\
\frac{d[D\phi c31]}{dt}&= k_{D\phi c31}[\phi c31]^2-k_{-D\phi c31}[D\phi c31]-k_{SA\phi c31}[D\phi c31][SI_{\phi c31}]+k_{-SA\phi c31}[SA_{\phi c31}]-d_{D\phi c31}[D\phi c31]\\
\frac{d[SI_{\phi c31}]}{dt}&=-k_{SA\phi c31}[D\phi c31][SI_{\phi c31}]+k_{-SA\phi c31}[SA_{\phi c31}]\\
\frac{d[SA_{\phi C31}]}{dt}&=k_{SA\phi C31}[D\phi C31][SI_{\phi C31}]-k_{-SA\phi C31}[SA_{\phi C31}] - 2 k_{Toff\phi C31} [SA_{\phi C31}]^2 [T_{on,i}] - 2 k_{-ToffBxb1} [SA_{\phi C31}]^2 [T_{offBxb1}]\\
\frac{d[SF_{\phi C31}]}{dt}&= 2 k_{Toff\phi C31} [SA_{\phi C31}]^2 [T_{on,i}] + 2 k_{-ToffBxb1} [SA_{\phi C31}]^2 [T_{offBxb1}]\\
\\
\frac{d[T_{on,i}]}{dt}&= -k_{ToffBxb1}[T_{on,i}][SA_{Bxb1}]^2 -k_{Toff\phi C31} [T_{on,i}][SA_{\phi C31}]^2 \\
\frac{d[T_{offBxb1}]}{dt}&=k_{ToffBxb1} [T_{on,i}][SA_{Bxb1}]^2 - k_{-ToffBxb1} [T_{offBxb1}] [SA_{\phi C31}]^2\\
\frac{d[T_{off\phi C31}]}{dt}&=k_{Toff\phi C31} [T_{on,i}][SA_{\phi C31}]^2 - k_{-Toff\phi C31} [T_{offBxb1}] [SA_{Bxb1}]^2\\
\frac{d[T_{on,f}]}{dt}&= k_{-ToffBxb1} [T_{offBxb1}] [SA_{\phi C31}]^2 + k_{-Toff\phi C31} [T_{offBxb1}] [SA_{Bxb1}]^2\\
\\
\frac{d[mRNA_{LasI}]}{dt}&= k_{mRNA_{LasI}} (\frac{[T_{offBxb1}]}{\theta + [T_{offBxb1}]} + \frac{[T_{off\phi C31}]}{\theta + [T_{off\phi C31}]}) - d_{mRNA_{LasI}}[mRNA_{LasI}]\\
\frac{d[LasI]}{dt}&= k_{LasI} [mRNA_{LasI}] - d_{LasI}[LasI]\\
\frac{d[mRNA_{GFP}]}{dt}&= k_{mRNA_{GFP}} (\frac{[T_{offBxb1}]}{\theta + [T_{offBxb1}]} + \frac{[T_{off\phi C31}]}{\theta + [T_{off\phi C31}]}) - d_{mRNA_{GFP}}[mRNA_{GFP}]\\
\frac{d[GFP]}{dt}&= k_{GFP} [mRNA_{GFP}] - d_{GFP}[GFP]\\
\end{align*}$$
Dynamics
Ideal case
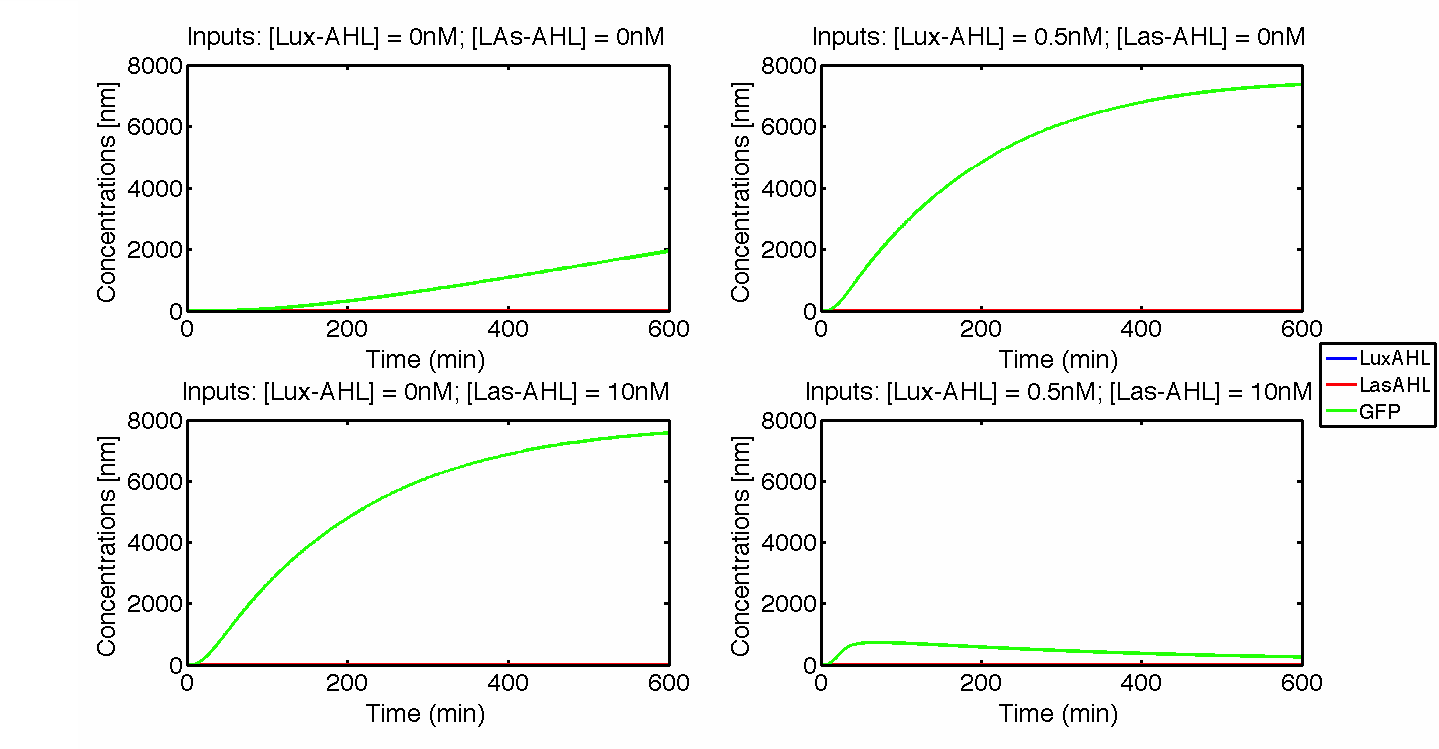
Under ideal conditions, there is no leakiness or cross-talk. The figure shows the production of GFP as a function of four different combinations of inputs in an ideal whole cell. No GFP is produced when there is no LasAHL and LuxAHL or when both are present. GFP is produced only when one of the input AHLs are present, thus emulating an XOR system. In the case with 0.5 nM of LuxAHL and no LasAHL as input, the rate of production of GFP reduces after six hours. This could be attributed to the delayed production of LasAHL from LasI. In the case with only LasAHL as input, there is a positive feedback of LasAHL.
In the case above, the cell receives only 0.5 nM of LuxAHL and no LasAHL but the cell produces LasI upon flipping of the terminator. The LasI produced in turn catalyses the production of LasAHL. The LasAHL activates PLas thus producing ΦC31. ΦC31 results in flipping of the terminator again thus, turning OFF the system. The dynamics of Toni, ToffBxb1, ToffΦC31 and Tonf is summarised in the figure above.
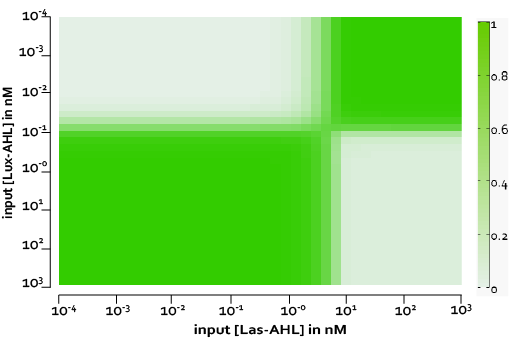
The figure above, shows the predicted XOR gate performance by a single cell model with no basal leakiness for different concentrations of LasAHL and LuxAHL.
With Leakiness
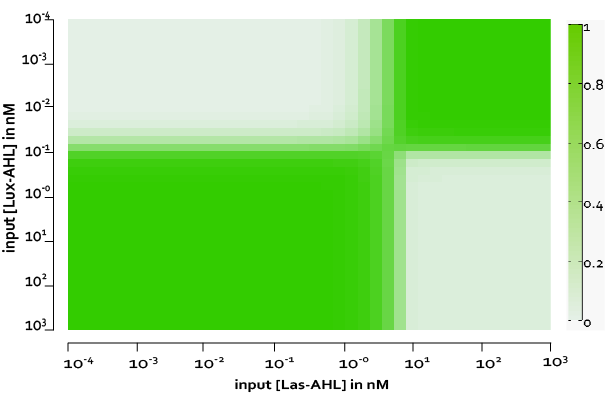
From our experimental data, we observed some basal leakiness for PLux and PLas even after using riboregulators. From the model we see that, this small basal leakiness is amplified downstream. The basal leakiness results in production of integrases which further act on the XOR module and cause the switching of the terminator. Thus, there is GFP and LasI produced, and the LasI produced further catalyses the production of more LasAHL. Thus, we observe some GFP even without inputs.
However, if we measure the fluorescence at around 300 mins, we observe a good and acceptable XOR behaviour. Therefore, one of the solutions we propose is to kill or freeze the cells in each row after 3 hours.
Initially from our model we observed that the feedback was rapid and hence, the amplification was much higher. However, from literature [9] we see that the XOR module is relatively slow. We were able to correct this by modelling transcription and translation steps. The delay introduced seems more reliable although we do not have our own experimental data to validate the same.
Alternate Design
We collaborated with the Edinburgh iGEM team 2014. Their project is focused on communication between bacteria using metabolites as communicating molecules. Combining one metabolite molecule with another quorum sensing molecule could be a solution to avoid cross-talk. We derived a model on the molecular level of their problem (see the Alternate Design section on the quorum sensing module page). We inserted the equations in our system, putting B (metabolite) instead of LasAHL as communicating molecule.
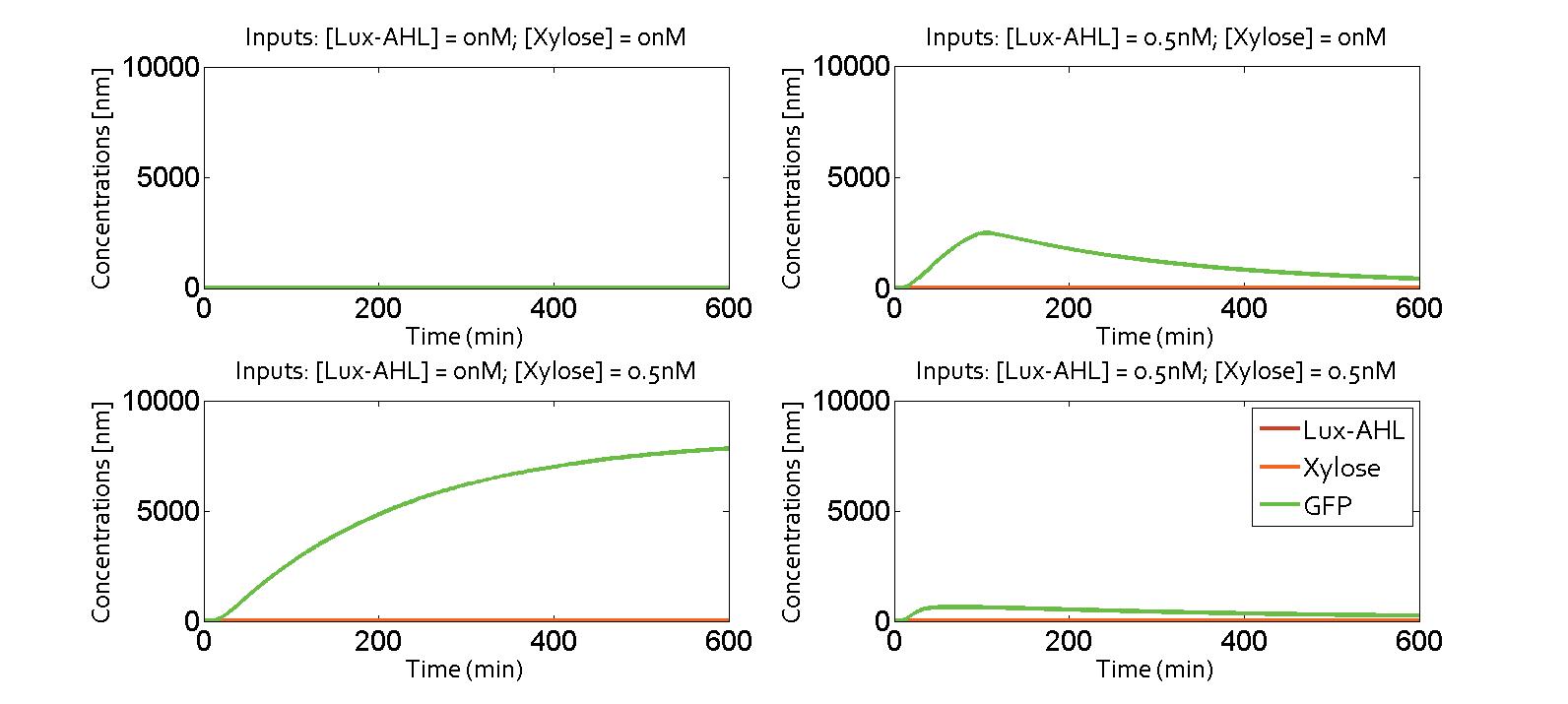
Here, we present the simulation results. The four possibilities of the XOR gate are presented in the following figure. The system was the one with a feedback corresponding to the metabolite pathway. The metabolite parameter taken was the one of xylose, which is supposed to be generic for all models
It exhibits an XOR behavior. It could be possible to combine quorum sensing and metabolite wiring to make the pattern appear.
 "
"