|
|
| (21 intermediate revisions not shown) |
| Line 23: |
Line 23: |
| | </html> | | </html> |
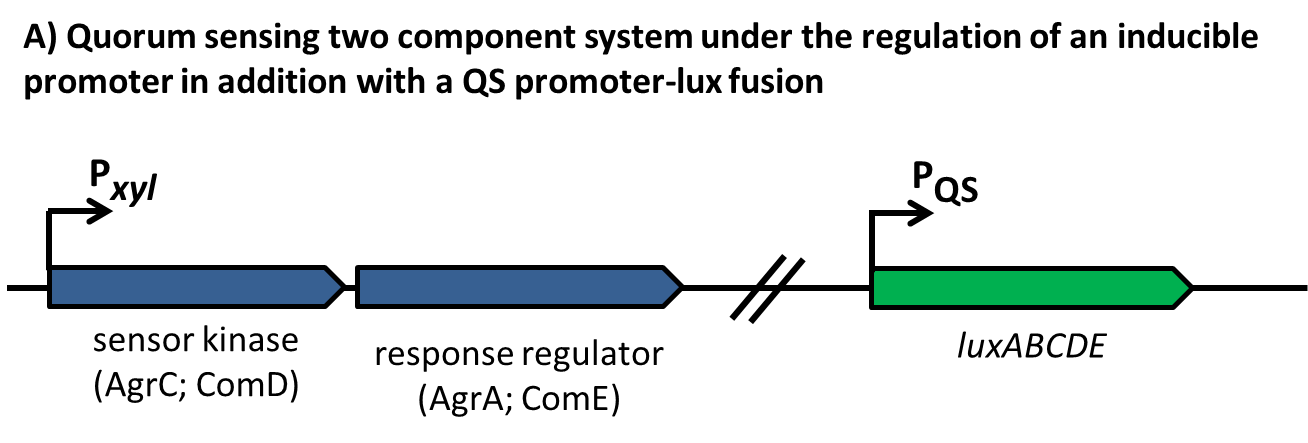
| | == Sensing == | | == Sensing == |
| - | This part of the BaKillus-concept focuses on specific pathogen detection. Here we use quorum sensing to specifically recognize the presence of certain pathogens. The major aim of this subproject is transferring the quorum sensing two-component systems AgrC-II/AgrA of ''Staphylococcus aureus'' and ComDE of ''Streptococcus pneumoniae'' into ''Bacillus subtilis'', thus creating a pathogen detecting strain. With this strategy we will trigger our killing strategies only if a threshold concentration of autoinducers, produced by pathogens, is present in the environment. Thereby we aim to drastically increase the specificity of our antimicrobial strategies compared to commonly used broad-spectrum antibiotic therapy. | + | This part of the BaKillus-concept focuses on specific pathogen detection. Here we use quorum sensing to specifically recognize the presence of certain pathogens. The major aim of this subproject is transferring the quorum sensing two-component systems AgrC-II/AgrA of ''Staphylococcus aureus'' and ComDE of ''Streptococcus pneumoniae'' into ''Bacillus subtilis'', thus creating a pathogen detecting strain. With this strategy we will trigger our killing strategies only if a threshold concentration of autoinducers, produced by pathogens, is present in the environment Thereby we aim to drastically increase the specificity of our antimicrobial strategies compared to commonly used broad-spectrum antibiotic therapy. |
| | | | |
| | <html> | | <html> |
| Line 648: |
Line 648: |
| | | | |
| | Subtilin is a small antimicrobial peptide (AMP) which belongs to the class of lanthionine-containing antibiotics (lantibiotics). These are heat-stable, ribosomally synthesized molecules with a molecular weight below 4 kDa. Their main characteristic is the high proportion of unusual amino acids, synthesized by post-translational side-chain modifications of precursor peptides. Most prominent examples are the polycyclic thioether amino acids, lanthionine and β–methyllanthionine and a number of dehydrated amino acids such as dehydroalanine (Dha) and dehydrobutyrine (Dhb). | | Subtilin is a small antimicrobial peptide (AMP) which belongs to the class of lanthionine-containing antibiotics (lantibiotics). These are heat-stable, ribosomally synthesized molecules with a molecular weight below 4 kDa. Their main characteristic is the high proportion of unusual amino acids, synthesized by post-translational side-chain modifications of precursor peptides. Most prominent examples are the polycyclic thioether amino acids, lanthionine and β–methyllanthionine and a number of dehydrated amino acids such as dehydroalanine (Dha) and dehydrobutyrine (Dhb). |
| - | Lantibiotics are promising candidates for future antimicrobials, as they inhibit the growth of many clinically relevant pathogens comprising even multidrug-resistant bacteria. Moreover, lantibiotics have only low tendency to generate resistance. Both features make them highly attractive for medical applications. [1][2] | + | Lantibiotics are promising candidates for future antimicrobials, as they inhibit the growth of many clinically relevant pathogens comprising even multidrug-resistant bacteria. Moreover, lantibiotics have only low tendency to generate resistance. Both features make them highly attractive for medical applications. [http://www.unboundmedicine.com/medline/citation/24210177/Lantibiotics:_promising_candidates_for_future_applications_in_health_care_[1]] [http://www.ncbi.nlm.nih.gov/pubmed/16205711[2]] |
| | | | |
| | [[File:LMU14 killing subtilin structure.png|thumb|800px|center|Fig. 1. Schematic representation of the structure of the lantibiotic subtilin. Dha represents a dehydroalanine residue, Dhb represents a dehydrobutirine residue, and Abu represents a dehydrobutirine residue that has formed a thioether β-methyl-lanthionine bridge with a cysteine residue. | | [[File:LMU14 killing subtilin structure.png|thumb|800px|center|Fig. 1. Schematic representation of the structure of the lantibiotic subtilin. Dha represents a dehydroalanine residue, Dhb represents a dehydrobutirine residue, and Abu represents a dehydrobutirine residue that has formed a thioether β-methyl-lanthionine bridge with a cysteine residue. |
| Line 704: |
Line 704: |
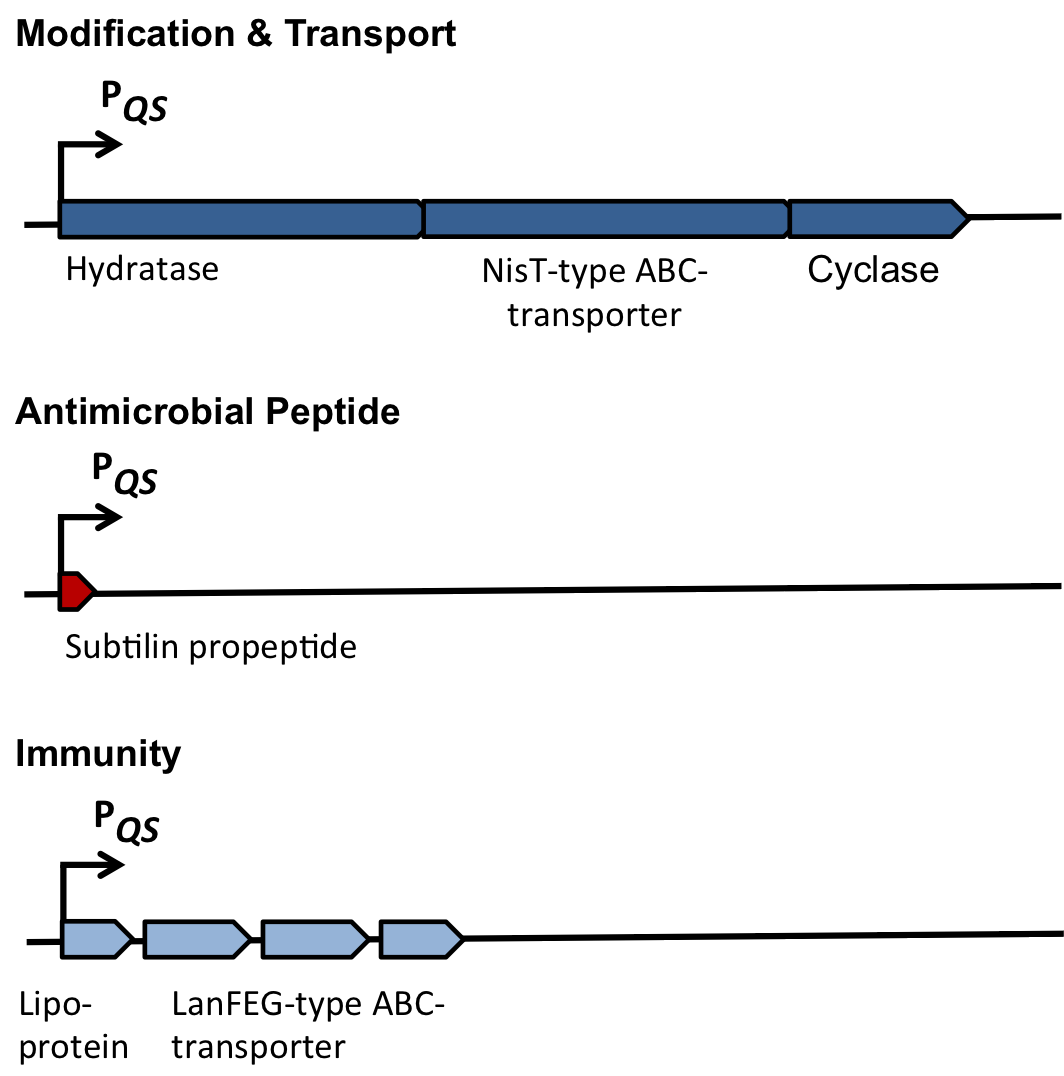
| | ===Self-protection from subtilin=== | | ===Self-protection from subtilin=== |
| | | | |
| - | Of course, the producer needs to be immune against the antimicrobial agent it produces. ''B. subtilis'' ATCC6633 protects itself against subtilin by two mechanisms that both act independently and confer some level of resistance, however full resistance is only achieved when both mechanisms are active. SpaI is a membrane-anchored lipoprotein. It is suggested that it binds the antimicrobial peptide and by this keeps it away from the membrane. SpaFEG is a LanFEG-like ABC-transporter that works by exporting subtilin from the cytoplasmic membrane to the culture supernatant. | + | Of course, the producer needs to be immune against the antimicrobial agent it produces. ''B. subtilis'' ATCC6633 protects itself against subtilin by two mechanisms that both act independently and confer some level of resistance, however full resistance is only achieved when both mechanisms are active. SpaI is a membrane-anchored lipoprotein. It is suggested that it binds the antimicrobial peptide and by this keeps it away from the membrane. SpaFEG is a LanFEG-like ABC-transporter that works by exporting subtilin from the cytoplasmic membrane to the culture supernatant. [http://www.ncbi.nlm.nih.gov/pubmed/15659659 [4]] [http://www.ncbi.nlm.nih.gov/pubmed/?term=Alkhatib%2C+Z.%3B+Abts%2C+A.%3B+Mavaro%2C+A.%3B+Schmitt%2C+L.%3B+Smits%2C+S.+H.%3A+Lantibiotics%3A+how+do+producers+become+self-protected%3F+Journal+of+biotechnology+2012%2C+159%2C+145-54. [5]] |
| | | | |
| | | | |
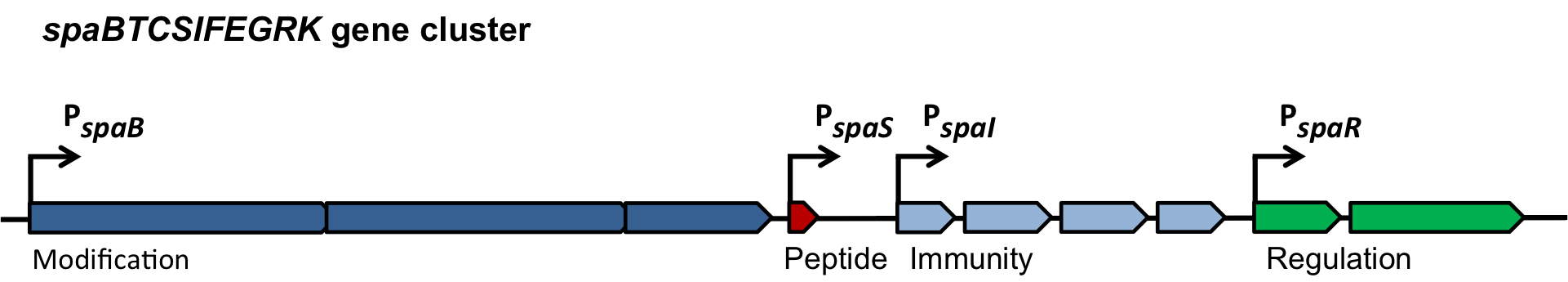
| | ===Dual regulation of subtilin biosynthesis and immunity=== | | ===Dual regulation of subtilin biosynthesis and immunity=== |
| | | | |
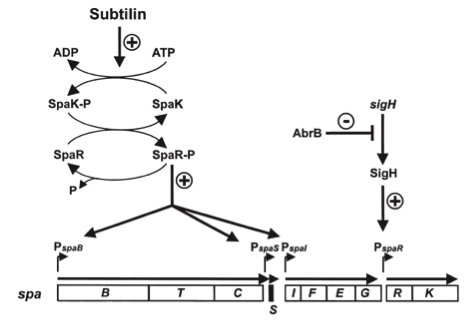
| - | [[File:LMU14 killing subtilin regulation.png|thumb|400px|right|Fig. 3. Dual control of subtilin biosynthesis and immunity. [http://www.ncbi.nlm.nih.gov/pubmed/11972779]]] | + | [[File:LMU14 killing subtilin regulation.png|thumb|400px|right|Fig. 3. Dual control of subtilin biosynthesis and immunity. [http://www.ncbi.nlm.nih.gov/pubmed/11972779 [6] ]]] |
| | | | |
| | Subtilin biosynthesis and immunity in ''B. subtilis'' ATCC6633 are subject to a dual control mechanism. | | Subtilin biosynthesis and immunity in ''B. subtilis'' ATCC6633 are subject to a dual control mechanism. |
| Line 716: |
Line 716: |
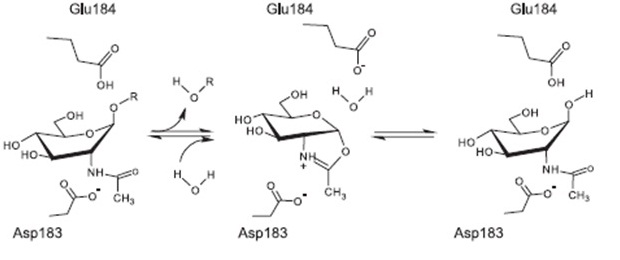
| | ===Mode of action=== | | ===Mode of action=== |
| | | | |
| - | Lantibiotics are active against a wide range of Gram-positive bacteria including multidrug-resistant pathogens. Some lantibiotics have a dual mode of action, subtilin is proposed to be among them. Subtilin forms a complex with the cell wall precursor lipid II, whereby the pyrophosphate moiety may play a key role for target recognition. Thereby, the cell wall biosynthesis is inhibited. Furthermore, the complexes aggregate and form a pore in the bacterial membrane by using lipid II as a docking molecule. This leads to a depolarisation of the membrane potential, the efflux of cytoplasma and in turn to rapid cell death. Just like nisin, which is highly similar, subtilin is still active in the nanomolar range and thus serves excellently as alternative killing strategy against multidrug-resistant pathogens. | + | Lantibiotics are active against a wide range of Gram-positive bacteria including multidrug-resistant pathogens. Some lantibiotics have a dual mode of action, subtilin is proposed to be among them. Subtilin forms a complex with the cell wall precursor lipid II, whereby the pyrophosphate moiety may play a key role for target recognition. Thereby, the cell wall biosynthesis is inhibited. Furthermore, the complexes aggregate and form a pore in the bacterial membrane by using lipid II as a docking molecule. This leads to a depolarisation of the membrane potential, the efflux of cytoplasma and in turn to rapid cell death. Just like nisin, which is highly similar, subtilin is still active in the nanomolar range and thus serves excellently as alternative killing strategy against multidrug-resistant pathogens. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Molecular+Mechanism+of+Target+parisot [7]] [http://www.researchgate.net/publication/7875075_Mode_of_action_of_lipid_II-targeting_lantibiotics [8]] |
| | | | |
| - | [[File:LMU14 killing subtilin modeofaction.png|frame|150px|center|Fig. 4. Proposed mechanism of pyrophosphate-mediated target engagement by the lanthionine antibiotic subtilin. Membrane breaches occur in a lipid II-mediated fashion and cell wall synthesis is impeded through engagement of pyrophosphate-containing intermediates into complexes. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Molecular+Mechanism+of+Target+parisot [A7]]]] | + | |
| | + | [[File:LMU14 killing subtilin modeofaction.png|frame|150px|center|Fig. 4. Proposed mechanism of pyrophosphate-mediated target engagement by the lanthionine antibiotic subtilin. Membrane breaches occur in a lipid II-mediated fashion and cell wall synthesis is impeded through engagement of pyrophosphate-containing intermediates into complexes. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Molecular+Mechanism+of+Target+parisot [7] ]]] |
| | | | |
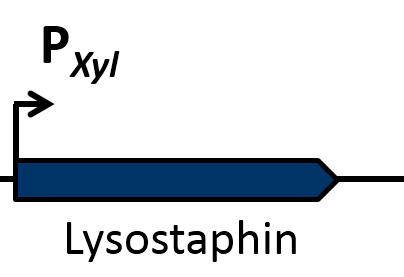
| | == Lysostaphin == | | == Lysostaphin == |
| Line 758: |
Line 759: |
| | [4] Stein, T.; Heinzmann, S.; Dusterhus, S.; Borchert, S.; Entian, K. D.: Expression and functional analysis of the subtilin immunity genes spaIFEG in the subtilin-sensitive host Bacillus subtilis MO1099. Journal of bacteriology 2005, 187, 822-8. | | [4] Stein, T.; Heinzmann, S.; Dusterhus, S.; Borchert, S.; Entian, K. D.: Expression and functional analysis of the subtilin immunity genes spaIFEG in the subtilin-sensitive host Bacillus subtilis MO1099. Journal of bacteriology 2005, 187, 822-8. |
| | | | |
| - | [5 ] Alkhatib, Z.; Abts, A.; Mavaro, A.; Schmitt, L.; Smits, S. H.: Lantibiotics: how do producers become self-protected? Journal of biotechnology 2012, 159, 145-54. | + | [5] Alkhatib, Z.; Abts, A.; Mavaro, A.; Schmitt, L.; Smits, S. H.: Lantibiotics: how do producers become self-protected? Journal of biotechnology 2012, 159, 145-54. |
| | | | |
| | [6] Stein, T.; Borchert, S.; Kiesau, P.; Heinzmann, S.; Kloss, S.; Klein, C.; Helfrich, M.; Entian, K. D.: Dual control of subtilin biosynthesis and immunity in Bacillus subtilis. Molecular microbiology 2002, 44, 403-16. | | [6] Stein, T.; Borchert, S.; Kiesau, P.; Heinzmann, S.; Kloss, S.; Klein, C.; Helfrich, M.; Entian, K. D.: Dual control of subtilin biosynthesis and immunity in Bacillus subtilis. Molecular microbiology 2002, 44, 403-16. |
| Line 852: |
Line 853: |
| | | | |
| | First of all, a growth curve for ''S. pneumoniae'' was recorded to learn about the growth behavior of this organism. It was measured in the plate reader in 100 µl THB broth at 37 °C without shaking and measurement was taken every 10 min. The mean value of 10 measurements were determined and plotted. | | First of all, a growth curve for ''S. pneumoniae'' was recorded to learn about the growth behavior of this organism. It was measured in the plate reader in 100 µl THB broth at 37 °C without shaking and measurement was taken every 10 min. The mean value of 10 measurements were determined and plotted. |
| - | [[File:LMU14 killing subtilin growthcurve pneumo.png|thumb|500px|right|Fig. 1. Growth curve of ''S. pneumoniae'' measured in the plate reader in 100 µl THB broth.]] | + | [[File:LMU14 killing subtilin growthcurve pneumo.png|thumb|600px|center|Fig. 1. Growth curve of ''S. pneumoniae'' measured in the plate reader in 100 µl THB broth.]] |
| - | [[File:LMU14 killing subtilin Pneumo-halo.png|thumb|700px|center|Fig. 2. The spot-on-lawn assay shows a clear halo around the ''B. subtilis'' ATCC6633 spot. No clearing zone is visible around ''B. subtilis'' W168.]] | + | [[File:LMU14 killing subtilin Pneumo-halo.png|thumb|400px|center|Fig. 2. The spot-on-lawn assay shows a clear halo around the ''B. subtilis'' ATCC6633 spot. No clearing zone is visible around ''B. subtilis'' W168.]] |
| | As expected, ''S. pneumoniae'' shows rapid growth during its exponential growth phase. It reaches its maximal OD<sub>600</sub> just under 5 hours. In contrast to other bacteria, for ''S. pneumoniae'', there is no plateau during stationary phase, as autolysis of the cells starts and the OD<sub>600</sub> decreases again. | | As expected, ''S. pneumoniae'' shows rapid growth during its exponential growth phase. It reaches its maximal OD<sub>600</sub> just under 5 hours. In contrast to other bacteria, for ''S. pneumoniae'', there is no plateau during stationary phase, as autolysis of the cells starts and the OD<sub>600</sub> decreases again. |
| | | | |
| Line 923: |
Line 924: |
| | Thanks to the numerous killing strategies listed above, BaKillus has specifically dispatched the targeted pathogens, thus its job is almost done. However, to ensure the temporary mode of action and to avoid uncontrollable spread, we implemented the so called “suicide switch“ leading to time-delayed cell death of BaKillus. | | Thanks to the numerous killing strategies listed above, BaKillus has specifically dispatched the targeted pathogens, thus its job is almost done. However, to ensure the temporary mode of action and to avoid uncontrollable spread, we implemented the so called “suicide switch“ leading to time-delayed cell death of BaKillus. |
| | | | |
| - | During stationary phase of a Bacillus subtilis culture it has been observed that some cells start producing a toxin which causes cell lysis of sister cells whereas the toxin-producer cells stay unaffected. [http://www.sciencemag.org/content/301/5632/510.full [1]] This social and heterogenous behavior is called cannibalism and seems to be an even more promising basis for constructing the BaKillus “suicide switch“ since it has been shown that the toxin also kills a variety of Gram-positive bacteria, including MRSA-strains. [http://www.pnas.org/content/107/37/16286.full [2]] | + | During stationary phase of a ''Bacillus subtilis'' culture it has been observed that some cells start producing a toxin which causes cell lysis of sister cells whereas the toxin-producer cells stay unaffected. [http://www.sciencemag.org/content/301/5632/510.full [1]] This social and heterogenous behavior is called cannibalism and seems to be an even more promising basis for constructing the BaKillus “suicide switch“ since it has been shown that the toxin also kills a variety of Gram-positive bacteria, including MRSA-strains. [http://www.pnas.org/content/107/37/16286.full [2]] |
| | | | |
| | So how does the suicide switch work? | | So how does the suicide switch work? |
| Line 941: |
Line 942: |
| | <article class="ac-small"> | | <article class="ac-small"> |
| | </html> | | </html> |
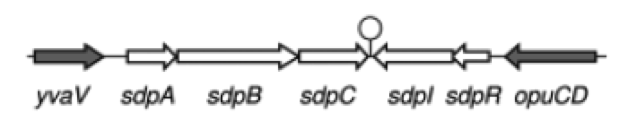
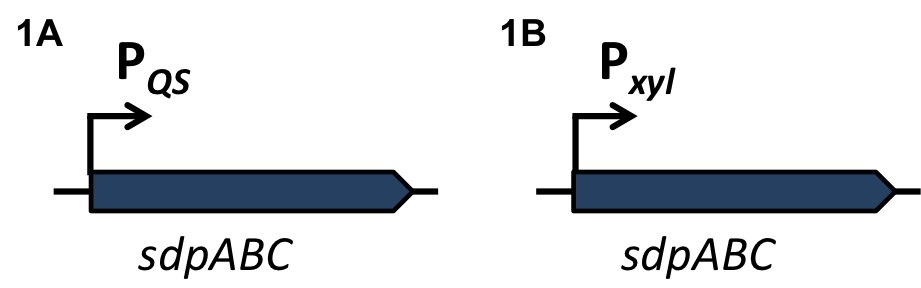
| - | '''The sdp-System of B. subtilis''' consists of two operons: The ''sdpABC'' operon, coding for the production and secretion of the cannibalism toxin SDP and the ''sdpRI'' operon responsible for the regulation and production of the immunity protein SdpI (Fig. 1). | + | '''The ''sdp''-System of ''B. subtilis''''' consists of two operons: The ''sdpABC'' operon, coding for the production and secretion of the cannibalism toxin SDP and the ''sdpRI'' operon responsible for the regulation and production of the immunity protein SdpI (Fig. 1). |
| | | | |
| - | [[File:LMU14 suicide background Fig.1.png|thumb|800px|center|Fig. 1. Gene organization for the sdpABC sdpRI operons. The hairpin symbolizes thetranscriptional terminators. [http://www.sciencemag.org/content/301/5632/510.full [1] ]]] | + | [[File:LMU14 suicide background Fig.1.png|thumb|800px|center|Fig. 1. Gene organization for the ''sdpABC'' ''sdpRI'' operons. The hairpin symbolizes thetranscriptional terminators. [http://www.sciencemag.org/content/301/5632/510.full [1] ]]] |
| | | | |
| | In vegetative cells, both operons are repressed by the unstable AbrB regulator. However, during early stages of sporulation AbrB itself is repressed by the master regulator of sporulation Spo0A, making ''sdpABC'' and ''sdpRI'' accessible for RNA polymerase. [http://www.sciencemag.org/content/301/5632/510.full [1]] | | In vegetative cells, both operons are repressed by the unstable AbrB regulator. However, during early stages of sporulation AbrB itself is repressed by the master regulator of sporulation Spo0A, making ''sdpABC'' and ''sdpRI'' accessible for RNA polymerase. [http://www.sciencemag.org/content/301/5632/510.full [1]] |
| Line 953: |
Line 954: |
| | [[File:Background Fig.2.png|thumb|800px|center|Fig. 2. SDP production requires multiple steps. In the cytosol, the full length SdpC (pro-SdpC1-203) is secreted via the Sec pathway. Following secretion, the signal peptidases SipS and SipT cleave the N-terminal signal peptide sequence of SdpC. Disulfide bond formation occurs independently of SdpAB. Finally, posttranslational cleavage of SdpC occurs via SdpAB to produce a 42-amino-acid SDP that will be secreted extracellulary as the active SDP peptide. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3697648/ [2] ]]] | | [[File:Background Fig.2.png|thumb|800px|center|Fig. 2. SDP production requires multiple steps. In the cytosol, the full length SdpC (pro-SdpC1-203) is secreted via the Sec pathway. Following secretion, the signal peptidases SipS and SipT cleave the N-terminal signal peptide sequence of SdpC. Disulfide bond formation occurs independently of SdpAB. Finally, posttranslational cleavage of SdpC occurs via SdpAB to produce a 42-amino-acid SDP that will be secreted extracellulary as the active SDP peptide. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3697648/ [2] ]]] |
| | | | |
| - | SDP has been shown to be a very effective AMP against a variety of Gram-positive bacteria in the Phylum of the Firmicutes (Fig. 3). It rapidly collapses the proton motive force (PMF), thus inducing autolysis. [3] | + | SDP has been shown to be a very effective AMP against a variety of Gram-positive bacteria in the Phylum of the Firmicutes (Fig. 3). It rapidly collapses the proton motive force (PMF), thus inducing autolysis. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3839633/ [3]] |
| | | | |
| - | [[File:Background Fig.3.png|thumb|600px|center|Fig. 3. SDP inhibition curves for pathogenic microbes. Relative growth of the strains named abovewith the presence of increasing concentrations of SDP is shown in the curve. As a negative control the gram-negative bacteria ''K. pneumoniae'' and ''P. aeruginosa'' are depicted, which are unaffected by the toxin SDP, as it specifically targets gram-positive bacteria. ''B. subtilis'' is a gram-positive bacteria, but expresses the immunity protein SdpI and is therefore relatively resistent to the toxin SDP. the ''Stapylococcus'' species (also MRSA) though are quite drastically reduced in the presence of the SDP. [4]]] | + | [[File:Background Fig.3.png|thumb|600px|center|Fig. 3. SDP inhibition curves for pathogenic microbes. Relative growth of the strains named abovewith the presence of increasing concentrations of SDP is shown in the curve. As a negative control the gram-negative bacteria ''K. pneumoniae'' and ''P. aeruginosa'' are depicted, which are unaffected by the toxin SDP, as it specifically targets gram-positive bacteria. ''B. subtilis'' is a gram-positive bacteria, but expresses the immunity protein SdpI and is therefore relatively resistent to the toxin SDP. the ''Stapylococcus'' species (also MRSA) though are quite drastically reduced in the presence of the SDP. [http://www.pnas.org/content/107/37/16286.full [4] ]]] |
| | | | |
| | === The ''sdpIR'' Operon – Production and Regulation of the Immunity Protein SdpI === | | === The ''sdpIR'' Operon – Production and Regulation of the Immunity Protein SdpI === |
| | | | |
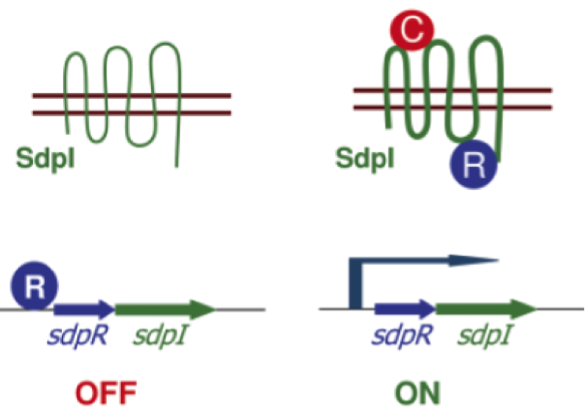
| - | In the absence of SDP the gene coding for the immunity protien DdpI is repressed by the autorepressor SdpR. If extracellular SDP binds the transmembrane protein SdpI it causes a conformational change. As a consequence SdpR is sequestered at the membrane, forming a complex with SDP and SdpI. Since the repression by SdpR is relieved the RNA polymerase can bind the ''sdpRI'' promotor region and start transcription (Fig. 4). [http://www.sciencemag.org/content/301/5632/510.full [1]] | + | In the absence of SDP the gene coding for the immunity protein SdpI is repressed by the autorepressor SdpR. If extracellular SDP binds the transmembrane protein SdpI it causes a conformational change. As a consequence SdpR is sequestered at the membrane, forming a complex with SDP and SdpI. Since the repression by SdpR is relieved the RNA polymerase can bind the ''sdpRI'' promotor region and start transcription (Fig. 4). [http://www.sciencemag.org/content/301/5632/510.full [1]] |
| | | | |
| | [[File:Background Fig.4.png|thumb|800px|center|Fig. 4. Model of how SDP induces expression of the ''sdpRI'' immunity operon. The signal SDP interacts with the immunity protein SdpI to induce a conformational change that allows SdpI to bind the repressor SdpR, which is sequestered at the membrane. [http://www.sciencemag.org/content/301/5632/510.full [1] ]]] | | [[File:Background Fig.4.png|thumb|800px|center|Fig. 4. Model of how SDP induces expression of the ''sdpRI'' immunity operon. The signal SDP interacts with the immunity protein SdpI to induce a conformational change that allows SdpI to bind the repressor SdpR, which is sequestered at the membrane. [http://www.sciencemag.org/content/301/5632/510.full [1] ]]] |
| Line 965: |
Line 966: |
| | === ECF41: An alternative sigma factor for building an orthogonal Delay Switch === | | === ECF41: An alternative sigma factor for building an orthogonal Delay Switch === |
| | | | |
| - | The alternative σ-factor ''ecf41''<sub>''bli aa 1-204''</sub>, which derives from ''B. licheniformis'' is a truncated version and binds to the target promoter P<sub>''ydfG''</sub>. It is an orthogonal σ-factor since there are no cross-reactions with other native promoters. [5] The introduction of this loop leads to a translational delay. | + | The alternative σ-factor ''ecf41''<sub>''bli aa 1-204''</sub>, which derives from ''B. licheniformis'' is a truncated version and binds to the target promoter P<sub>''ydfG''</sub>. It is an orthogonal σ-factor since there are no cross-reactions with other native promoters. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3426412/ [5]] The introduction of this loop leads to a translational delay. |
| | | | |
| | === FsrA: a small RNA Represses Expression of Succinate Dehydrogenase === | | === FsrA: a small RNA Represses Expression of Succinate Dehydrogenase === |
| | | | |
| - | Regulation of bacterial iron homeostasis is often controlled by the iron-sensing ferric uptake repressor (Fur). The ''Bacillus subtilis'' Fur protein acts as an iron-dependent repressor for siderophore biosynthesis and iron transport proteins. Furthermore it also coordinates an iron-sparing response that acts to repress the expression of iron-rich proteins (e.g. succinate dehydrogenase) when iron is limited. It has been shown that the downregulation of succinate dehydrogenase (SDH) is most likely caused by complementary binding of a small RNA, named fsrA, to the leader region of the sdhCAB mRNA, namely SdhC' (Fig. 5). In this project this principle is exploited to downregulate the translation of the immunity protein SdpI in a delayed response to the detection of the QS-molecules of ''S. aureus'' or ''S. pneumoniae''. [6] | + | Regulation of bacterial iron homeostasis is often controlled by the iron-sensing ferric uptake repressor (Fur). The ''Bacillus subtilis'' Fur protein acts as an iron-dependent repressor for siderophore biosynthesis and iron transport proteins. Furthermore it also coordinates an iron-sparing response that acts to repress the expression of iron-rich proteins (e.g. succinate dehydrogenase) when iron is limited. It has been shown that the downregulation of succinate dehydrogenase (SDH) is most likely caused by complementary binding of a small RNA, named fsrA, to the leader region of the sdhCAB mRNA, namely SdhC' (Fig. 5). In this project this principle is exploited to downregulate the translation of the immunity protein SdpI in a delayed response to the detection of the QS-molecules of ''S. aureus'' or ''S. pneumoniae''. [http://www.pnas.org/content/105/33/11927.full.pdf [6]] |
| | | | |
| - | [[File:Background Fig.5.png|thumb|800px|center|Fig. 5. Predicted pairing between fsrA and the 5’-leader region of sdhC, the first gene of the sdhCAB operon. [6]]] | + | [[File:Background Fig.5.png|thumb|800px|center|Fig. 5. Predicted pairing between fsrA and the 5’-leader region of sdhC, the first gene of the ''sdhCAB'' operon. [http://www.pnas.org/content/105/33/11927.full.pdf [6] ]]] |
| | | | |
| | === Sources === | | === Sources === |
| Line 981: |
Line 982: |
| | [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3839633/ [3] Lamsa, A., et al. (2012). "The Bacillus subtilis cannibalism toxin SDP collapses the proton motive force and induces autolysis." Mol Microbiol 84(3): 486-500. | | [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3839633/ [3] Lamsa, A., et al. (2012). "The Bacillus subtilis cannibalism toxin SDP collapses the proton motive force and induces autolysis." Mol Microbiol 84(3): 486-500. |
| | | | |
| - | [4] Liu, W. T., et al. (2010). "Imaging mass spectrometry of intraspecies metabolic exchange revealed the cannibalistic factors of Bacillus subtilis." Proc Natl Acad Sci U S A | + | [http://www.pnas.org/content/107/37/16286.full [4]] Liu, W. T., et al. (2010). "Imaging mass spectrometry of intraspecies metabolic exchange revealed the cannibalistic factors of Bacillus subtilis." Proc Natl Acad Sci U S A |
| | | | |
| - | [5] Wecke, T., et al. (2012). "Extracytoplasmic function sigma factors of the widely distributed group ECF41 contain a fused regulatory domain." Microbiologyopen 1(2): 194-213. | + | [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3426412/ [5]] Wecke, T., et al. (2012). "Extracytoplasmic function sigma factors of the widely distributed group ECF41 contain a fused regulatory domain." Microbiologyopen 1(2): 194-213. |
| | | | |
| - | [6] Gaballa, A., et al. (2008). "The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins." Proc Natl Acad Sci U S A 105(33): 11927-11932. | + | [http://www.pnas.org/content/105/33/11927.full.pdf [6]] Gaballa, A., et al. (2008). "The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins." Proc Natl Acad Sci U S A 105(33): 11927-11932. |
| | | | |
| | | | |
| Line 1,018: |
Line 1,019: |
| | [[File:Design Fig.3.png|600px|center|Fig. 3.]] | | [[File:Design Fig.3.png|600px|center|Fig. 3.]] |
| | | | |
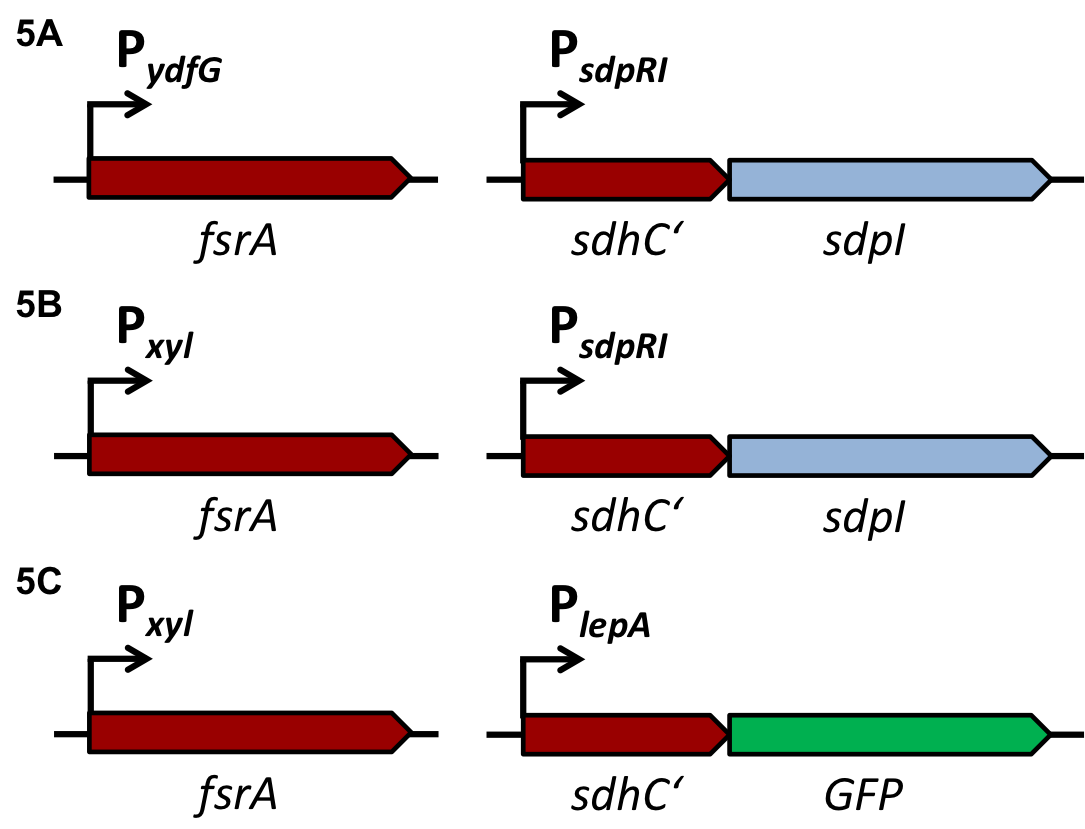
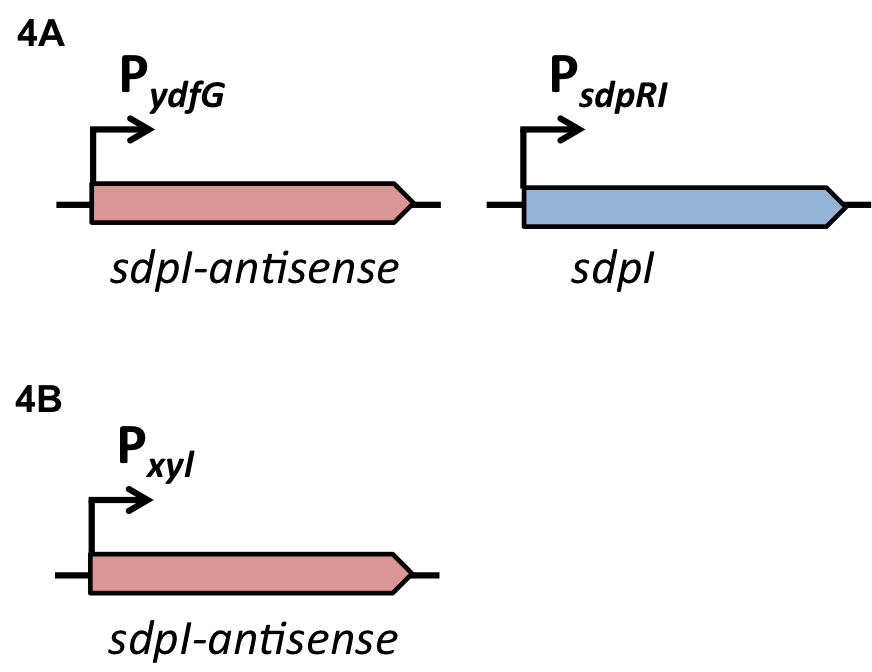
| - | For the loss of immunity we implemented two strategies, both causing translational inhibition of the sdpI mRNA. | + | For the loss of immunity we implemented two strategies, both causing translational inhibition of the ''sdpI'' mRNA. |
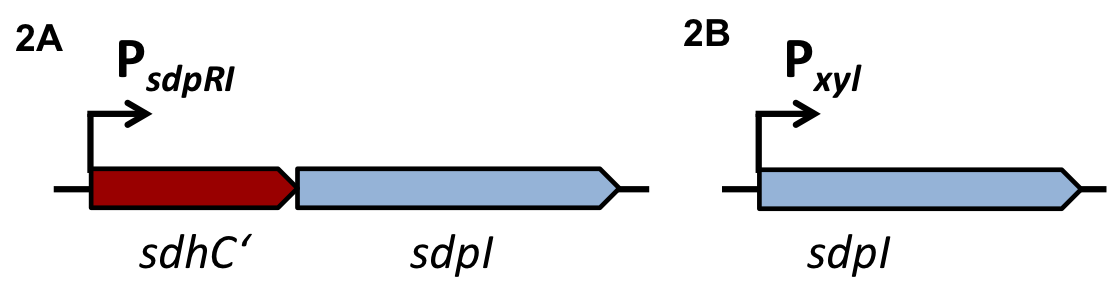
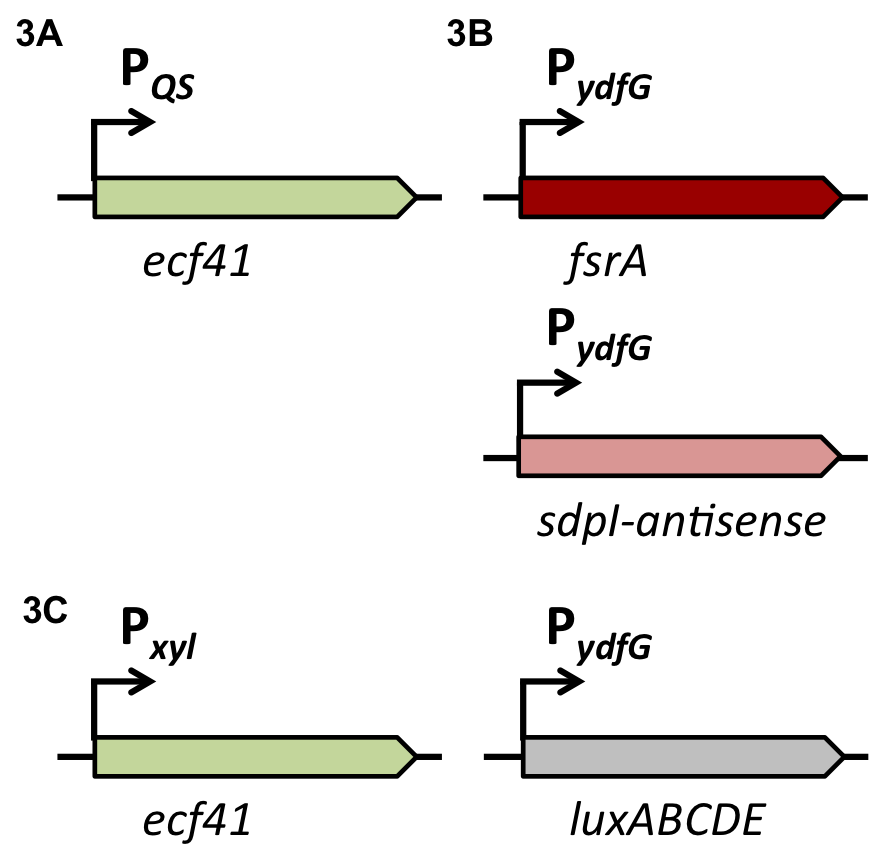
| | The first strategy is based on an artificial synthesized RNA called sdpI-antisense RNA, which is complementary to the sdpI mRNA sequence. The sdpI-antisense sequence is regulated by P<sub>''ydfG''</sub> (Fig. 4A) which is activated by the binding of Ecf41<sub>''bli aa 1-204''</sub>. As a consequence the sdpI-antisense RNA and the sdpI mRNA form a double-stranded duplex which is inaccessible for the ribosome making the cell SDP-sensitive. | | The first strategy is based on an artificial synthesized RNA called sdpI-antisense RNA, which is complementary to the sdpI mRNA sequence. The sdpI-antisense sequence is regulated by P<sub>''ydfG''</sub> (Fig. 4A) which is activated by the binding of Ecf41<sub>''bli aa 1-204''</sub>. As a consequence the sdpI-antisense RNA and the sdpI mRNA form a double-stranded duplex which is inaccessible for the ribosome making the cell SDP-sensitive. |
| | We tested the sdpI-antisense RNA in B. subtilis W168 with the xylose-inducable promoter P<sub>''xyl''</sub>. (Fig. 4B) | | We tested the sdpI-antisense RNA in B. subtilis W168 with the xylose-inducable promoter P<sub>''xyl''</sub>. (Fig. 4B) |
| Line 2,665: |
Line 2,666: |
| | The graph shows wildtype <i>Bacillus subtilis</i>, the ΔsdpI mutant as well as a ΔsdpI mutant carrying our generated </html>([http://parts.igem.org/Part:BBa_K1351017 BBa_K1351017]) in addition the graph shows a wildype <i>Bacillus subtilis</i> carrying an antisense RNA [http://parts.igem.org/Part:BBa_K1351019 (BBa_K1351019)] sequence which should bind the mRNA of the sdpI immunity. The LUMI-assay shows a normal growth curve of <i>Bacillus subtilis</i> and a ΔsdpI mutant which degrades in early to late stationary phase. <html>The two from us constructed devices show in case of the inducible immunity in the ΔsdpI mutant an almost recovery in growth behavior when compared to wild type. The antisense carrying W168-strain showed normal wildtype growth and it seems like that an translational inhibition of the immunity was not functioning. <br><br><br> | | The graph shows wildtype <i>Bacillus subtilis</i>, the ΔsdpI mutant as well as a ΔsdpI mutant carrying our generated </html>([http://parts.igem.org/Part:BBa_K1351017 BBa_K1351017]) in addition the graph shows a wildype <i>Bacillus subtilis</i> carrying an antisense RNA [http://parts.igem.org/Part:BBa_K1351019 (BBa_K1351019)] sequence which should bind the mRNA of the sdpI immunity. The LUMI-assay shows a normal growth curve of <i>Bacillus subtilis</i> and a ΔsdpI mutant which degrades in early to late stationary phase. <html>The two from us constructed devices show in case of the inducible immunity in the ΔsdpI mutant an almost recovery in growth behavior when compared to wild type. The antisense carrying W168-strain showed normal wildtype growth and it seems like that an translational inhibition of the immunity was not functioning. <br><br><br> |
| | | | |
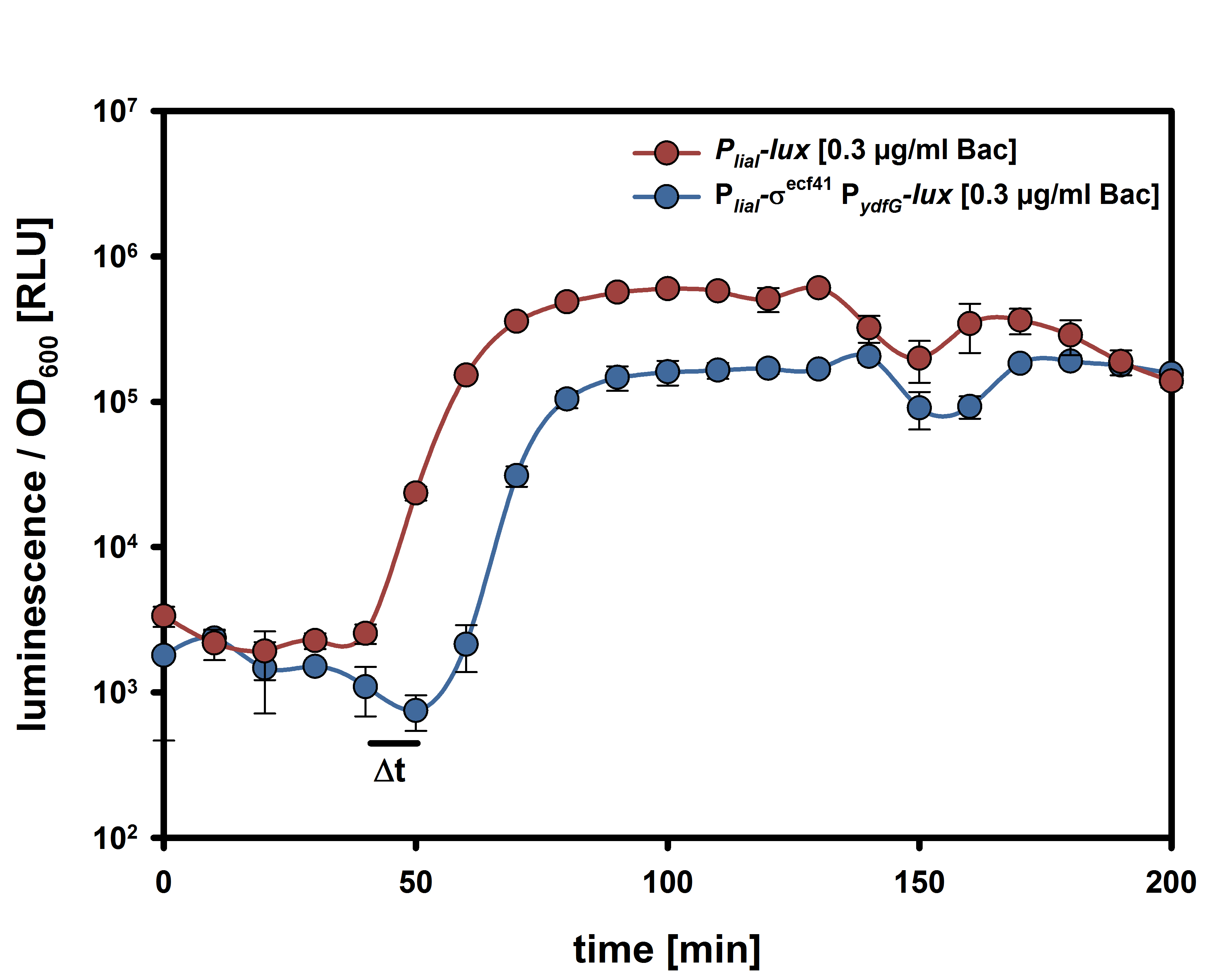
| - | For the suicide switch we figured that a time delay was necessary before the suicide switch kicks in so that BaKillus has enough time to do his job. Therefore we decided to test if an insertion of an alternative sigma factor would lead to a time delay. For this purpose we designed two constructs: one having an inducible promoter in front of a lux-readout and the other device carrying an alternative sigma (ECF41 </html>[http://parts.igem.org/Part:BBa_K823043 BBa_K823043]<html>) factor after the same inducible promoter. The sigma factor then has to find its own specific promoter (P<sub>ydfG</sub> </html>[http://parts.igem.org/Part:BBa_K823041 BBa_K823041]<html>) to give then a lux-readout.<br><br> | + | For the suicide switch we figured that a time delay was necessary before the suicide switch kicks in so that BaKillus has enough time to do his job. Therefore we decided to test if an insertion of an alternative sigma factor would lead to a time delay. For this purpose we designed two constructs: one having an inducible promoter in front of a lux-readout and the other device carrying an alternative sigma (ECF41 </html>[http://parts.igem.org/Part:BBa_K823043 BBa_K823043]<html>) factor after the same inducible promoter. The sigma factor has to find its own specific promoter again (P<sub>ydfG</sub> </html>[http://parts.igem.org/Part:BBa_K823041 BBa_K823041]<html>) which was cloned in front of the lux-operon.<br><br> |
| - | The LUMI-assay in the graph shows that we could achieve a time delay of about ten minutes using the alternative sigma factor route. For a reasonable delay switch we intend to implant a series of alternative sigma factors and their target promoters to achieve a cascade. | + | The </html>[https://static.igem.org/mediawiki/2014/c/c7/LMU-Munich14_Luminescence_Assay.pdf LUMI-assay]<html> in the graph shows that we could achieve a time delay of about ten minutes using the alternative sigma factor route. For a reasonable delay we intend to implement a series of alternative sigma factors and their target promoters to achieve a cascade and therefore increase the time delay. |
| | <br> | | <br> |
| | </html> | | </html> |
 "
"