Team:Heidelberg
From 2014.igem.org
(Difference between revisions)
| Line 24: | Line 24: | ||
<h3>iGEM TEAM <span>HEIDELBERG</span> 2014</h3> | <h3>iGEM TEAM <span>HEIDELBERG</span> 2014</h3> | ||
<h1>THE RING<br />OF <span>FIRE</span></h1> | <h1>THE RING<br />OF <span>FIRE</span></h1> | ||
| - | <p | + | <p>Welcome to the proteins of tomorrow.<br/>This is the iGEM team Heidelberg‘s wiki page.</p> |
<p> Click <a href="/Team:Heidelberg/Project#Abstract">here</a> to view our abstract. | <p> Click <a href="/Team:Heidelberg/Project#Abstract">here</a> to view our abstract. | ||
<br/> | <br/> | ||
| Line 40: | Line 40: | ||
<div class="col-lg-9 col-md-9 col-sm-12" style="text-align: right;"> | <div class="col-lg-9 col-md-9 col-sm-12" style="text-align: right;"> | ||
<h1 style="text-align:right;"><span class="red-text" style="font-size:1.3em;">PLACEHOLDER</span></h1> | <h1 style="text-align:right;"><span class="red-text" style="font-size:1.3em;">PLACEHOLDER</span></h1> | ||
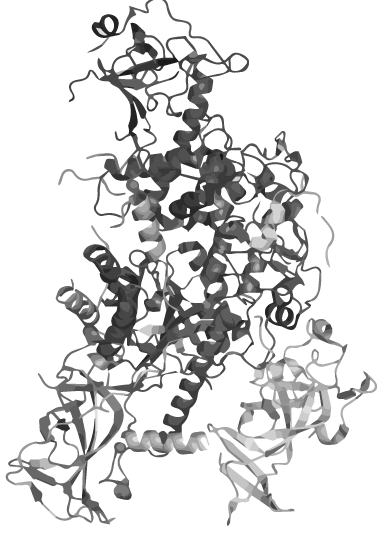
| - | <p><span class="red-text" style="font-size: 1.5em;">Nature</span> has made many curious inventions. One of these are circular proteins, which are conventional peptides that neither have a beginning, nor an ending | + | <p><span class="red-text" style="font-size: 1.5em;">Nature</span> has made many curious inventions. One of these are circular proteins, which are conventional peptides that neither have a beginning, nor an ending</p> |
<p>Apart from other features, these proteins are extremely stable against | <p>Apart from other features, these proteins are extremely stable against | ||
high temperatures, pH and proteases.</p> | high temperatures, pH and proteases.</p> | ||
| - | <p style="font-weight: | + | <p style="font-weight:bold;">We seeked to apply this principle of circularization in Synthetic Biology and create a way of rendering any protein heatstable.</p> |
</div> | </div> | ||
| Line 67: | Line 67: | ||
</div> | </div> | ||
</div> | </div> | ||
| - | |||
| - | |||
| - | |||
<div class="row"> | <div class="row"> | ||
<div class="jumbotron slide epic-background" style="color:white;"> | <div class="jumbotron slide epic-background" style="color:white;"> | ||
| Line 112: | Line 109: | ||
</div> | </div> | ||
<div style="position:relative;" class="row"> | <div style="position:relative;" class="row"> | ||
| - | <div class="jumbotron slide | + | <div class="jumbotron slide grey"> |
<div> | <div> | ||
<div class="col-md-4 col-md-push-8 col-md-offset-0 col-xs-offset-3 col-xs-6" style="z-index:5;"> | <div class="col-md-4 col-md-push-8 col-md-offset-0 col-xs-offset-3 col-xs-6" style="z-index:5;"> | ||
| Line 118: | Line 115: | ||
</div> | </div> | ||
<div class="col-md-8 col-md-pull-4 col-xs-12"> | <div class="col-md-8 col-md-pull-4 col-xs-12"> | ||
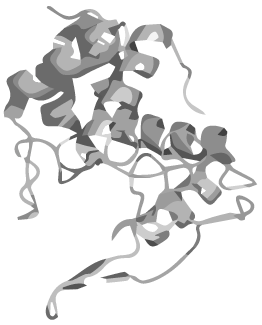
| - | <h1 style=" | + | <h1 class="dark-grey-text" style="text-align: right;"><span style="font-size: 0.8em;">circular <span class="red-text">heat-stable</span></span><br>DNMT1</h1> |
<p>Wouldn´t it be great to amplify DNA in a normal PCR maintaining the epigenetic information coded in methylation patterns?</p> | <p>Wouldn´t it be great to amplify DNA in a normal PCR maintaining the epigenetic information coded in methylation patterns?</p> | ||
<p>The problem: DNMT 1, an enzyme which is responsible for the establishment and maintenance of the individual methylation pattern of different cell types, is not heat stable. | <p>The problem: DNMT 1, an enzyme which is responsible for the establishment and maintenance of the individual methylation pattern of different cell types, is not heat stable. | ||
Revision as of 15:08, 13 October 2014
 "
"


 CIRCULARIZATION
CIRCULARIZATION
 OLIGOMERIZATION
OLIGOMERIZATION
 FUSION
FUSION
 ON/OFF
ON/OFF
 PURIFICATION
PURIFICATION