Team:IvyTech SouthBend IN/progress
From 2014.igem.org
Data Review 10/14/14

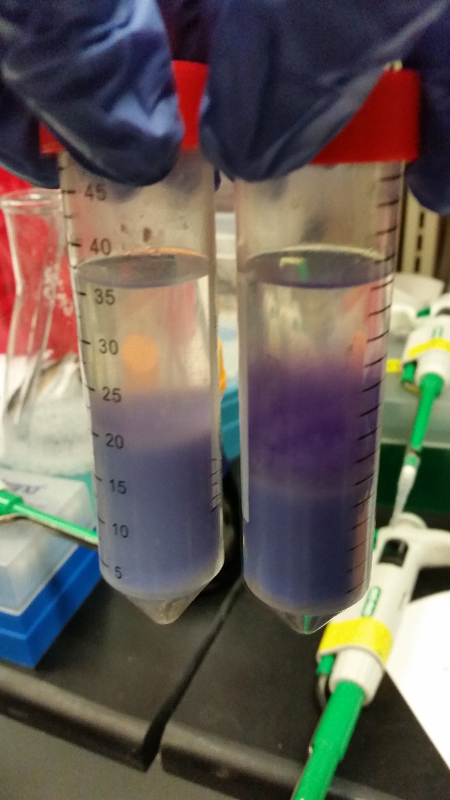
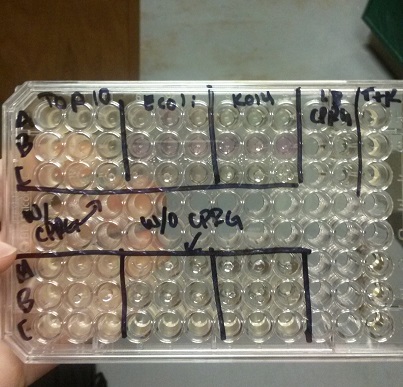
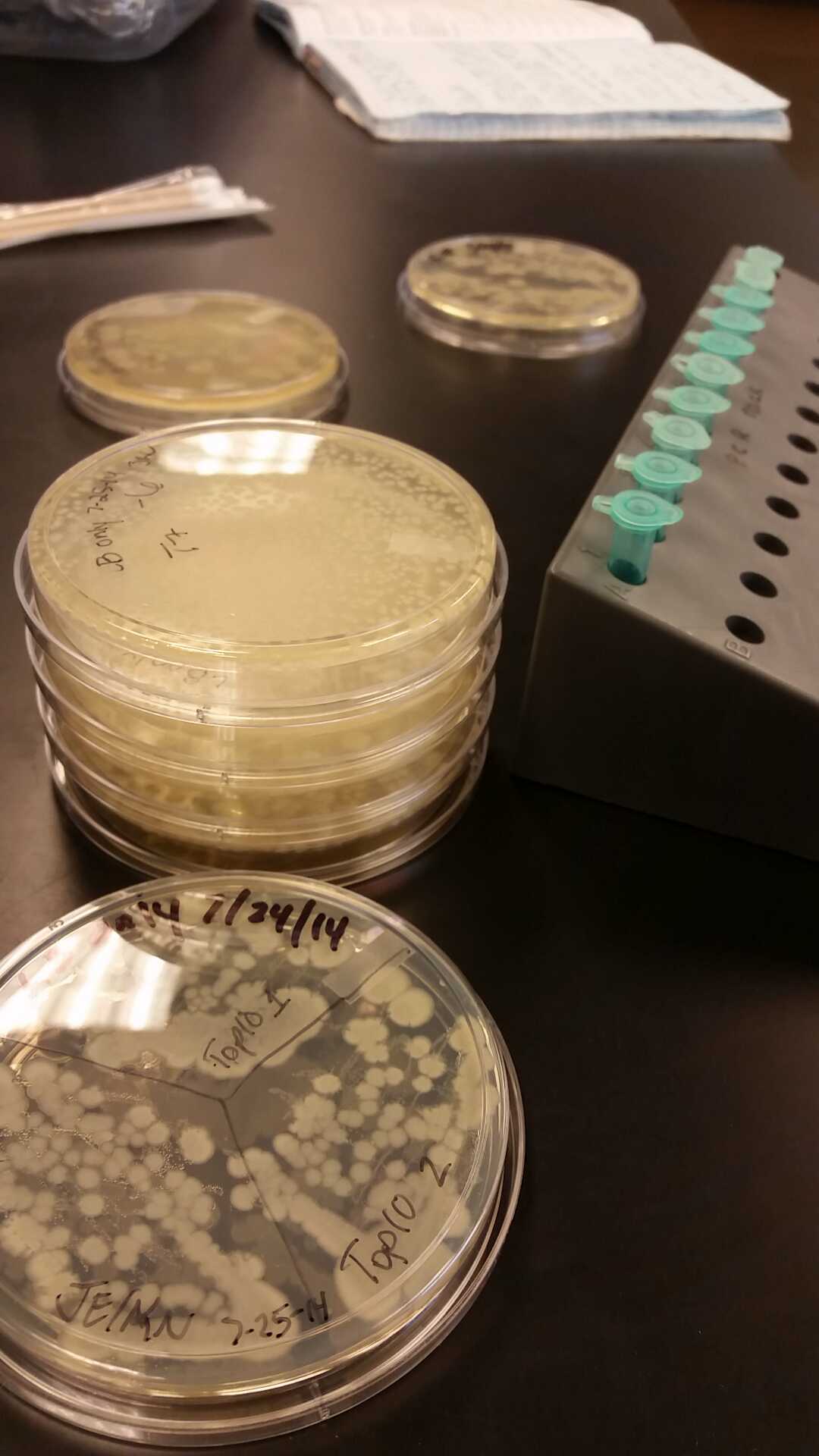
- Data from a replication of Stanick’s assay with K1477014: Cells were grown overnight in LB or LB/glucose. Cells in LB were stimulated with 200 uM IPTG for 1 hour and then all cell cultures were divided into 1mL aliquots to which calcium chloride and magnesium sulfate were added. T7 bacteriophage (1 x 10^5 pfu) was added to the appropriate culture and the absorbance at A600 was followed until virus lysis was evident. Cell cultures were then transferred to SpinX filter devices and the media was separated from the cells. Aliquots of filtrates from cells only, T7-treated cultures, or from T7-treated culture were transferred to a microtiter plate. Buffer Z with 2 mercaptoethanol was added and absorbance at 415 nm was determined after 30 minutes.
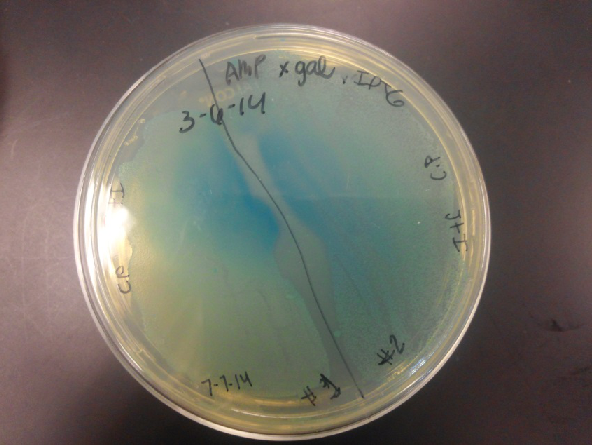
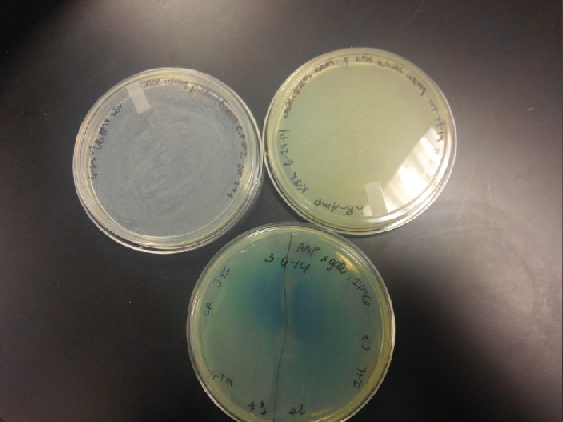
- Results: Both E.coli C and K1477014 cells stimulated with IPTG “leaked” β-gal activity (Cells only). Top10 cells were negative as expected. β-gal activity was suppressed in all cell lines by glucose as expected. T7 treatment resulted in an increased release of β-gal activity from both E. coli C and K1477014. The combination of T7 lysates from Top10 cells stimulated with IPTG and K1477014 cells grown in glucose did not show any β-gal activity. .
- Conclusion: Cells stimulated with IPTG and subjected to this protocol release β-gal activity. This might be due to β-gal activity that accumulated in the media as the cells grew. Another possibility is that β-gal activity was released when the calcium chloride or magnesium sulfate solutions were added before virus lysis or perhaps released during centrifugation of the cells in the SpinX columns.
- Most important however is that β-gal activity was found in the medium of lysed and unlysed K1477014 suggesting that the LacZ alpha and omega polypeptides stay combined once they are released! That is great news!
- T7 treatment caused an increase in the release of β-gal enzyme from the E. coli C and the β-gal enzyme (alpha-omega polypeptide combo) from K1477014.
- Unfortunately, the combination of T7 lysates of Top 10 cells treated with IPTG, which should have formed LacZ omega polypeptides and the K1477014 cells treated with glucose, which should have formed only LacZ alpha polypeptides, did not have any detectable β-gal activity.
- This lack of activity could be due to the slow association of the alpha and omega fragments, maybe slowed by the Buffer Z and calcium and magnesium salts or, worse, it means that the alpha and omega polypeptides have to associate inside a cell to be joined into a working enzyme. We can study this further by letting the combination of fragments incubate together for hours in hopes that β-gal activity appears. If it does that is great because that is the proof of principle that we are after.
Week of 8/3/14 through 8/9/14
- During this week, the team realized that the contamination was coming from our stock of cells that was being kept in the -80C freezer. We have no idea how they became contaminated, seeing as continued to use the line of cells we froze for a week after we froze them, but before any contamination occurred. Nonetheless, one of our advisers has procured a new stock of top 10 E. coli cells for us to use.
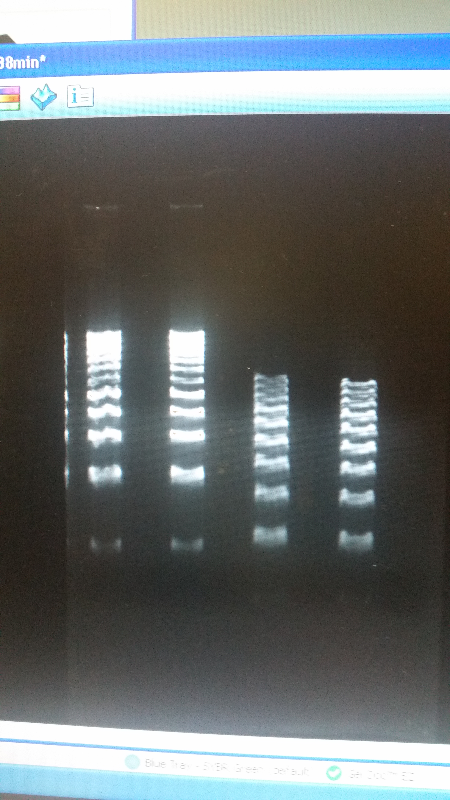
- After unsuccessful runs of electrophoresis gels, we have run out of DNA to try to measure. So, we took some time out this week to streak, grow, and purify lines of our K1477014 and K1477030 parts on both pSB1C3 and pSB1AT3 backbones.
Week of 7/13/14 through 7/19/14
- This week we worked towards switching our part's backbone from pSB1AT3 to pSB1C3. We had some difficulty doing this. The first attempt was fruitless, but on our second attempt we had success and we now have a stock of our part on a chlor-resistant backbone.
- Unfortunately, We no longer have any stock of our original part. So, now we are again following the 3A Biolabs Biobrick Assembly Manual to reconstruct our parts K1477014 and K1477030. Our first try yielded nothing.
- Our second run through of the protocol resulted in the growth of red colonies on chlor plates after electroporation of our part into Top 10 cells. However, we need white colonies to grow because this will point towards the rfp being repressed when the three parts come together. Red colonies are a sign that we did not have a successful assembly of our parts.
Week of 5/25/14 through 5/31/14
- We transformed Top 10 cells through electroporation with registry parts K112806 (Endolysin,) I732020 (LacZoperon Mutant,) J23102 (p-con) and I712074 (T7 promoter.)
- After streaking on plates, and leaving them in the incubator for a day, there was growth on all plates, showing that a successful transformation occured.
- Plates containing J23102 were red-another sign of success. Unfortunately, it looks like contamination has arisen as well, and is spreading on plates with I712074 and K112806 growth.
- The contamination persisted all week, but we were able to get plates with transformed parts that contained no contamination growth.
- We then ran a DNA purification protocol of J23102 and I732020 to get them ready for a biobrick assembly protocol.
 "
"