Team:Aachen/Project/2D Biosensor
From 2014.igem.org
(→Principle of Operation) |
(→Principle of Operation) |
||
| Line 69: | Line 69: | ||
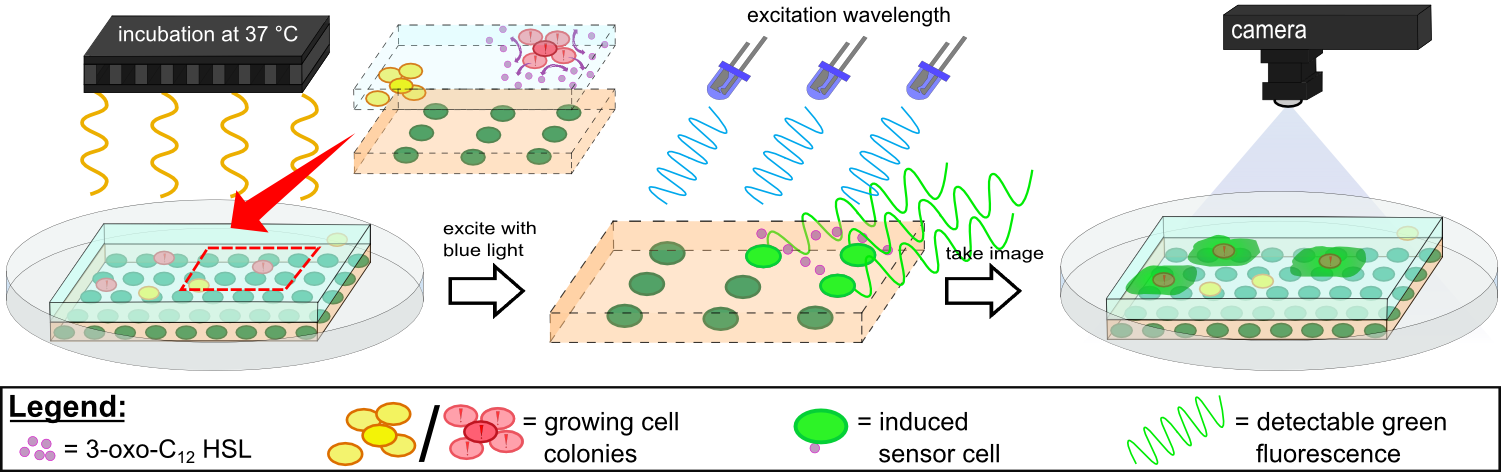
{{Team:Aachen/Figure|Aachen 14-10-14 Flowsheet OD-device part2 ipo.png|title=Mode of action inside WatsOn.|width=1000px}} | {{Team:Aachen/Figure|Aachen 14-10-14 Flowsheet OD-device part2 ipo.png|title=Mode of action inside WatsOn.|width=1000px}} | ||
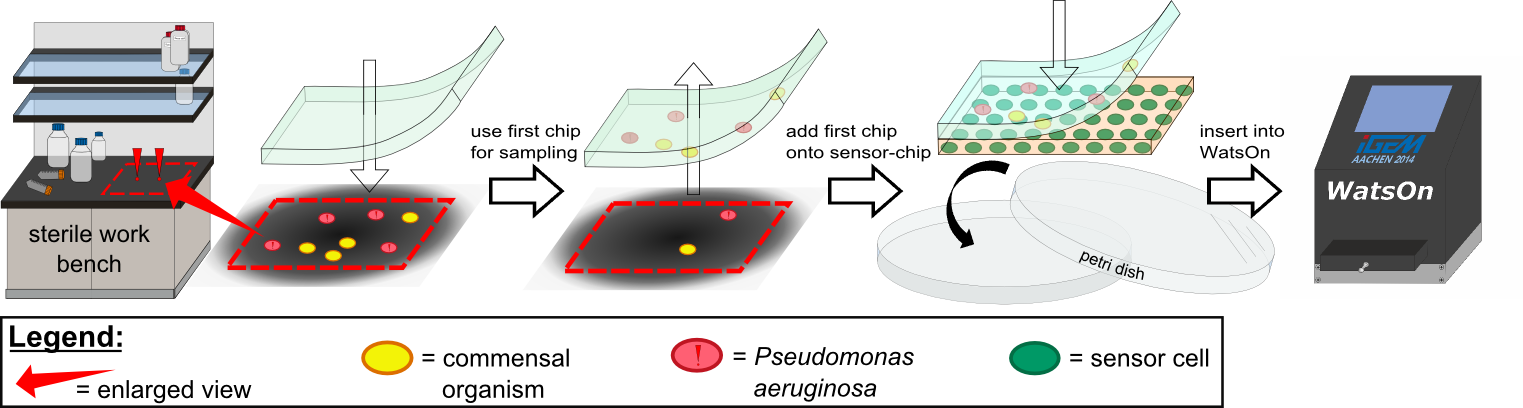
| - | Inside ''WatsOn'' the chips are incubated at 37 °C and populations of microorganisms on the sampling | + | Inside ''WatsOn'' the chips are incubated at 37 °C and populations of microorganisms on the sampling chip start to multiply. ''P. aeruginosa'' secrets an increasing number of 3-oxo-C{{sub|12}} homoserine lactones (HSLs) when multiplying. These HSLs induce the generation of a fluorescent signal in our sensor cells, which is described in more detail in the [https://2014.igem.org/Team:Aachen/Project/FRET_Reporter REACh Construct] section. |
The chips can be illuminated with blue light at any time, while ''WatsOn'' takes a picture of the chip. The software ''Measurarty'' then analyzes any fluorescent signal. Depending on the intensity of the signal and the size of the spot, ''Cellock Holmes'' can '''calculate concentration and distribution of ''P. aeruginosa'' ''' on the sampled surface. | The chips can be illuminated with blue light at any time, while ''WatsOn'' takes a picture of the chip. The software ''Measurarty'' then analyzes any fluorescent signal. Depending on the intensity of the signal and the size of the spot, ''Cellock Holmes'' can '''calculate concentration and distribution of ''P. aeruginosa'' ''' on the sampled surface. | ||
Revision as of 15:56, 15 October 2014
|
|
|
 "
"