Team:Aachen/Project/2D Biosensor
From 2014.igem.org
(→Agar Concentration) |
(→Achievements) |
||
| Line 186: | Line 186: | ||
<html></br></br></br></br></br></br></br></br></br></br></br></br></br></br></br></br></html> | <html></br></br></br></br></br></br></br></br></br></br></br></br></br></br></br></br></html> | ||
| - | === Detecting IPTG with | + | === Detecting IPTG with Sensor Chips === |
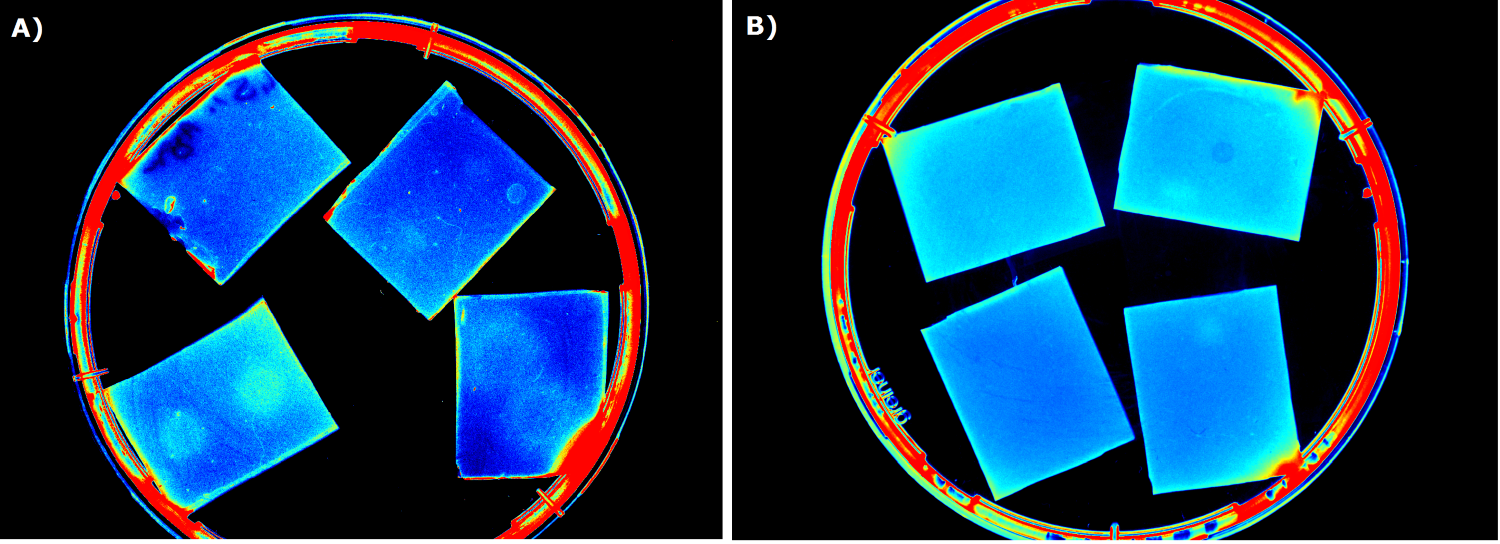
{{Team:Aachen/FigureFloat|Aachen_I746909_slower_reduced.gif|title=IPTG inducible superfolder GFP (I746909) in sensor chips|subtitle=IPTG inducible superfolder GFP (I746909) is induced with IPTG (2 µl, 100mM) on the right chip with a non induced chip on the left|width=480px}} | {{Team:Aachen/FigureFloat|Aachen_I746909_slower_reduced.gif|title=IPTG inducible superfolder GFP (I746909) in sensor chips|subtitle=IPTG inducible superfolder GFP (I746909) is induced with IPTG (2 µl, 100mM) on the right chip with a non induced chip on the left|width=480px}} | ||
This video shows the construct [http://parts.igem.org/Part:BBa_I746909 I746909] from the 2007 iGEM Team Cambridge. It is a producer of superfolder GFP under the control of a T7 promoter. This construct was introduced into BL21(DE3) cells making the expression IPTG inducible due to the T7 RNA Polymerase encoded in the genome of BL21(DE3) under the control of a lacI promoter. | This video shows the construct [http://parts.igem.org/Part:BBa_I746909 I746909] from the 2007 iGEM Team Cambridge. It is a producer of superfolder GFP under the control of a T7 promoter. This construct was introduced into BL21(DE3) cells making the expression IPTG inducible due to the T7 RNA Polymerase encoded in the genome of BL21(DE3) under the control of a lacI promoter. | ||
| Line 194: | Line 194: | ||
</html> | </html> | ||
| - | ===Detecting the 3-oxo-C{{sub|12}} HSL with K131026 in our | + | ===Detecting the 3-oxo-C{{sub|12}} HSL with K131026 in our Sensor Chips with WatsOn=== |
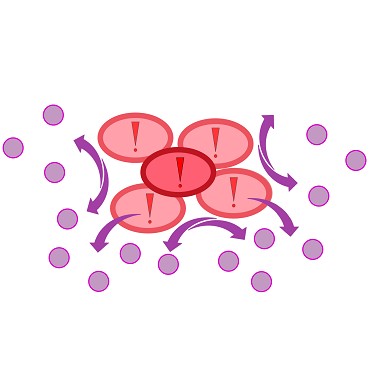
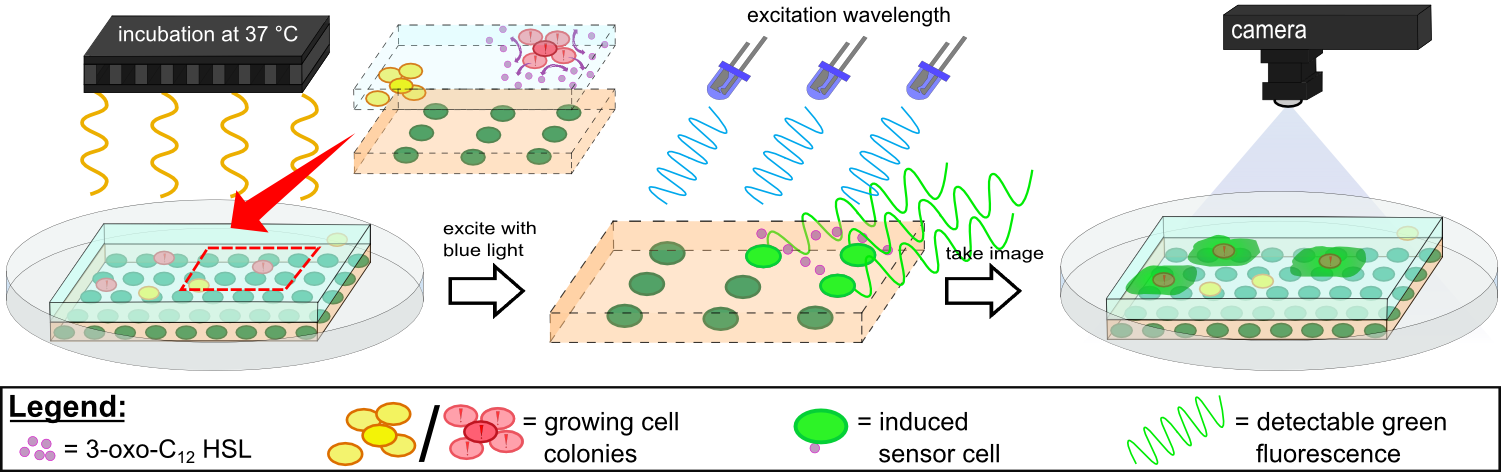
{{Team:Aachen/FigureFloatRight|Aachen_K131026_HSLdetection_slow.gif|title=Detection of 3-oxo-C{{sub|12}} HSL with K131026|subtitle=0.2 µL of 3-oxo-C{{sub|12}} HSL was placed in the middle of the chip and then incubated at 37 °C in WatsOn.|width=480px}} | {{Team:Aachen/FigureFloatRight|Aachen_K131026_HSLdetection_slow.gif|title=Detection of 3-oxo-C{{sub|12}} HSL with K131026|subtitle=0.2 µL of 3-oxo-C{{sub|12}} HSL was placed in the middle of the chip and then incubated at 37 °C in WatsOn.|width=480px}} | ||
| Line 205: | Line 205: | ||
</html> | </html> | ||
| - | ===Detecting Pseudomonas aeruginosa with K131026 in our | + | ===Detecting ''Pseudomonas aeruginosa'' with K131026 in our Sensor Chip with WatsOn=== |
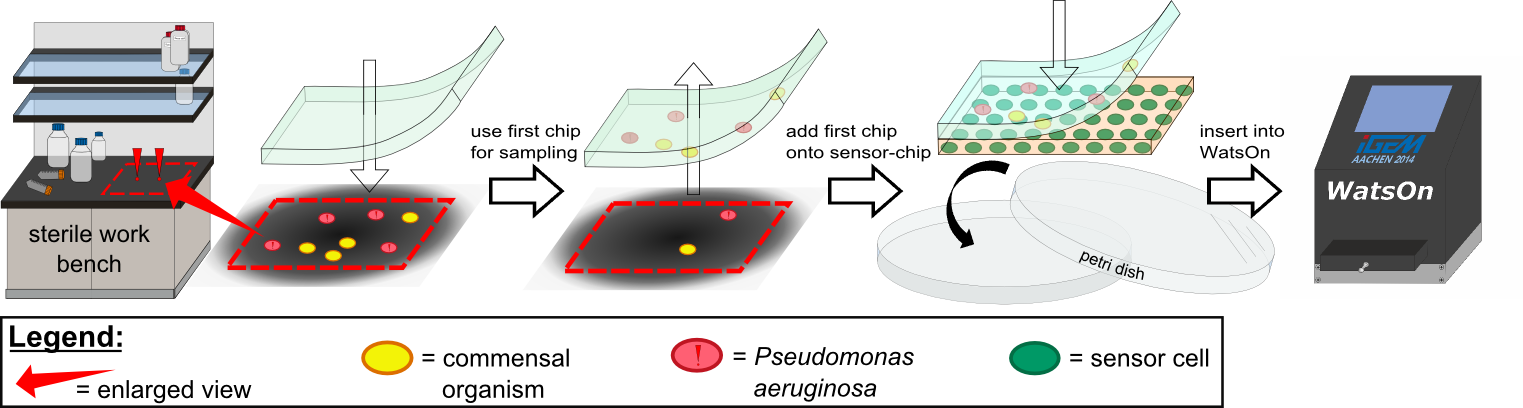
{{Team:Aachen/FigureFloat|Aachen_K131026_Pseudomonas_aeruginosa_detection.gif|title=Detection of ''Pseudomonas aeruginosa'' with K131026|subtitle=Direct detection of ''Pseudomonas aeruginosa'' on our sensor chips. Sensor cell used were K131026.|width=480px}} | {{Team:Aachen/FigureFloat|Aachen_K131026_Pseudomonas_aeruginosa_detection.gif|title=Detection of ''Pseudomonas aeruginosa'' with K131026|subtitle=Direct detection of ''Pseudomonas aeruginosa'' on our sensor chips. Sensor cell used were K131026.|width=480px}} | ||
| Line 215: | Line 215: | ||
</html> | </html> | ||
| - | === Comparing the REACh | + | === Comparing the REACh Construct with K731520 and I746909 === |
More information about the kinetic differences between these construct in our sensor chips, look under | More information about the kinetic differences between these construct in our sensor chips, look under | ||
Revision as of 10:32, 17 October 2014
|
|
|
|
|
|
 "
"