Team:CU-Boulder/Project/Modeling
From 2014.igem.org
(Created page with "{{:Team:CU-Boulder/Wiki}} {{Template:UCB-NavBar}} <html>") |
|||
| (9 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | {{: | + | {{Template:UCB-Main}} |
{{Template:UCB-NavBar}} | {{Template:UCB-NavBar}} | ||
| - | < | + | |
| + | == '''Modeling''' == | ||
| + | |||
| + | ===Intro=== | ||
| + | The CRISPR Cas9 system is comprised of the Cas9 endonuclease and a guide RNA that is coded by a 20nt spacer sequence. The complex formed by these parts identifies target sites by scanning the DNA for a NGG motif (referred to as a Protospacer Adjacent Motif, or PAM site). If found, the complex checks the adjacent DNA sequence for complementarity to the guiding RNA. If the DNA-RNA annealing interaction is sufficient, the Cas9 endonuclease can successfully bind and cleave the DNA. | ||
| + | The guiding RNA is coded by a 20 nucleotide sequence called a protospacer. Positions within this sequence carry varying weights of significance. The seed region (the last eight nucleotides) must be an exact match for the Cas9 protein to bind. The 12 nucleotides preceding the seed region can tolerate up to five mismatches; however, each mismatch decreases the probability of Cas9 binding. This degeneracy allows for off-targeting of similar sequences, presenting a problem in a mixed population of bacteria where the preservation of certain strains may be desired. | ||
| + | |||
| + | |||
| + | [[File:UCB-modeling wiki content-01-141013.JPG]] | ||
| + | |||
| + | |||
| + | http://eendb.zfgenetics.org/casot/ | ||
| + | |||
| + | <br> | ||
| + | ===The Model=== | ||
| + | |||
| + | Successful binding of Cas9 depends on the presence of a PAM site and the strength of the guide RNA-DNA interaction. These two features make it possible to predict successful CRISPR-Cas9 binding. | ||
| + | If a spacer is to be used in a mixed population of bacteria comprised of strains to ‘kill’ and strains to ‘keep’, the ideal spacer sequence can be computed programmatically. The spacer sequence should be present and adjacent to a PAM site within the ‘kill’ set genome, but absent from the genomes of the ‘keep’ set. This allows the Cas9 protein, programmed with this sequence, to target only the desired genomes. The model described below determines a sequence that is unique to the ‘kill’ set, and absent from the ‘keep’ set. | ||
| + | |||
| + | The program accepts the fasta files containing the target genomes and the files containing the other non-targeted genome. The program then finds every protospacer adjacent to an NGG in the genomes of the target bacteria and sorts the sequences by the seed region. Each sequence containing a particular seed region is scored or ranked based on the number of genomes it is found in, if it is a perfect match or an off-target site, and the number of time it is found in a given genome. Each of these 20mers is scored similarly against every 20mer found in the genome that is not targeted. The non-target score for each sequence is then subtracted from the target score to calculate a total score for each protospacer. The sequences are then ranked by this total score. | ||
| + | |||
| + | |||
| + | <br> | ||
| + | ===Results=== | ||
| + | |||
| + | |||
| + | To test the model’s ability to design unique spacers a neomycin phosphotransferase gene was run as the target genome. The ‘keep’ set contained ''E. coli'' K-12 and ''E. coli'' MG1655. The following is a subsection of the output: | ||
| + | |||
| + | [[File:UCB-modeling wiki content-02-141013.JPG]] | ||
| + | |||
| + | The first line represents the protospacer that is found in the target genome. The next line is the total score of the protospacer. The sequences are sorted by these total scores. The third line is the score representing how well the protospacer binds to the target genome. The second indentation is the genome the protospacer is found in. The third indentation is the binding site found within the genome. The fourth indentation represents how well the Cas9 protein will bind. First the results found in the target genomes are shown, followed by the results for the non-target genomes, for each protospacer. | ||
| + | |||
| + | <br> | ||
| + | ===Future Directions=== | ||
| + | |||
| + | |||
| + | Improvements still need to be made on the web interface. For instance, the user will be able to upload fasta files for their ‘keep’ and ‘kill’ sets. In addition, the program will be parallelized to allow many genomes to be run at the same time, decreasing computation time. | ||
| + | |||
| + | <br> | ||
| + | ===Assumptions=== | ||
| + | |||
| + | #A bacteria with multiple off-target sites will be killed better than a bacteria with a single off-target site. | ||
| + | |||
| + | |||
| + | {{Template:UCB-Footer}} | ||
Latest revision as of 09:36, 17 October 2014
Contents |
Modeling
Intro
The CRISPR Cas9 system is comprised of the Cas9 endonuclease and a guide RNA that is coded by a 20nt spacer sequence. The complex formed by these parts identifies target sites by scanning the DNA for a NGG motif (referred to as a Protospacer Adjacent Motif, or PAM site). If found, the complex checks the adjacent DNA sequence for complementarity to the guiding RNA. If the DNA-RNA annealing interaction is sufficient, the Cas9 endonuclease can successfully bind and cleave the DNA. The guiding RNA is coded by a 20 nucleotide sequence called a protospacer. Positions within this sequence carry varying weights of significance. The seed region (the last eight nucleotides) must be an exact match for the Cas9 protein to bind. The 12 nucleotides preceding the seed region can tolerate up to five mismatches; however, each mismatch decreases the probability of Cas9 binding. This degeneracy allows for off-targeting of similar sequences, presenting a problem in a mixed population of bacteria where the preservation of certain strains may be desired.
http://eendb.zfgenetics.org/casot/
The Model
Successful binding of Cas9 depends on the presence of a PAM site and the strength of the guide RNA-DNA interaction. These two features make it possible to predict successful CRISPR-Cas9 binding. If a spacer is to be used in a mixed population of bacteria comprised of strains to ‘kill’ and strains to ‘keep’, the ideal spacer sequence can be computed programmatically. The spacer sequence should be present and adjacent to a PAM site within the ‘kill’ set genome, but absent from the genomes of the ‘keep’ set. This allows the Cas9 protein, programmed with this sequence, to target only the desired genomes. The model described below determines a sequence that is unique to the ‘kill’ set, and absent from the ‘keep’ set.
The program accepts the fasta files containing the target genomes and the files containing the other non-targeted genome. The program then finds every protospacer adjacent to an NGG in the genomes of the target bacteria and sorts the sequences by the seed region. Each sequence containing a particular seed region is scored or ranked based on the number of genomes it is found in, if it is a perfect match or an off-target site, and the number of time it is found in a given genome. Each of these 20mers is scored similarly against every 20mer found in the genome that is not targeted. The non-target score for each sequence is then subtracted from the target score to calculate a total score for each protospacer. The sequences are then ranked by this total score.
Results
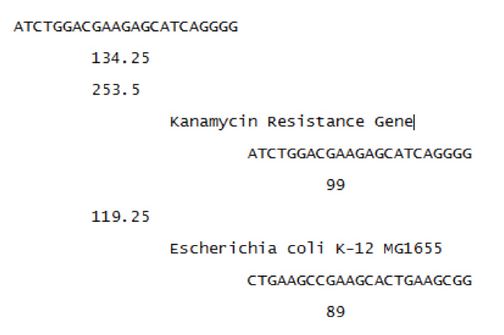
To test the model’s ability to design unique spacers a neomycin phosphotransferase gene was run as the target genome. The ‘keep’ set contained E. coli K-12 and E. coli MG1655. The following is a subsection of the output:
The first line represents the protospacer that is found in the target genome. The next line is the total score of the protospacer. The sequences are sorted by these total scores. The third line is the score representing how well the protospacer binds to the target genome. The second indentation is the genome the protospacer is found in. The third indentation is the binding site found within the genome. The fourth indentation represents how well the Cas9 protein will bind. First the results found in the target genomes are shown, followed by the results for the non-target genomes, for each protospacer.
Future Directions
Improvements still need to be made on the web interface. For instance, the user will be able to upload fasta files for their ‘keep’ and ‘kill’ sets. In addition, the program will be parallelized to allow many genomes to be run at the same time, decreasing computation time.
Assumptions
- A bacteria with multiple off-target sites will be killed better than a bacteria with a single off-target site.
 "
"