Team:Aachen/Project/FRET Reporter
From 2014.igem.org
(→The FRET (Förster Resonance Energy Transfer) System) |
|||
| Line 135: | Line 135: | ||
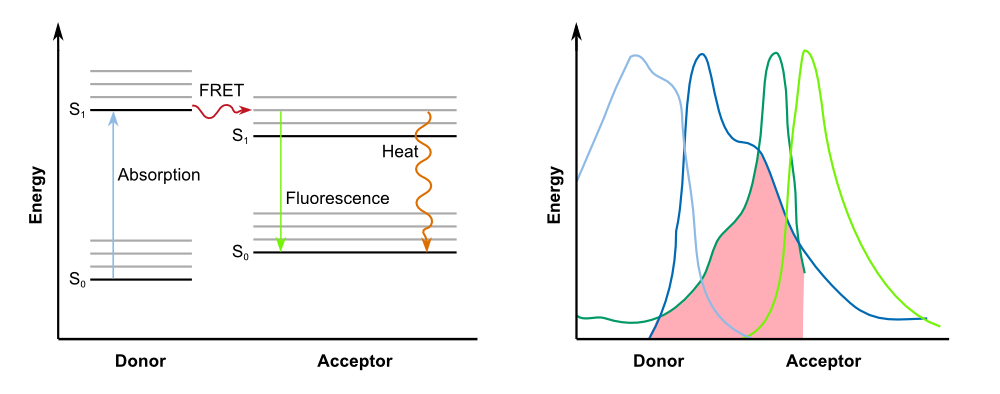
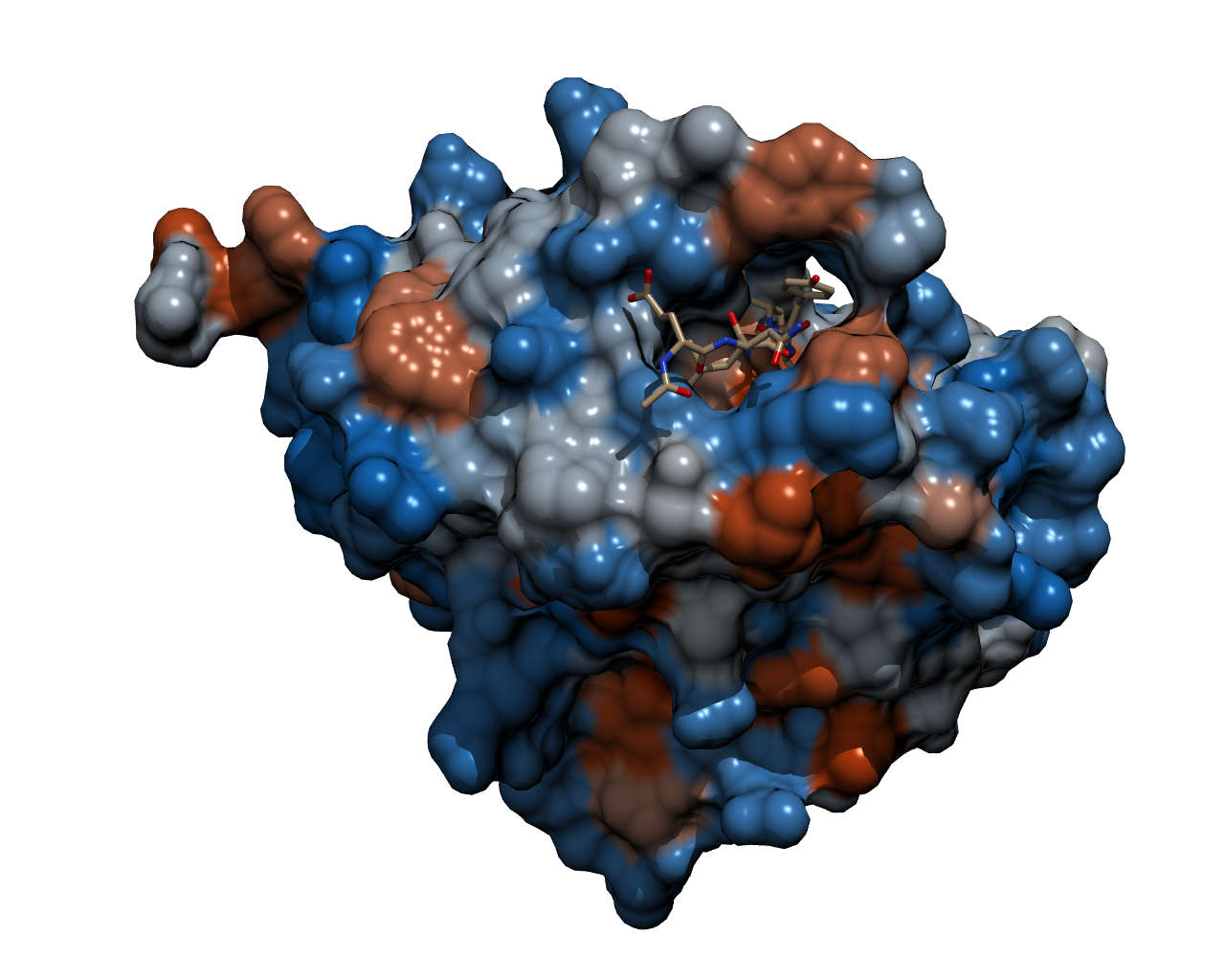
To optimize the spectral overlap of this FRET pair, the group obtained '''a genetically modified YFP acceptor'''. Mutations of amino acid residues that stabilize the excited state of the chromophore in enhanced YFP (EYFP) resulted in a non-fluorescent chromoprotein. Two mutations, H148V and Y145W, reduced the fluorescence emission by 82 and 98%, respectively. Ganesan et al. chose the Y145W mutant and the Y145W/H148V double mutant as FRET acceptors, and named them REACh1 and REACh2, respectively. '''Both REACh1 and REACh2 act as dark quenchers of GFP'''. | To optimize the spectral overlap of this FRET pair, the group obtained '''a genetically modified YFP acceptor'''. Mutations of amino acid residues that stabilize the excited state of the chromophore in enhanced YFP (EYFP) resulted in a non-fluorescent chromoprotein. Two mutations, H148V and Y145W, reduced the fluorescence emission by 82 and 98%, respectively. Ganesan et al. chose the Y145W mutant and the Y145W/H148V double mutant as FRET acceptors, and named them REACh1 and REACh2, respectively. '''Both REACh1 and REACh2 act as dark quenchers of GFP'''. | ||
| + | |||
{{Team:Aachen/BlockSeparator}} | {{Team:Aachen/BlockSeparator}} | ||
| Line 148: | Line 149: | ||
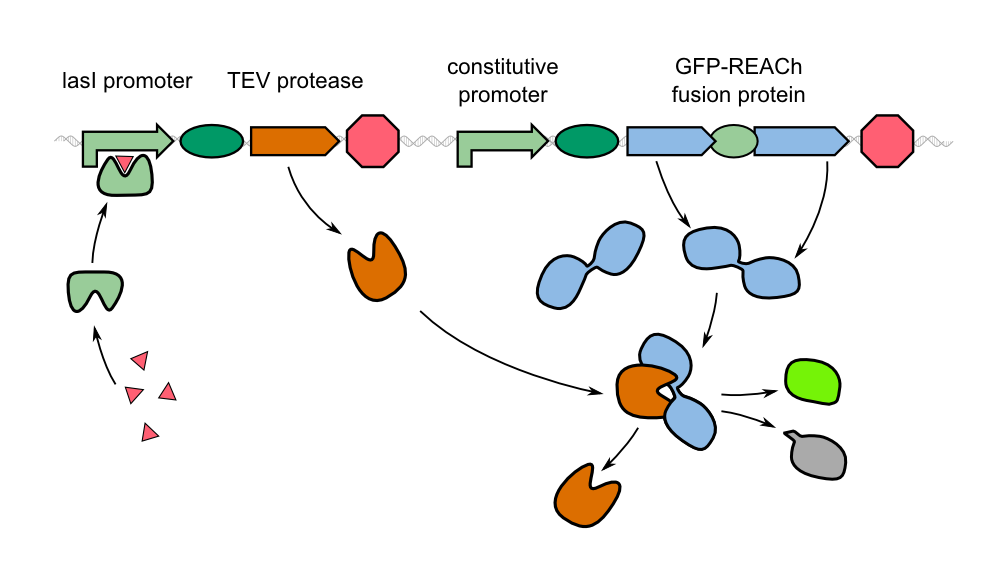
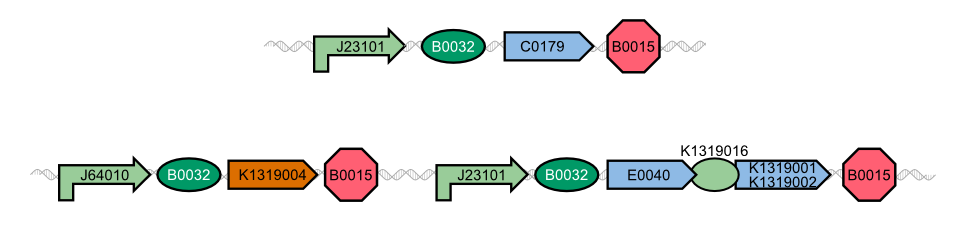
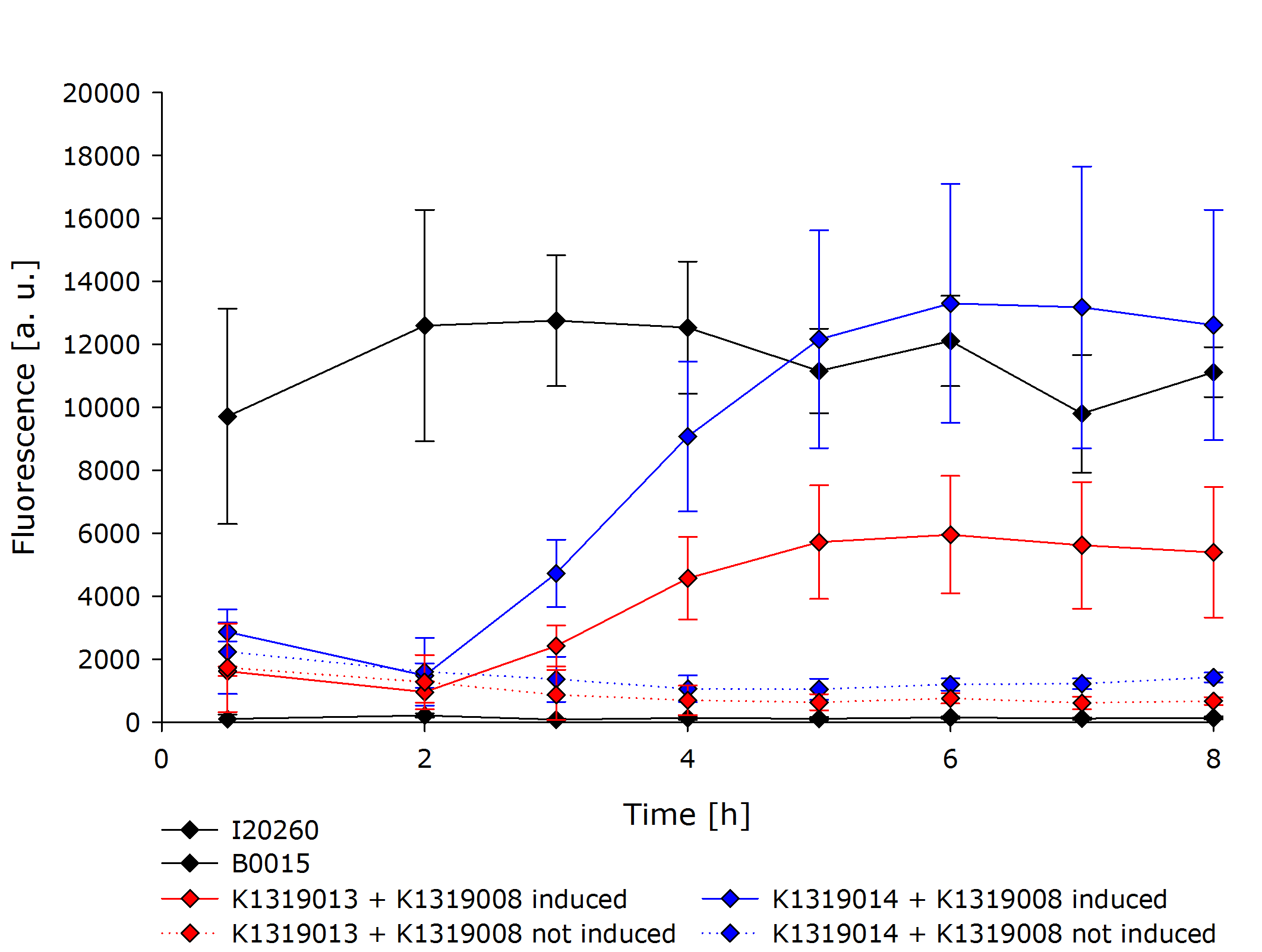
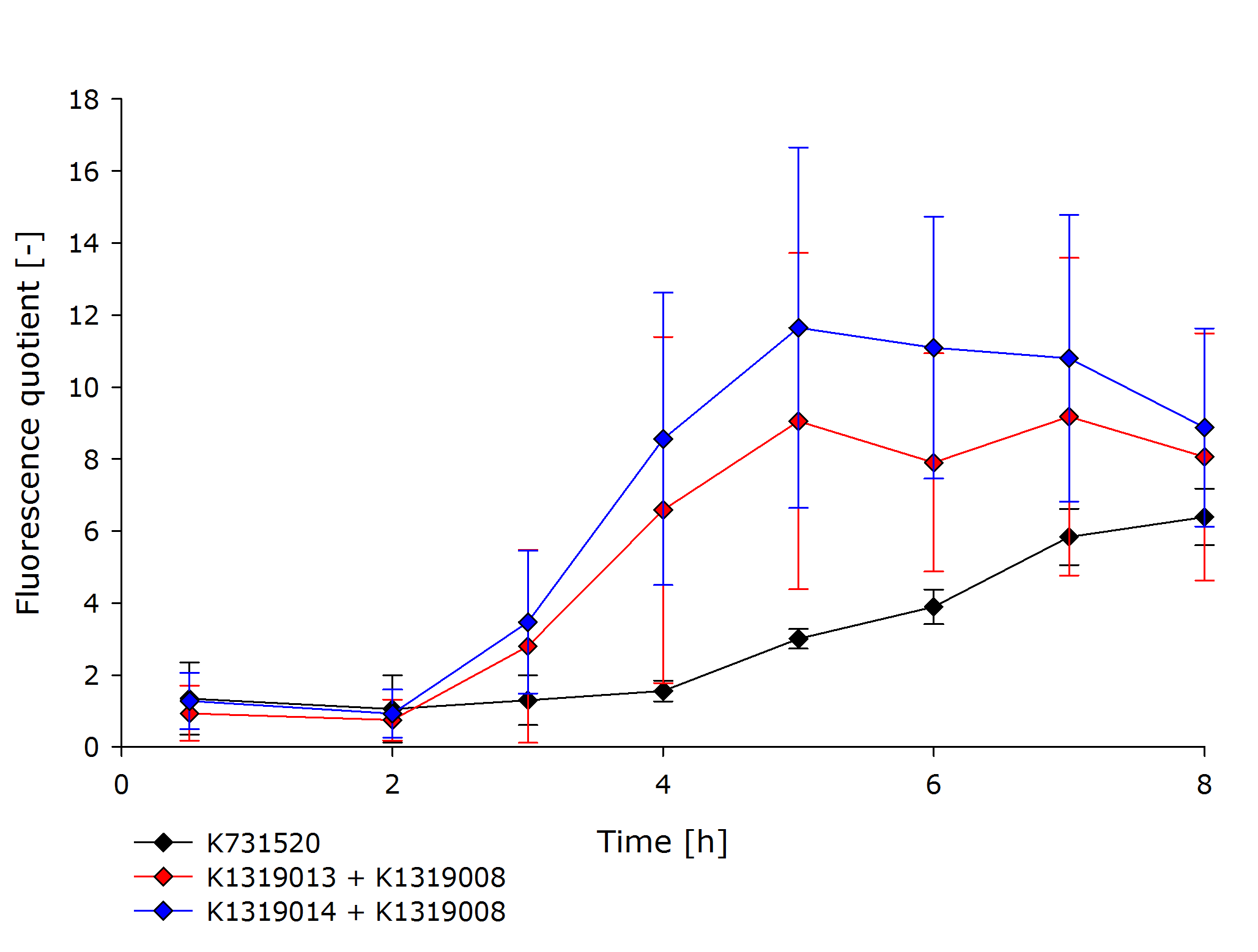
The resulting fusion proteins were labelled as [http://parts.igem.org/Part:BBa_K1319013 K1319013] (GFP fused with REACh1) and [http://parts.igem.org/Part:BBa_K1319014 K1319014] (GFP fused with REACh 2) and the linker between the proteins is labelled as [http://parts.igem.org/Part:BBa_K1319013 K1319016]. | The resulting fusion proteins were labelled as [http://parts.igem.org/Part:BBa_K1319013 K1319013] (GFP fused with REACh1) and [http://parts.igem.org/Part:BBa_K1319014 K1319014] (GFP fused with REACh 2) and the linker between the proteins is labelled as [http://parts.igem.org/Part:BBa_K1319013 K1319016]. | ||
| + | |||
{{Team:Aachen/BlockSeparator}} | {{Team:Aachen/BlockSeparator}} | ||
| Line 161: | Line 163: | ||
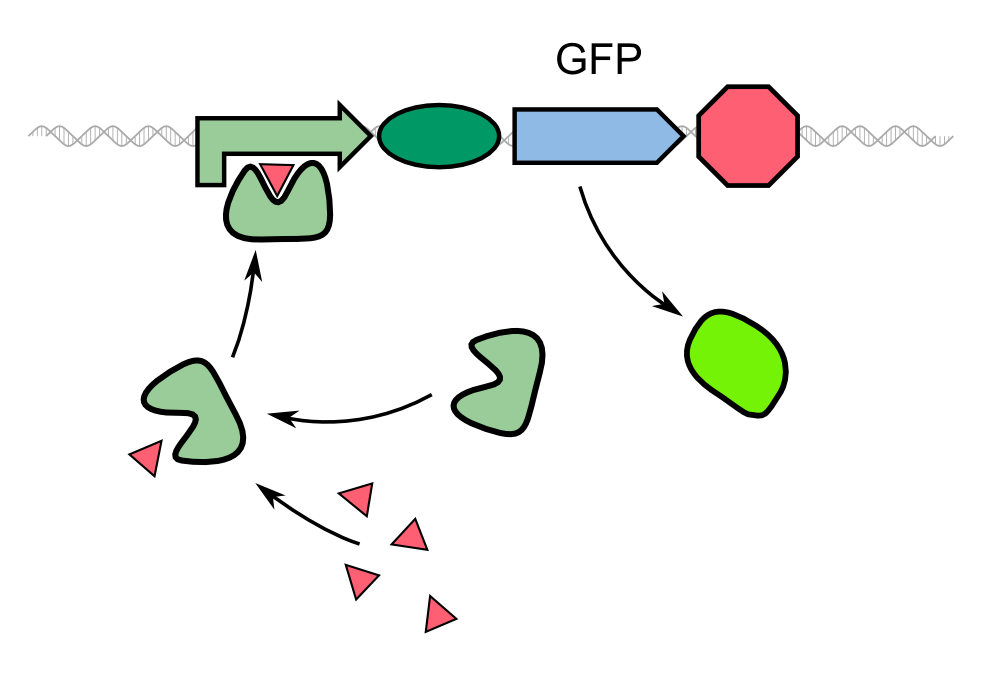
Though quite popular in molecular biology, the TEV protease is not avaiable as a BioBrick yet. Hence, the Aachen team introduces a protease with anti-self cleavage mutation S219V and codon optimized for ''E. coli'' [http://parts.igem.org/Part:BBa_K1319004 '''to the Parts Registry this year.'''] | Though quite popular in molecular biology, the TEV protease is not avaiable as a BioBrick yet. Hence, the Aachen team introduces a protease with anti-self cleavage mutation S219V and codon optimized for ''E. coli'' [http://parts.igem.org/Part:BBa_K1319004 '''to the Parts Registry this year.'''] | ||
| + | |||
{{Team:Aachen/BlockSeparator}} | {{Team:Aachen/BlockSeparator}} | ||
| Line 228: | Line 231: | ||
</div> | </div> | ||
</center> | </center> | ||
| + | |||
{{Team:Aachen/BlockSeparator}} | {{Team:Aachen/BlockSeparator}} | ||
| Line 241: | Line 245: | ||
Also finding and testing different promoters to induce the TEV protease is planned to be able to detect not only ''Pseudomonas aeruginosa'' but also other pathogens or other relevant molecules in general so that we can establish a concept for faster recognition for a variety of uses. | Also finding and testing different promoters to induce the TEV protease is planned to be able to detect not only ''Pseudomonas aeruginosa'' but also other pathogens or other relevant molecules in general so that we can establish a concept for faster recognition for a variety of uses. | ||
| + | |||
{{Team:Aachen/BlockSeparator}} | {{Team:Aachen/BlockSeparator}} | ||
| Line 263: | Line 268: | ||
'''POV-Ray''' | '''POV-Ray''' | ||
* Persistence of Vision Pty. Ltd. (2004) Persistence of Vision Raytracer (Version 3.7) [Computer software]. Retrieved from http://www.povray.org/download/ | * Persistence of Vision Pty. Ltd. (2004) Persistence of Vision Raytracer (Version 3.7) [Computer software]. Retrieved from http://www.povray.org/download/ | ||
| + | |||
{{Team:Aachen/Footer}} | {{Team:Aachen/Footer}} | ||
Revision as of 14:44, 17 October 2014
|
|
|
|
|
|
|
|
|
 "
"