Team:Aachen/Project/2D Biosensor
From 2014.igem.org
(→Achievements) |
(→Principle of Operation) |
||
| Line 83: | Line 83: | ||
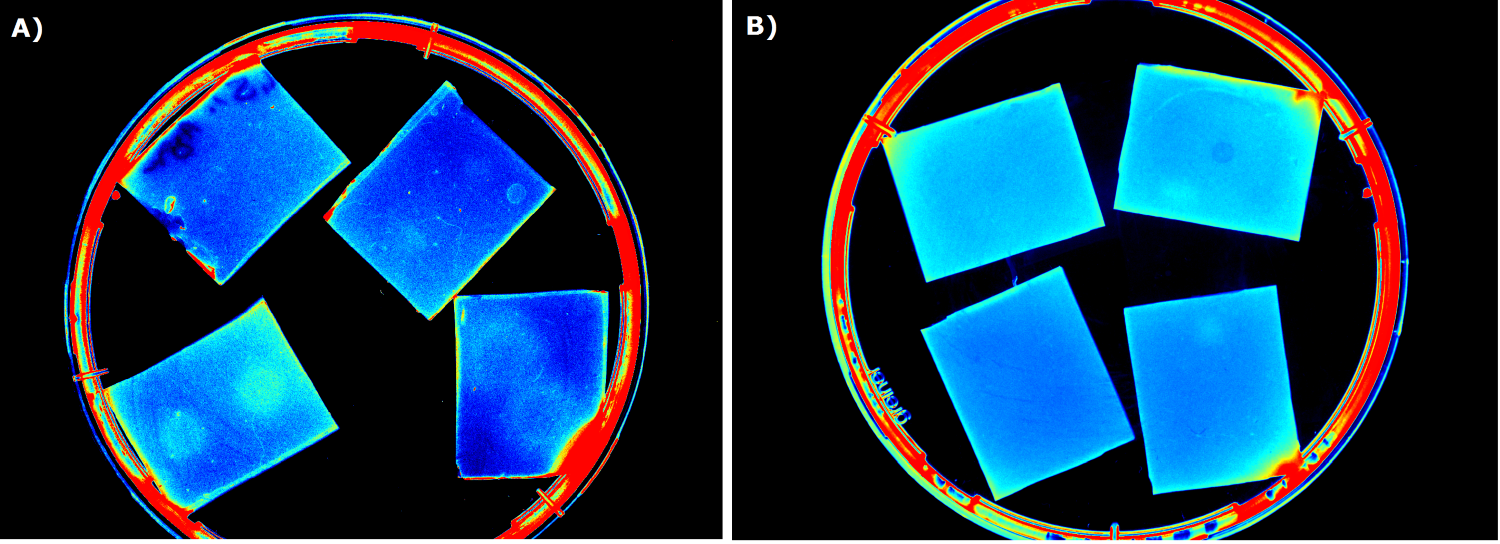
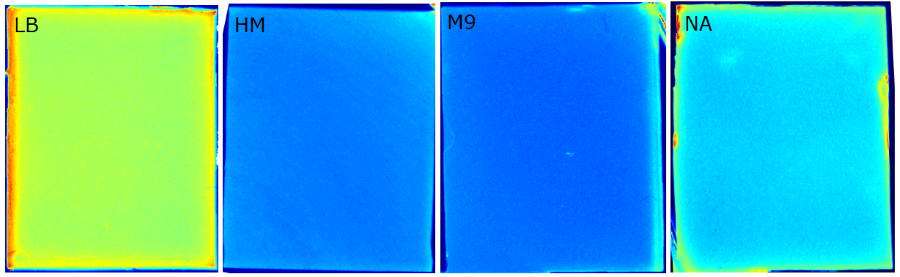
Our '''sensor cells, ''Cellocks'', are immobilized in agar chips'''. To make the chips, we mix the ''Cellocks'' with liquid LB agar. | Our '''sensor cells, ''Cellocks'', are immobilized in agar chips'''. To make the chips, we mix the ''Cellocks'' with liquid LB agar. | ||
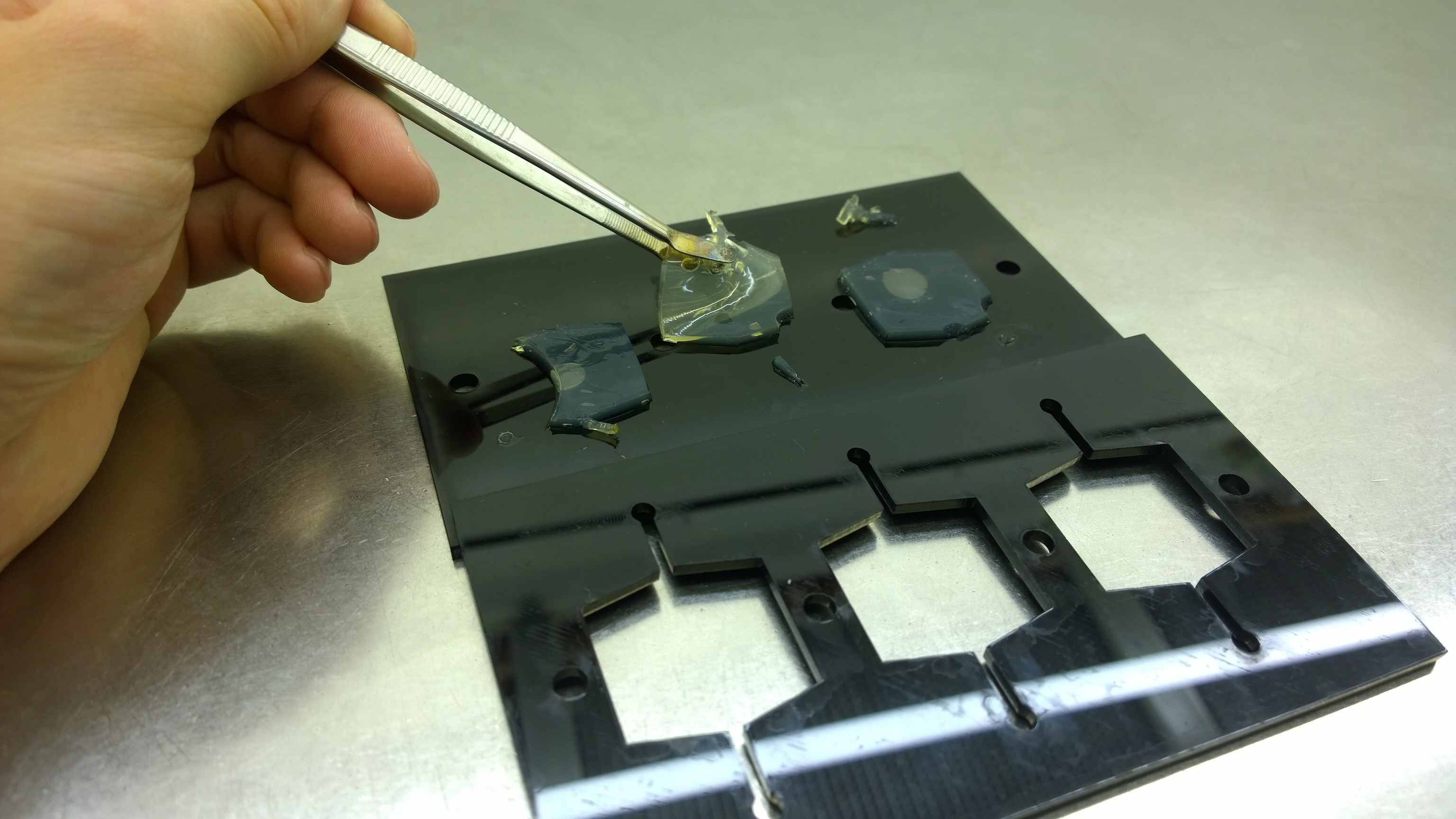
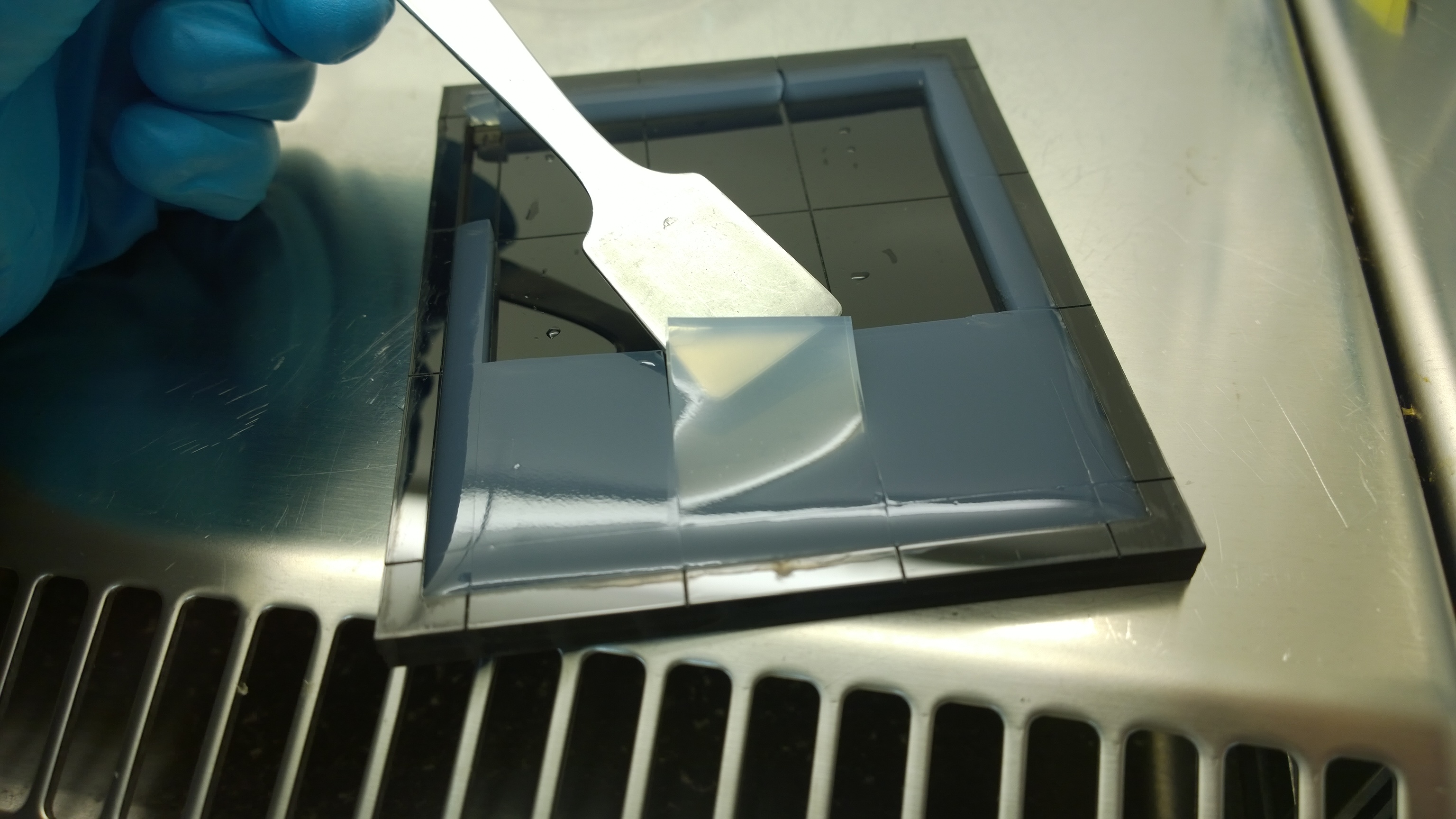
| - | In the course of our project, we designed a casting mold specifically for the production of our agar chips. When the agar has cooled down, the chips are cut out of the mold and are ready to use. Storage of the readily usable sensor chips is possible for up to | + | In the course of our project, we designed a casting mold specifically for the production of our agar chips. When the agar has cooled down, the chips are cut out of the mold and are ready to use. Storage of the readily usable sensor chips is possible for up to two days at 4°C when using LB medium or up to 5 days if TB medium is used. A detailed description of the sensor chip manufacturing can be found in our [https://2014.igem.org/Team:Aachen/Notebook/Protocols/detection Protocols] section. |
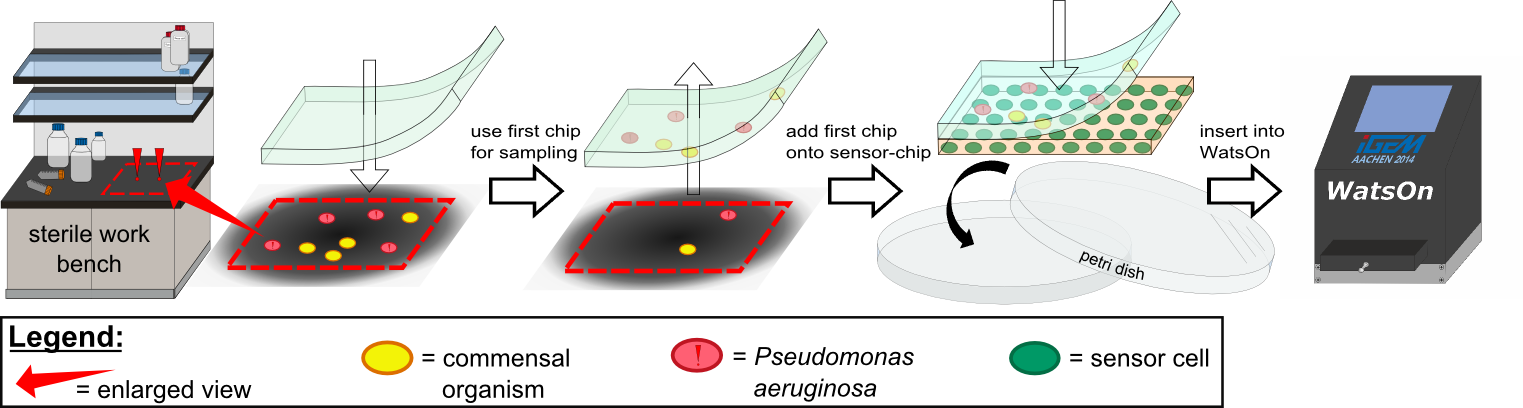
{{Team:Aachen/Figure|Aachen 14-10-14 Flowsheet OD-device part1 ipo.png|title=Assay to detect ''P. aeruginosa'' using ''Cellock Holmes''|subtitle=This flow sheet shows the procedure to sample and detect ''P. aeruginosa'': A sampling chip is briefly put onto the potentially contaminated surface, added onto one of our sensor chips and inserted into ''WatsOn''.|width=900px}} | {{Team:Aachen/Figure|Aachen 14-10-14 Flowsheet OD-device part1 ipo.png|title=Assay to detect ''P. aeruginosa'' using ''Cellock Holmes''|subtitle=This flow sheet shows the procedure to sample and detect ''P. aeruginosa'': A sampling chip is briefly put onto the potentially contaminated surface, added onto one of our sensor chips and inserted into ''WatsOn''.|width=900px}} | ||
| - | Using ''Cellock Holmes'', we developed a simple assay to detect ''P. aeruginosa''. | + | Using ''Cellock Holmes'', we developed a simple assay to detect ''P. aeruginosa''. Initially, a so called sampling chip is placed on a solid surface that is potentially contaminated with the pathogen. Subsequently, the sampling chip is removed from the surface and put onto one of our sensor chips. Theorectically, the sensor chips could be directly used for sampling, however, this was avoided in our project to '''match [https://2014.igem.org/Team:Aachen/Safety biosafety regulations]''' and to prevent the spread of genetically modified organisms (GMOs) into the environment. The two layered chip-stack is then put into a petri dish which is inserted into our measurement device [https://2014.igem.org/Team:Aachen/Project/Measurement_Device ''WatsOn''] for evalutation. |
| - | + | ||
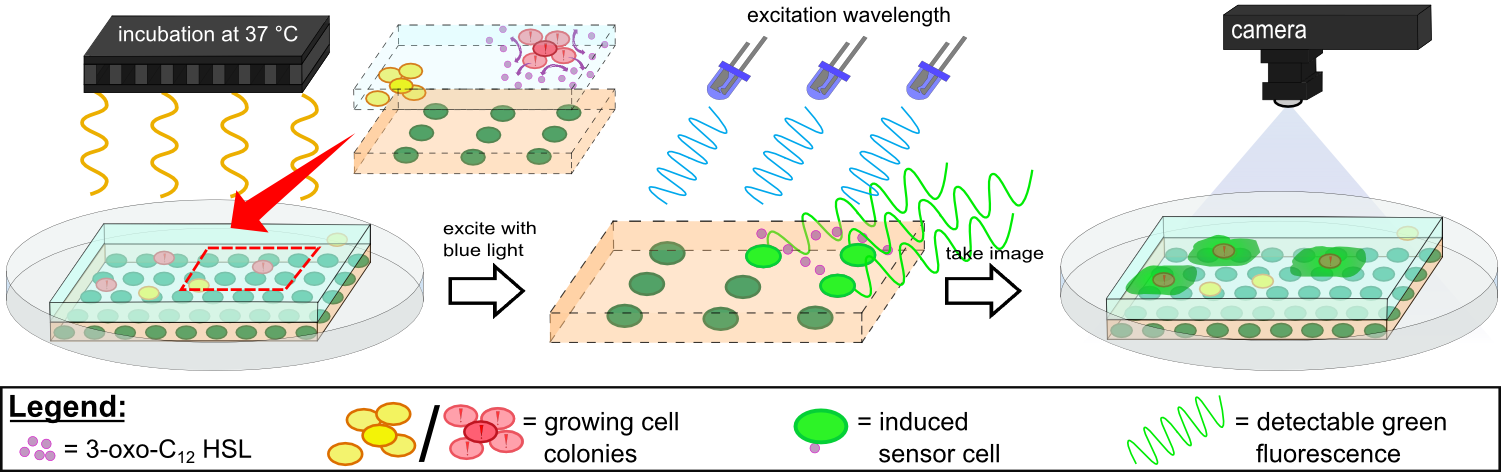
{{Team:Aachen/Figure|Aachen 14-10-14 Flowsheet OD-device part2 ipo.png|title=Mode of action inside ''WatsOn''|subtitle=Chips are incubated at 37°C to stimulate cell growth and then illuminated with blue light to excite fluorescence. A picture is taken and analyzed for fluorescence signals using the software ''Measurarty''.|width=900px}} | {{Team:Aachen/Figure|Aachen 14-10-14 Flowsheet OD-device part2 ipo.png|title=Mode of action inside ''WatsOn''|subtitle=Chips are incubated at 37°C to stimulate cell growth and then illuminated with blue light to excite fluorescence. A picture is taken and analyzed for fluorescence signals using the software ''Measurarty''.|width=900px}} | ||
Revision as of 01:34, 18 October 2014
|
|
|
|
|
|
 "
"