Team:Heidelberg/pages/Circularization
From 2014.igem.org
Contents |
Circularization
Introduction
Proteins normally appear in nature with two ends, the N-terminal amino and the C-terminal carboxylic groups. By linking those two ends, it is possible circularize proteins and introduce interesting properties. Circular proteins can be superior to linear proteins in terms of thermostability [1][2][3], resistance against chemical denaturation [4] and protection from exopeptidases [2][4], while the biological function of he linear counterpart is conserved [5]. Improving thermostability of enzymes is an essential aspect for industrial applications [5]. Indeed, in many biotechnological applications, enzymes are required to work at temperatures that would normally cause denaturation [6]. Other recombinant proteins, for example enzymes used in laboratories, are required to be as stable as possible at moderate temperatures [6]. Moreover, a circular backbone can improve in vivo stability of therapeutical proteins and peptides [5]. Our circularization tools allow easy expression of circular proteins in E. coli. Choose a protein you want to circularize and use our Toolbox Guide!
Circular Proteins in Nature
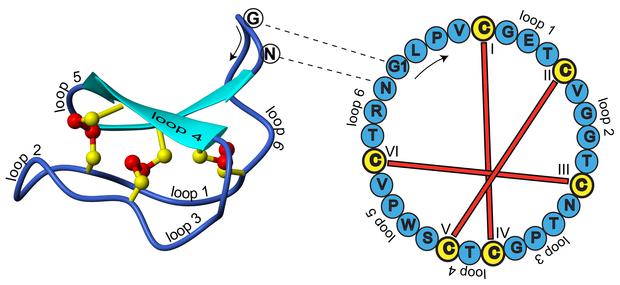
A different type of circular peptides, the cyclotides, were discovered more recently [8]. Those circular peptides are true proteins, since their linear precursors are produced by ribosomes [7]. The peptide ligase catalyzing the circularization of cyclotides, butelase 1, was discovered a few months ago [X]. Cyclotides are rather small (<79 aa) and most often them are additionally stabilized by disulfide bonds [7]. The first cyclotide to be discovered, katala B1, has been used by African women to accelerate childbirth and was obtained by boiling the dry plant leaves, demonstrating its considerable thermostability [8]. The unique cyclotide framework (Fig 1) is used in the development of drugs against HIV and cancer [Y].
Even in primates circular proteins have been discovered. Rhesus θ-defensin-1, found in rhesus macaque leukocytes, is a highly potent antibiotic peptide [9]. In the human genome there is a similar gene that is disrupted by a stop codon. The peptide called retrocyclin was synthesized chemically and it was shown to protect human lymphocytes from HIV-1 infection in vitro [Z].
Importantly, naturally occurring circularization of more complex and larger proteins have not yet been described. Considering the benefit that one can expect from improved protein stability, we anticipated that developing tools to circularize proteins would be of great help for the scientific and industrial communities.
Synthetically Circularized Proteins
Using [split inteins] or [sortase], it is possible to cirularize linear proteins. The most important proteins that were circularized using split inteins are dihydrofolate reductase [1] and GFP [4]. Those proteins are rather small and their termini are in close proximity. Both circular dihydrofolate reductase and GFP showed enhanced resistance against proteases and denaturation. The largest protein circular protein yet is maltose-binding protein (MBP) (395 aa) from E. coli [10]. However, stability was not tested in this case. A GFP variant was also circularized in vitro using sortase. In this case, faster recovery of fluorescence after denaturation was observed in comparison with a linear control [11].
Our Circularization Project
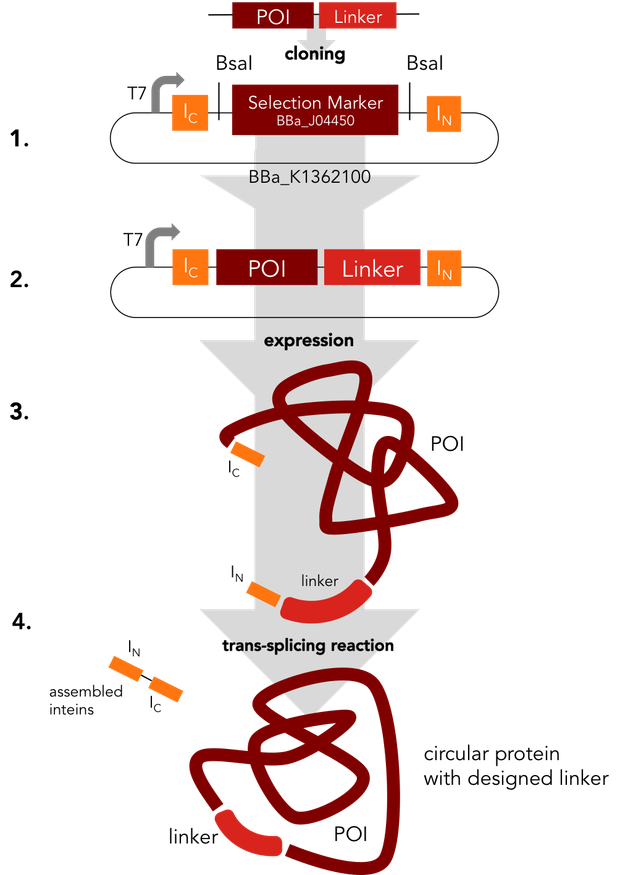
In a single step the protein of interest is cloned into the split intein circularization construct (1.). The resulting plasmid (2.) can be used to express a fusion protein (3.). In an autocatalytic in vivo reaction, the circular protein (4.) is formed.
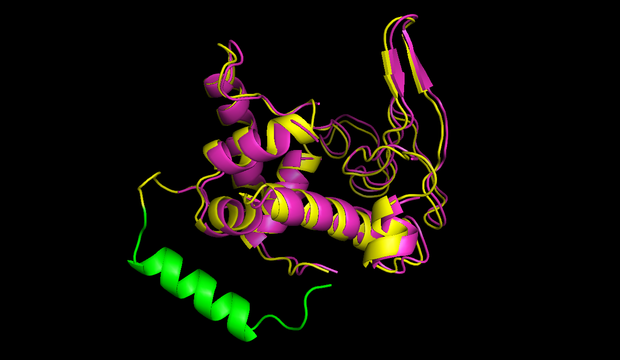
We decided to standardize the cloning process that is necessary to express circular proteins. Several circularization constructs based on the split intein Npu DnaE and Sortase A were designed. By Golden Gate cloning an mRFP selection marker can easily be exchanged with the protein to be circularized. Using our [http://parts.igem.org/Part:BBa_K1362000 split intein circularization construct], a fusion protein of the C-intein, the protein to be circularized and the N-intein is created. Upon expression, the C- and the N-intein assemble and catalyze their own excision and the formation of a peptide bond circularizing the protein. Our Sortase A based construct adds TEV and Sortase A recognition sites to the protein to be circularized. The expressed protein has to be purified and treated with TEV protease and Sortase A to be circularized in vitro. We used GFP to test both split intein and sortase circularization.
The Impact of Circularization on Thermostability
Backbone cyclization of proteins increases the thermostability of a protein in a similar way to the introduction of disulfide bridges. Either way, a covalent linkage is introduced that decreases the entropy of the unfolded state of the protein [6]. This principle has already been shown in 1956 in the context of fibrous proteins with covalent linkages [3]. A short explanation of "the entropy of the unfolded state is decreased": an unfolded linear polypeptide can move around, since each angle between the peptide bonds can adopt any possible state. If the polypeptide is circular, the state of those angles is much more restricted. Consequently, an unfolded state is less likely to occur, resulting in a higher thermostability.
Another factor that might improve thermostability is the fixation of the termini. They are usually flexible and thus a possible starting point of the denaturation process [6]. Especially if the termini are close together or a rigid linker is used to join them, this should cause an additional increase of thermostability.
What could be done?
- somehow depends on our final results as well -
References
[1] Scott, C. P., Abel-Santos, E., Wall, M., Wahnon, D. C. & Benkovic, S. J. Production of cyclic peptides and proteins in vivo. Proc. Natl. Acad. Sci. 96, 13638–13643 (1999).
[2] Iwai, H. & Plückthun, A. Circular beta-lactamase: stability enhancement by cyclizing the backbone. FEBS Lett. 459, 166–72 (1999).
[3] Flory, J. & Yol, S. Theory of Elastic Mechanisms in Fibrous Proteins. 715, 5222–5235 (1956)
[4] Iwai, H., Lingel, a & Pluckthun, a. Cyclic green fluorescent protein produced in vivo using an artificially split PI-PfuI intein from Pyrococcus furiosus. J. Biol. Chem. 276, 16548–54 (2001).
[5] Trabi, M. & Craik, D. J. Circular proteins--no end in sight. Trends Biochem. Sci. 27, 132–8 (2002).
[6] Vieille, C. & Zeikus, G. J. Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol. Mol. Biol. Rev. 65, 1–43 (2001).
[7] Craik, D. J. Chemistry. Seamless proteins tie up their loose ends. Science 311, 1563–4 (2006).
[8] A, P. K. B. et al. Elucidation of the Primary and Three-Dimensional Structure of the Uterotonic. 4147–4158 (1995).
[9] Tang, Y. A Cyclic Antimicrobial Peptide Produced in Primate Leukocytes by the Ligation of Two Truncated -Defensins. Science (80-. ). 286, 498–502 (1999).
[10] Evans, T. C. Protein trans-Splicing and Cyclization by a Naturally Split Intein from the dnaE Gene of Synechocystis Species PCC6803. J. Biol. Chem. 275, 9091–9094 (2000).
[11] Antos, J. M. et al. A straight path to circular proteins. J. Biol. Chem. 284, 16028–36 (2009).
[X] Nguyen, G. K. T. et al. Butelase 1 is an Asx-specific ligase enabling peptide macrocyclization and synthesis. Nat. Chem. Biol. 10, 732–8 (2014).
[Y] Henriques, S. T. & Craik, D. J. Cyclotides as templates in drug design. Drug Discov. Today 15, 57–64 (2010).
[Z] Cole, A. M. et al. Retrocyclin: a primate peptide that protects cells from infection by T- and M-tropic strains of HIV-1. Proc. Natl. Acad. Sci. U. S. A. 99, 1813–8 (2002).
[A] KatalaB1. Cyclotide structure. (2007). at <http://en.wikipedia.org/wiki/Cyclotide#mediaviewer/File:Cyclotide_structure.jpg>
 "
"
 [A]
[A]