Team:Aachen/Collaborations/Freiburg
From 2014.igem.org
m (→AcCELLoMatrix Reader App) |
(→References) |
||
| (25 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
{{Team:Aachen/Header}} | {{Team:Aachen/Header}} | ||
| - | = AcCELLoMatrix | + | = AcCELLoMatrix for [[Team:Freiburg/Team/Collaboration|Team Freiburg]]= |
| - | At the iGEM meetup in Munich in May , members of both of our teams realized that our projects share a common objective: Analyzing 2-dimensional, visual signals. For the following months, we stayed in contact and developed a concept to encode information in 384-well plates in the form of data matrix codes. | + | At the iGEM meetup in Munich in May, members of both of our teams realized that our projects share a common objective: Analyzing 2-dimensional, visual signals. For the following months, we stayed in contact and developed a concept to encode information in 384-well plates in the form of data matrix codes. |
At the beginning of September, Michael spontaneously took a train to Freiburg and we met face-to-face for several hours to exchange details on our cooperation and general iGEM experiences. | At the beginning of September, Michael spontaneously took a train to Freiburg and we met face-to-face for several hours to exchange details on our cooperation and general iGEM experiences. | ||
| + | |||
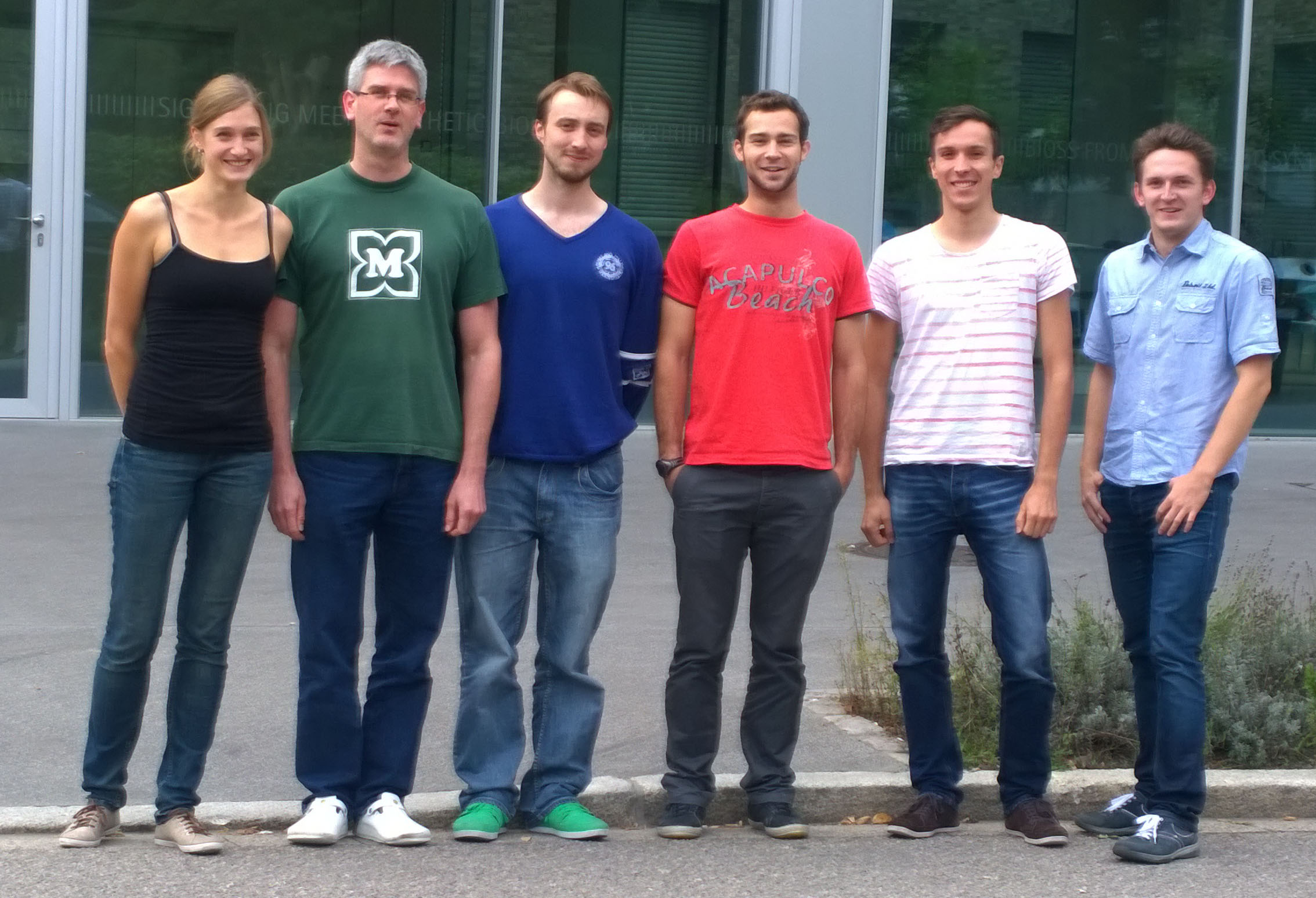
{{Team:Aachen/Figure|Aachen_FR-collaboration_group_photo.jpg|align=center|title=Spontaneous group photo|subtitle=Freíburg iGEMers and Michael (right) meeting to discuss our cooperation|width=350px}} | {{Team:Aachen/Figure|Aachen_FR-collaboration_group_photo.jpg|align=center|title=Spontaneous group photo|subtitle=Freíburg iGEMers and Michael (right) meeting to discuss our cooperation|width=350px}} | ||
| - | == Data Matrix Masks | + | |
| + | == Data Matrix Masks == | ||
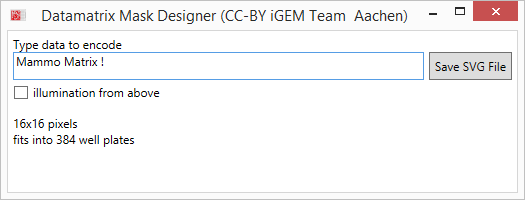
We decided to use masks to locally induce the cells in 384-well plates in a data matrix pattern. To enable us to quickly design masks for different data to be encoded, we wrote a software to easily generate SVG-files for a laser cutter. | We decided to use masks to locally induce the cells in 384-well plates in a data matrix pattern. To enable us to quickly design masks for different data to be encoded, we wrote a software to easily generate SVG-files for a laser cutter. | ||
| Line 26: | Line 28: | ||
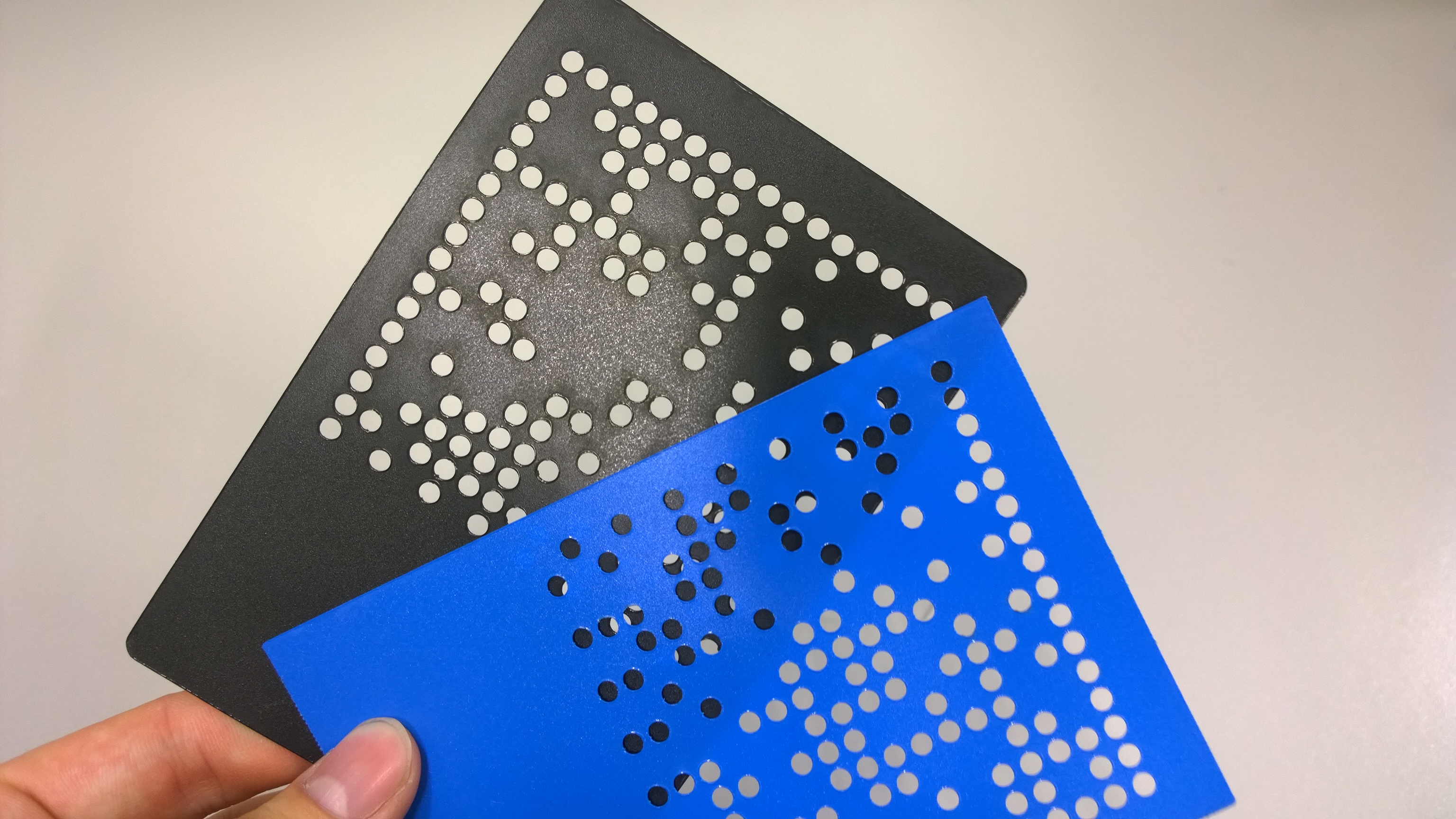
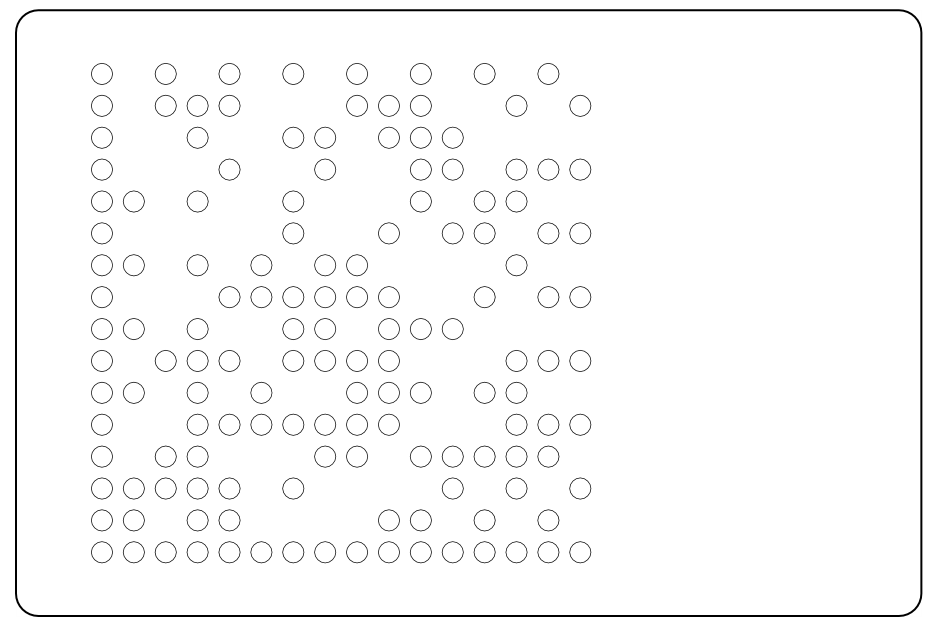
For the first experiments, we cut two different data matrix patterns from polypropylen foil. The masks can be used to cover the plates from the top or from the bottom. | For the first experiments, we cut two different data matrix patterns from polypropylen foil. The masks can be used to cover the plates from the top or from the bottom. | ||
| - | {{Team:Aachen/Figure|Aachen_MammoMatrixMasks1.jpg|align=center|title=AcCELLoMatrix masks|subtitle=These masks can be placed on top or beneath 384-well plates to facilitate localized illumination of wells.|width=350px}} | + | {{Team:Aachen/Figure|Aachen_MammoMatrixMasks1.jpg|align=center|title=AcCELLoMatrix masks|subtitle=These masks can be placed on top or beneath 384-well plates to facilitate localized illumination of wells.|width=350px}} |
== AcCELLoMatrix Reader App == | == AcCELLoMatrix Reader App == | ||
| - | After mask have been designed and the information was encoded into the well plate by selectively illuminating the cells, how do you get it out again? We made an app for that: the AcCELLoMatrix Reader for Windows Phone. | + | After mask have been designed and the information was encoded into the well plate by selectively illuminating the cells, how do you get it out again? We made an app for that: the AcCELLoMatrix Reader for Windows Phone 8. |
We designed the app to be easy to use: A user can simply open the app | We designed the app to be easy to use: A user can simply open the app | ||
| - | {{Team:Aachen/Figure|Aachen_Collaboration- | + | {{Team:Aachen/Figure|Aachen_Collaboration-FRScreenshots.png|title=Usage of AcCELLoMatrix Reader|width=1018px|subtitle=(1) Start the app and read through some brief instructions (2). Then start scanning and aim for a datamatrix in the wild (3). As soon as a datamatrix is detected in the preprocessed frames (4, small rectangle) by the ZXing algorithm, the decoded text is displayed to the user (5).}} |
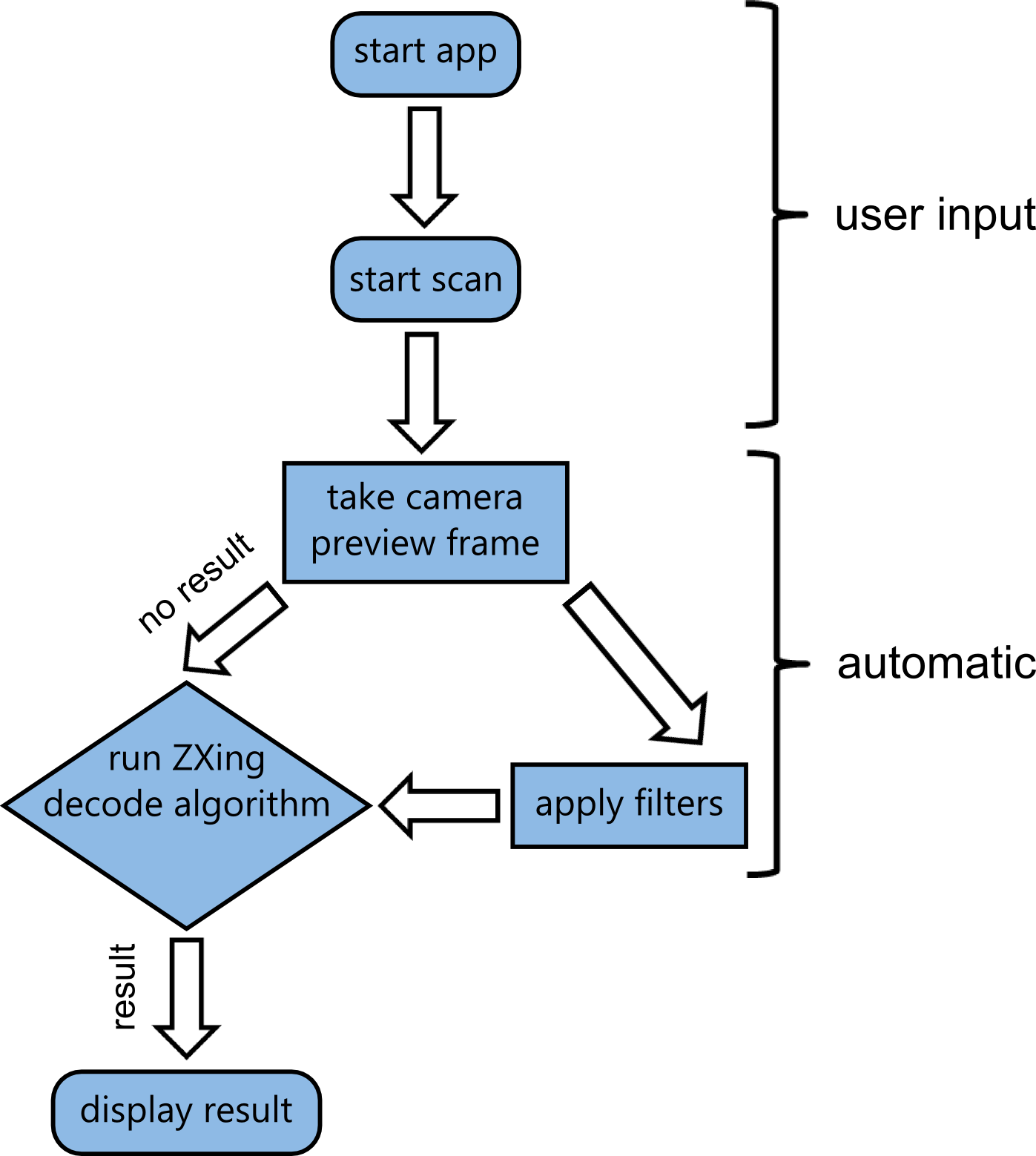
| - | + | While the apps frontend is designed to be very minimalistic, there's actually going on a lot in the backend. Visual data coming from the camera feed has to be processed and fed into the ZXing decoding algorithm. | |
| - | {{Team:Aachen/Figure|Aachen_Collaboration- | + | {{Team:Aachen/Figure|Aachen_Collaboration-FRAMRflow.png|title=Activity diagram of the app|width=350px|subtitle=After the app was started and the scanning initiated by the user, a loop is initiated. Filters are applied to camera preview frames and the result is fed into the decoding algorithm. As soon as a datamatrix is detected, the loop is interrupted and the result presented to the user.}} |
| - | . | + | == Outlook == |
| + | Due to the circumstance that this collaboration was started relatively late, we have not been able to test the decoding of real AcCELLoMatrixes. But we are confident, that with some threshold adjustments to the AcCELLoMatrixes Reader app and maybe the hole-size of the polypropylen masks, the coding and decoding of short text information in the genetic state of mammalian cells in a 384-well plate is perfectly possible. | ||
| - | + | == References == | |
| + | [http://www.nuget.org/packages/ZXing.Net/ ZXing.NET] by Michael Jahn is a port of [https://github.com/zxing/zxing ZXing] ("zebra crossing"), both licensed under [http://www.apache.org/licenses/LICENSE-2.0 Apache 2.0] | ||
| - | + | {{Team:Aachen/BlockSeparator}} | |
| - | + | = Characterization of our OD Device by Team Freiburg = | |
| + | As we were helping the Freiburg team out with the encoding/decoding of text as datamatrixes, they also wanted to help us in a useful way. Our team was committed to characterize the OD measurement device, so we sent them one of prototypes and asked them to try a measurement of their mammalian cells. | ||
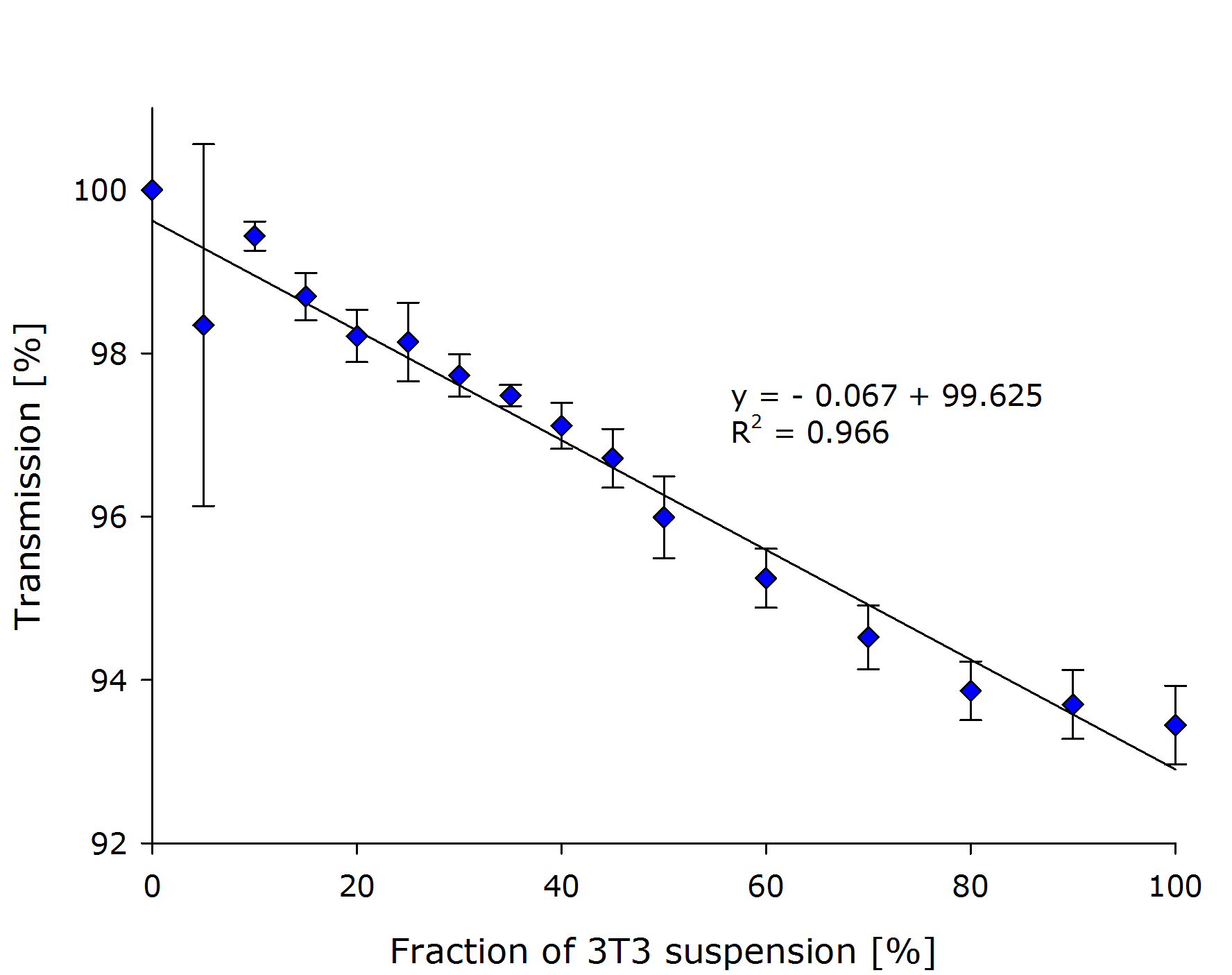
| - | == | + | They prepared a dilution series with biological triplicates of their NIH3T3 mouse fibroblasts in the respective growth medium. The absolute cell number was determined at 10{{sup|6}} cells/ml. The transmission of light that was measured by our prototype, is plotted against the fraction of cell culture in the sample in the left chart: |
| + | |||
| + | |||
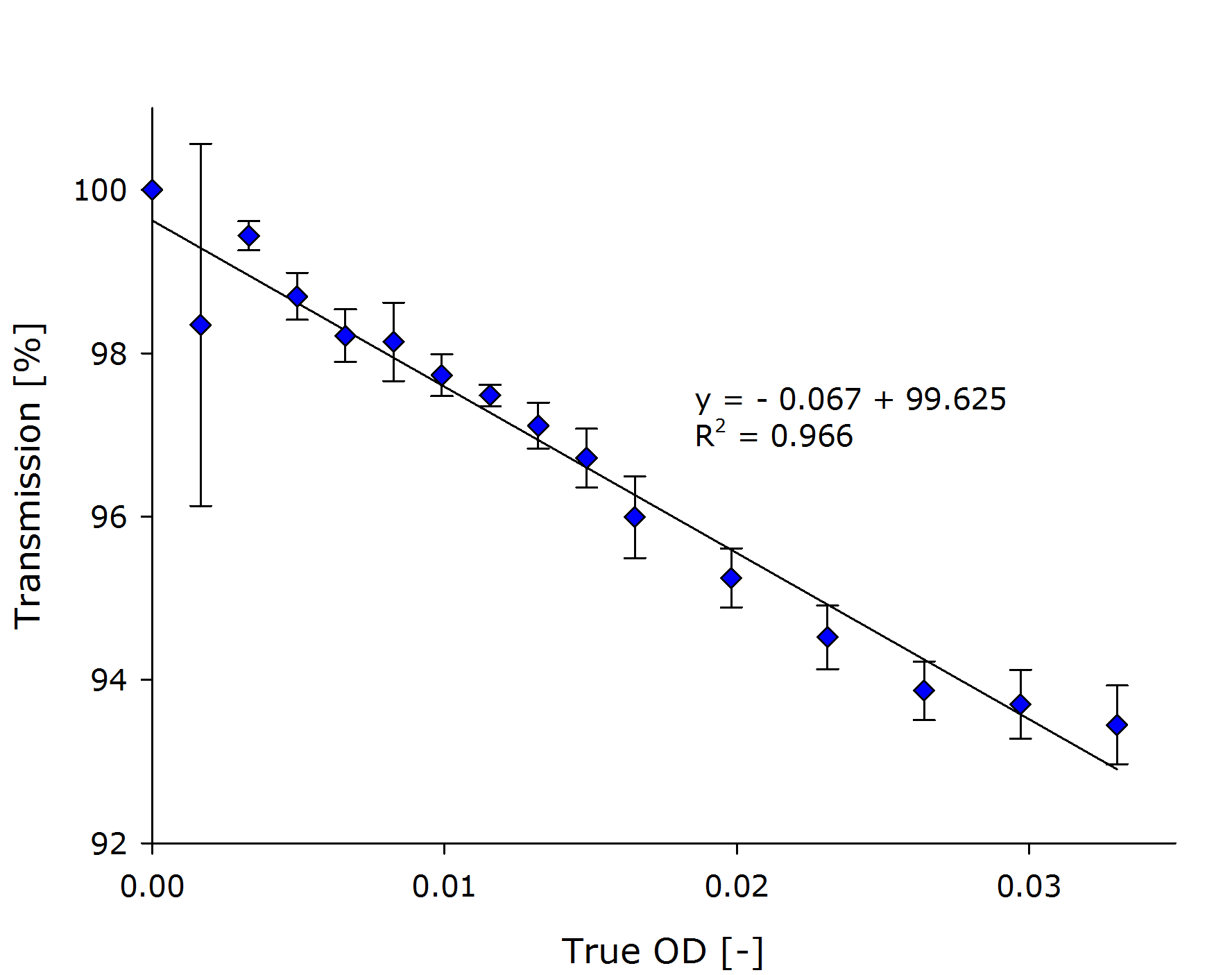
| + | {{Team:Aachen/FigureDual|Aachen_Collaboration-FR_NIH3T3_1.png|Aachen_Collaboration-FR_NIH3T3_2.png|title1=Transmission at different concentrations|title2=Transmission versus true OD of the sample|subtitle1=A linear dependency was observed between the calculated concentration of cells transmission [%].|subtitle2=Calculation of the true ODs revealed, that the optical density of the samples was very low.|width=425px}} | ||
| + | |||
| + | From the transmittance data of the dilution series, we calculated the [[Team:Aachen/Notebook/Engineering/ODF#Evaluation|true OD]] of the sample (right chart above). | ||
| + | |||
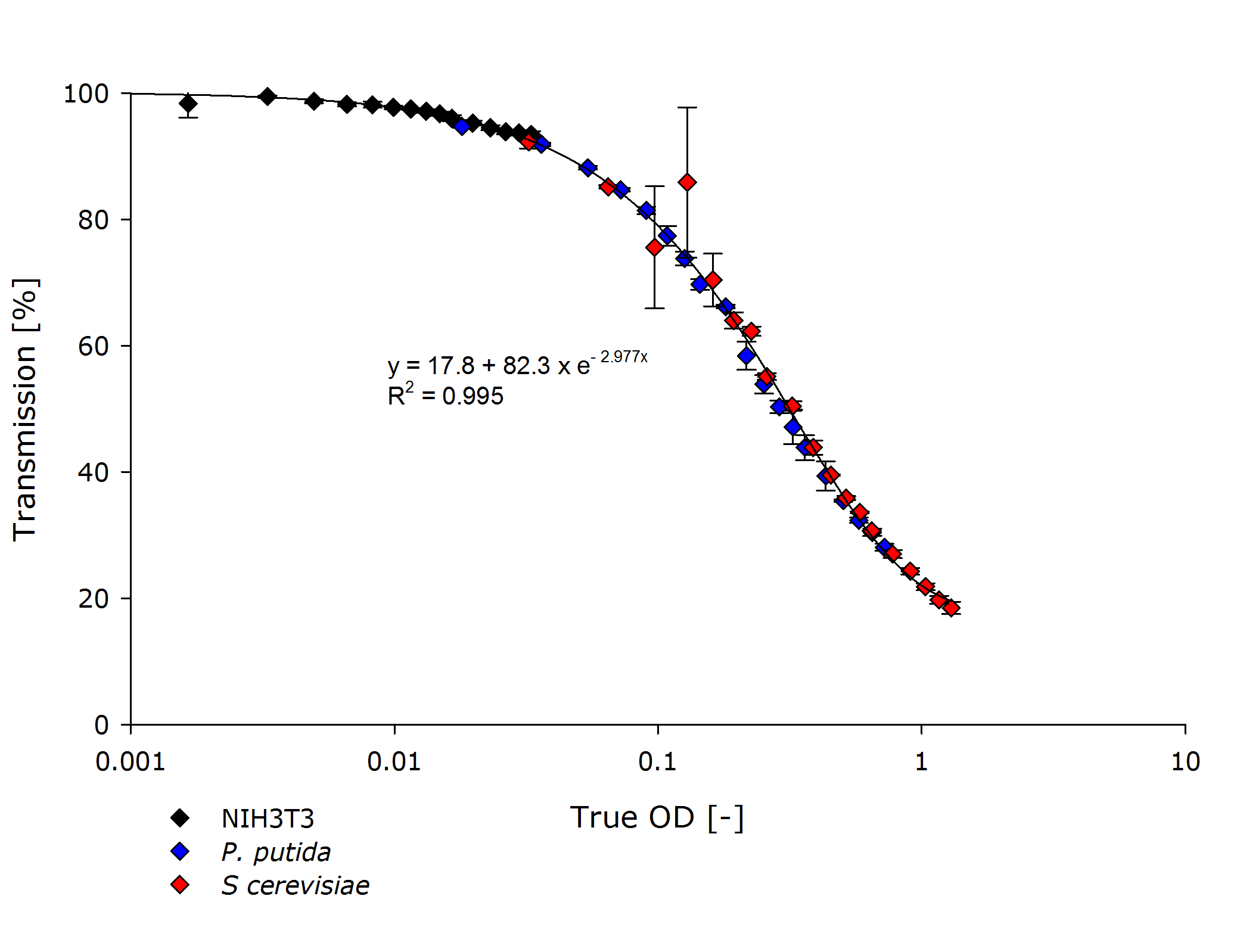
| + | Finally, we incorporated the measurements by team Freiburg into our characterization of the OD-measurement device that now covers multiple strains over three orders of magnitude: | ||
| + | |||
| + | {{Team:Aachen/Figure|Aachen_ODallstrains1.png|title=Transmission of different cell types at OD-values from 0.001-1|subtitle=The transmittance data of NIH 3T3 cells align with the transmittance of ''P. putida'' and ''S. cerevisiae'' strains, even though the measured optical densities are lower by 1-2 orders of magnitude.|width=800px}} | ||
| + | |||
| + | Our friends in Freiburg also tested a Nanodrop, but reported that this commercial device was unable to quantify the diluted samples. In the end we were very satisfied with the results of our cooperation! | ||
| - | |||
| - | |||
{{Team:Aachen/Footer}} | {{Team:Aachen/Footer}} | ||
Latest revision as of 03:38, 18 October 2014
|
|
 "
"