|
|
| Line 32: |
Line 32: |
| | The ''upp'' gene from Bacillus subtilis W168 encodes for a Uracilphosphoribosyl transferase (UPRTase). | | The ''upp'' gene from Bacillus subtilis W168 encodes for a Uracilphosphoribosyl transferase (UPRTase). |
| | Its key reaction in uracil salvage is the reaction of a uracil molecule with a 5'-phosphoribosyl-a-1- pyrophosphate (PRPP) molecule, resulting in the formation of UMP. | | Its key reaction in uracil salvage is the reaction of a uracil molecule with a 5'-phosphoribosyl-a-1- pyrophosphate (PRPP) molecule, resulting in the formation of UMP. |
| - | A second locus, the pyrR Gene, encoding a second UPRTase has been identified. However it has been shown, that the UPRTase derived from pyrR locus has an influence on overall UPRTase activity of < 1 %. (J Martinussen, P Glaser, P S Andersen and H H Saxild J. Bacteriol. 1995, 177(1):271. ) | + | A second locus, the ''pyrR'' gene, encoding a second UPRTase has been identified. However it has been shown, that the UPRTase derived from ''pyrR'' locus has an influence on overall UPRTase activity of < 1 %. (J Martinussen, P Glaser, P S Andersen and H H Saxild J. Bacteriol. 1995, 177(1):271.) |
| | This makes the ''upp''-derived UPRTase the only physiologically relevant catalyst for UPRTase activity. | | This makes the ''upp''-derived UPRTase the only physiologically relevant catalyst for UPRTase activity. |
| - | Exposure to the pyrimidine analogue 5-Fluorouracil UPRTase results in production of 5-fluoro-dUMP, a very potent inhibitor of the thymidylate synthase (Neuhard, J. (1983) Utilization of preformed pyrimidine bases and nucleosides. In Metabolism of Nucleotides, Nucleosides and Nucleobases in Microorganisms. Munch- Petersen, A. (eds). New York: Academic Press, pp. 95– 148. ). As a result 5-FU is toxic to the Bacillus subtilis W168 strain. | + | Exposure to the pyrimidine analogue 5-Fluorouracil UPRTase results in production of 5-fluoro-dUMP, a very potent inhibitor of the thymidylate synthase (Neuhard, J. (1983) Utilization of preformed pyrimidine bases and nucleosides. In Metabolism of Nucleotides, Nucleosides and Nucleobases in Microorganisms. Munch- Petersen, A. (eds). New York: Academic Press, pp. 95– 148. ). As a result 5-FU is toxic to the ''Bacillus subtilis'' W168 strain. |
| - | B. subtilis 5FU-resistant (5FUR) mutants selected on low drug concentration (10 mM 5FU) are UPRTase-defective. (Nygaard, P. (1993) Purine and pyrimidine salvage pathways. In Bacillus Subtilis and Other Gram-Positive Bacteria: Biochemistry, Physiology, and Molecular Genetics. Sonenshein, A.L., Hoch, J.A. and Losick, R., (eds). Washington, DC: American Society for Microbiology, pp. 359–378. ) | + | ''B. subtilis'' 5FU-resistant (5FUR) mutants selected on low drug concentration (10 mM 5FU) are UPRTase-defective. (Nygaard, P. (1993) Purine and pyrimidine salvage pathways. In Bacillus Subtilis and Other Gram-Positive Bacteria: Biochemistry, Physiology, and Molecular Genetics. Sonenshein, A.L., Hoch, J.A. and Losick, R., (eds). Washington, DC: American Society for Microbiology, pp. 359–378. ) |
| - | This has made ''upp'' a go to choice for negative selection in combination with an B. subtilis W168 Δ''upp'' strain. So far it has been used to make clean in-frame deletions and point mutations. (A new mutation delivery system for genome-scale approaches in Bacillus subtilis Céline Fabret,† S. Dusko Ehrlich and Philippe Noirot* Génétique Microbienne, INRA, Domaine de Vilvert, 78352 Jouy en Josas Cedex, France. ) | + | This has made ''upp'' a go to choice for negative selection in combination with an ''B. subtilis'' W168 ''Δupp'' strain. So far it has been used to make clean in-frame deletions and point mutations. (A new mutation delivery system for genome-scale approaches in Bacillus subtilis Céline Fabret,† S. Dusko Ehrlich and Philippe Noirot* Génétique Microbienne, INRA, Domaine de Vilvert, 78352 Jouy en Josas Cedex, France. ) |
| | | | |
| | However, to our knowledge, no application using ''upp'' for clean insertions has been established so far. | | However, to our knowledge, no application using ''upp'' for clean insertions has been established so far. |
| | | | |
| - | The I-SceI Restricition endonuclease has a highly specific recognition sequence of 18 nucleotides. No such sequence is present in the W168 strain. It creates a double strand break at targeted location, which leads to an increased rate of repair at the specific site. By this, the rate of homologous recombination is increased by an factor of 100x. (Establishment of a Markerless Mutation Delivery System in Bacillus subtilis Stimulated by a Double-Strand Break in the Chromosome Ting Shi1,2,3,4., Guanglu Wang1,2,3,4., Zhiwen Wang1,2,3,4*, Jing Fu1,2,3,4, Tao Chen1,2,3,4, Xueming Zhao1,2,3,4 ) | + | The I-SceI Restricition endonuclease has a highly specific recognition sequence of 18 nucleotides. No such sequence is present in the ''B.subtilis'' W168 strain. It creates a double strand break at targeted location, which leads to an increased rate of repair at the specific site. By this, the rate of homologous recombination is increased by a factor of 100. (Establishment of a Markerless Mutation Delivery System in Bacillus subtilis Stimulated by a Double-Strand Break in the Chromosome Ting Shi1,2,3,4., Guanglu Wang1,2,3,4., Zhiwen Wang1,2,3,4*, Jing Fu1,2,3,4, Tao Chen1,2,3,4, Xueming Zhao1,2,3,4 ) |
| | <html> | | <html> |
| | </article> | | </article> |
| Line 49: |
Line 49: |
| | <article class="ac-small"> | | <article class="ac-small"> |
| | </html> | | </html> |
| - | The Plan was to restructure the BioBrick compatible, integrative ''Bacillus subtilis'' vectors [http://parts.igem.org/Part:BBa_J179000 pSB1C], [http://parts.igem.org/Part:BBa_J179001 pSB2E] and [http://parts.igem.org/Part:BBa_J179002 pSB4S] in a fashion that leads to deletion of the antibiotic resistance after insertion of a gene of interest has been made. | + | The Plan was to restructure the BioBrick compatible, integrative ''Bacillus subtilis'' vectors [http://parts.igem.org/Part:BBa_J179000 pSB1C], [http://parts.igem.org/Part:BBa_J179001 pSB2E] and [http://parts.igem.org/Part:BBa_J179002 pSB4S] in a fashion that leads to deletion of the antibiotic resistance after an insertion of a gene of interest has been generated. |
| - | Integration of those basic vectors is achieved via homologous recombination between the ''amyE''/''lacA''/''thrC'' locus and the corresponding up and down fragments on the vector. This leads to an insertion of the gene of interest within the RCF10 compatible multiple cloning site and the resistance for positive selection (CAM/MLS/Spec). | + | Integration of those basic vectors is achieved via homologous recombination between the ''amyE''/''lacA''/''thrC'' locus, respectively, and the corresponding up and down fragments on the vector. This leads to an insertion of the gene of interest within the RCF10 compatible multiple cloning site and the resistance for positive selection (Cm<sup>r</sup>/MLS<sup>r</sup>/Spec<sup>r</sup>). |
| | | | |
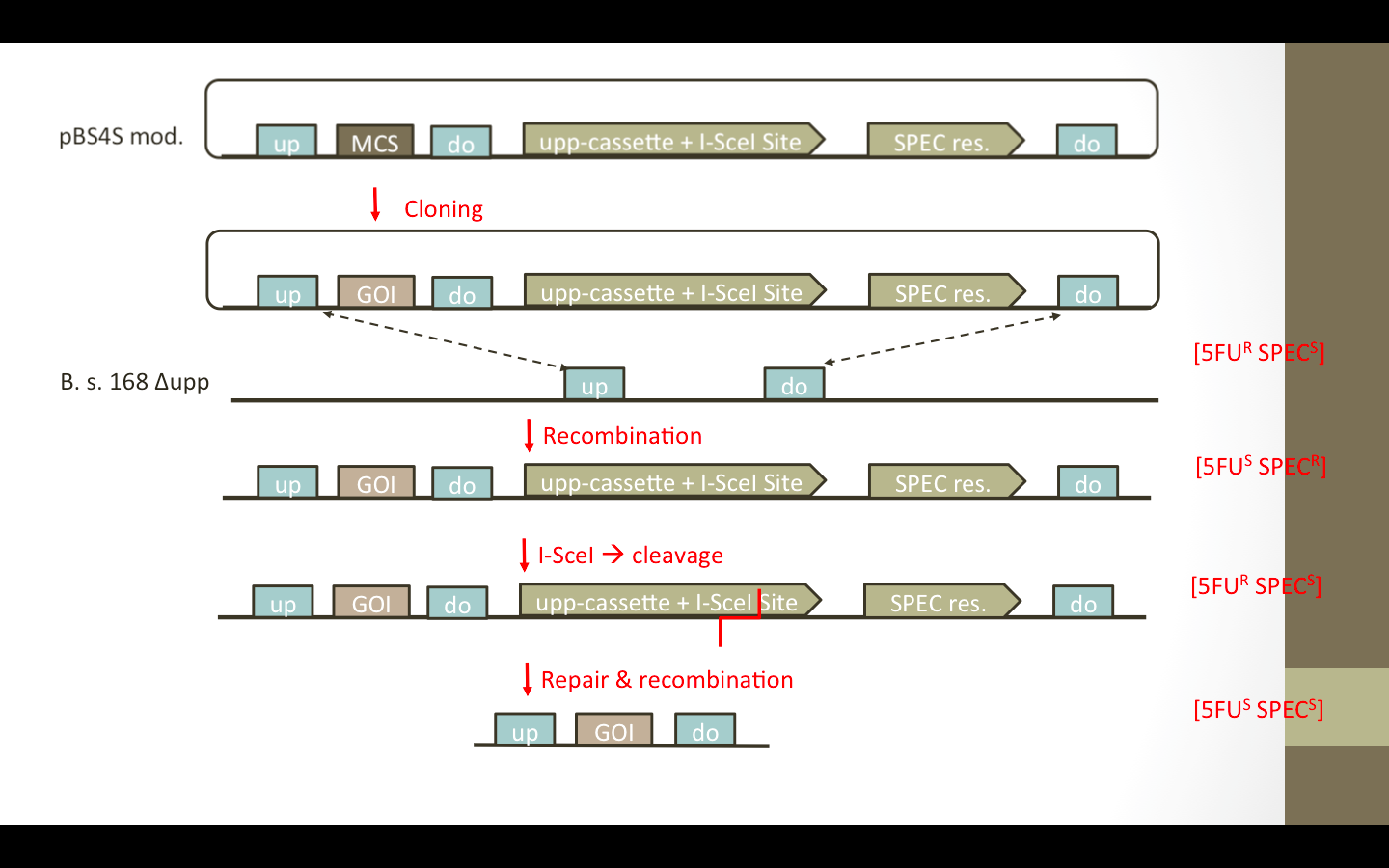
| - | We wanted to add the ''upp''-cassette, containing an I-SceIsite, as well as an additional up/down fragment to the vector. The desired vectors are presented in Fig. 1. The ''upp''-cassette will allow negative selection in 5FU media. The contained I-SceI site will be cut by the I-SceI restriction endonuclease which is encoded on helper plasmid pEBS-cop1. (nochmal auf das paper verweisen) (Figure) | + | We wanted to add the ''upp''-cassette, containing an I-SceI site, as well as an additional up/down fragment to the vector. The desired vectors are presented in Fig. 1. The ''upp''-cassette will allow negative selection in 5FU media. The contained I-SceI site will be cut by the I-SceI restriction endonuclease which is encoded on a helper plasmid pEBS-cop1. (nochmal auf das paper verweisen) (Figure) |
| | | | |
| | Cloning Strategy | | Cloning Strategy |
 "
"