Team:Heidelberg
From 2014.igem.org
(Difference between revisions)
| Line 154: | Line 154: | ||
</div> | </div> | ||
<div class="arrow arrow-left"></div> | <div class="arrow arrow-left"></div> | ||
| + | </div> | ||
| + | </div> | ||
| + | <div class="row"> | ||
| + | <div class="jumbotron slide grey"> | ||
| + | <div> | ||
| + | <div> | ||
| + | <div class="col-md-3 col-md-offset-0 col-xs-6 col-xs-offset-3" style="z-index:5;"> | ||
| + | <img class="img-responsive" src="/wiki/images/e/e5/Heidelberg_Frontpage_igemathome.png" /> | ||
| + | </div> | ||
| + | <div class="col-md-9 col-xs-12"> | ||
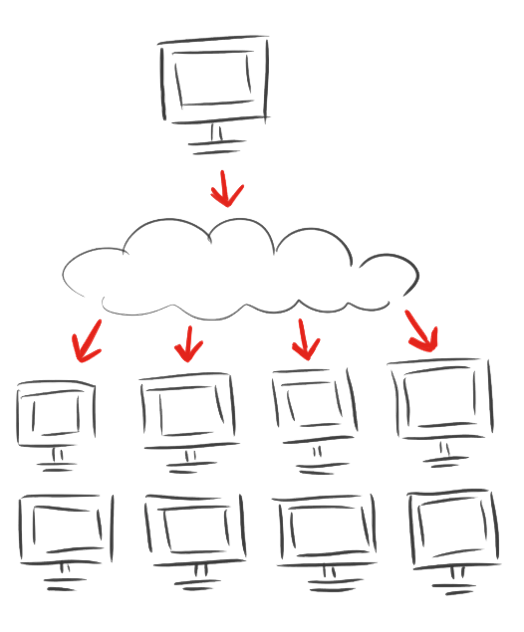
| + | <p class="align-right"><span class="larger">As calculations on protein structures are costly<br />we involved the rest of the world with</span></p> | ||
| + | <h1 class="very-large-text red-text"><span style="font-size:1.5em;">iGEM@home</span></h1> | ||
| + | </div> | ||
| + | <div class="clearfix"></div> | ||
| + | </div> | ||
| + | <div class="col-md-8 col-xs-12"> | ||
| + | <h3 class="normal-medium-text">Empowering science!</h3> | ||
| + | <p>NEW PLATFORM for distributed computing.</p> | ||
| + | <p>Volunteers provide the idle calacity of their home computers.</p> | ||
| + | <p>And, with an established user base with more than <span class="larger red-text">1000</span> computers, we effectively bring synthetic biology to society in a new way</p> | ||
| + | </div> | ||
| + | <div class="col-md-4 col-md-offset-0 col-sm-offset-3 col-sm-6 col-xs-offset-2 col-xs-8"> | ||
| + | <img style="position:relative; top: -35px;" class="img-responsive" src="/wiki/images/5/52/Heidelberg_Frontpage_igemathome_cloud.png" /> | ||
| + | </div> | ||
| + | <div class="clearfix"></div> | ||
| + | </div> | ||
| + | <div class="arrow arrow-left"></div> | ||
| + | </div> | ||
| + | </div> | ||
| + | <div style="position:relative;" class="row"> | ||
| + | <div class="jumbotron slide grey"> | ||
| + | <div> | ||
| + | <div class="col-md-4 col-md-push-8 col-md-offset-0 col-xs-offset-3 col-xs-6" style="z-index:5;"> | ||
| + | <img class="img-responsive" src="/wiki/images/a/a6/Heidelberg_Project_Dnmt1.png" /> | ||
| + | </div> | ||
| + | <div class="col-md-8 col-md-pull-4 col-xs-12"> | ||
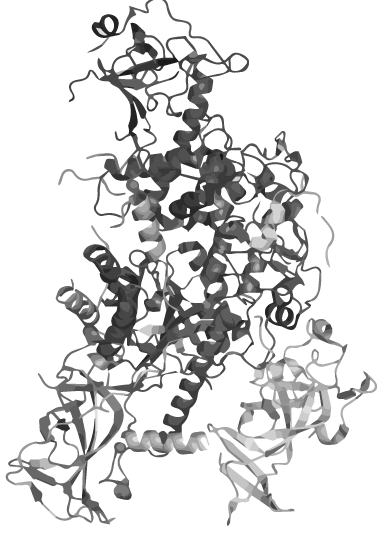
| + | <h1 class="dark-grey-text" style="text-align: right;"><span style="font-size: 0.8em;">circular <span class="red-text">heat-stable</span></span><br>DNMT1</h1> | ||
| + | <p>Wouldn´t it be great to amplify DNA in a normal PCR maintaining the epigenetic information coded in methylation patterns?</p> | ||
| + | <p>The problem: DNMT 1, an enzyme which is responsible for the establishment and maintenance of the individual methylation pattern of different cell types, is not heat stable. | ||
| + | For iGEM 2014 we therefore create a PCR 2.0 with heat-stable DNMT 1 by circularization. | ||
| + | </p> | ||
| + | </div> | ||
| + | <div class="clearfix"></div> | ||
| + | </div> | ||
| + | <div> | ||
| + | <div class="title-wrapper-dnmt1"> | ||
| + | <span class="title-dnmt1">PCR 2.0</span> | ||
| + | <span class="special-span-dnmt1"></span> | ||
| + | </div> | ||
| + | </div> | ||
| + | <div class="arrow arrow-right"></div> | ||
</div> | </div> | ||
</div> | </div> | ||
| Line 195: | Line 246: | ||
<div class="clearfix"></div> | <div class="clearfix"></div> | ||
<div class="arrow arrow-left"></div> | <div class="arrow arrow-left"></div> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
</div> | </div> | ||
</div> | </div> | ||
| Line 238: | Line 265: | ||
<span class="title-dnmt1">INDUSTRY</span> | <span class="title-dnmt1">INDUSTRY</span> | ||
<span class="special-span-dnmt1"></span> | <span class="special-span-dnmt1"></span> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
</div> | </div> | ||
<div class="arrow arrow-left"></div> | <div class="arrow arrow-left"></div> | ||
Revision as of 14:38, 15 October 2014
 "
"



 CIRCULARIZATION
CIRCULARIZATION
 OLIGOMERIZATION
OLIGOMERIZATION
 FUSION
FUSION
 ON/OFF
ON/OFF
 PURIFICATION
PURIFICATION

