Team:Aachen/Project/2D Biosensor
From 2014.igem.org
(→Principle of Operation) |
(→Principle of Operation) |
||
| Line 55: | Line 55: | ||
<span class="anchor" id="biosensorpoo"></span> | <span class="anchor" id="biosensorpoo"></span> | ||
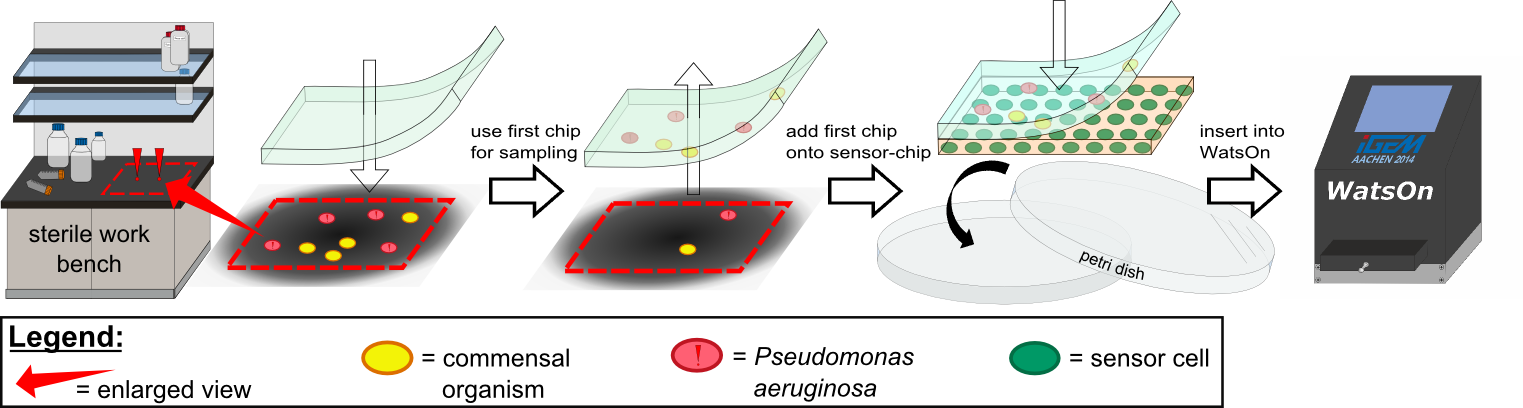
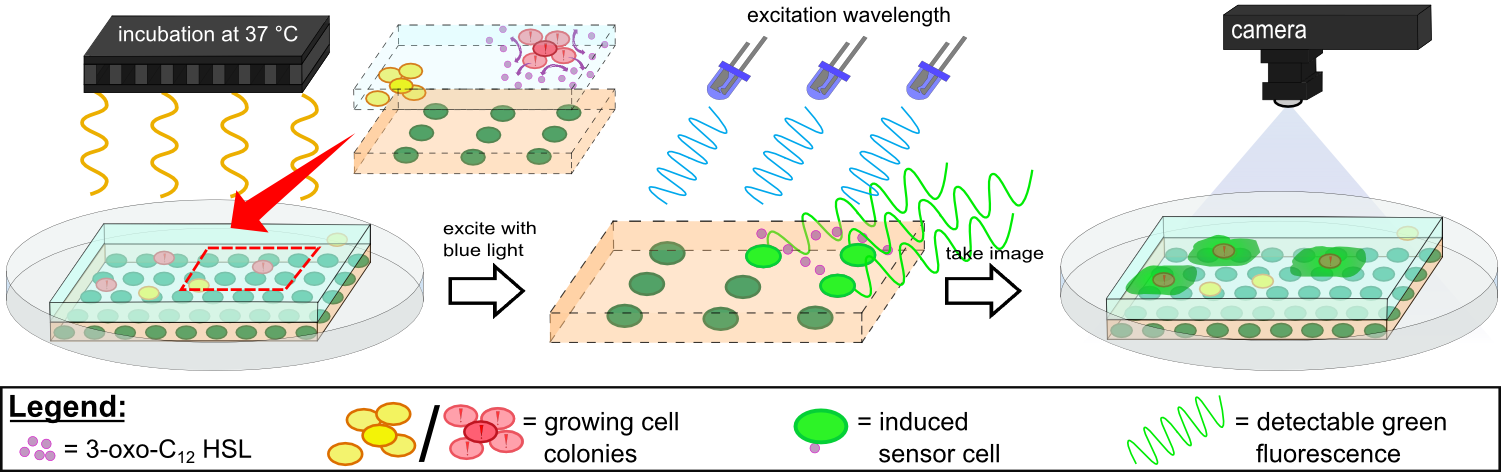
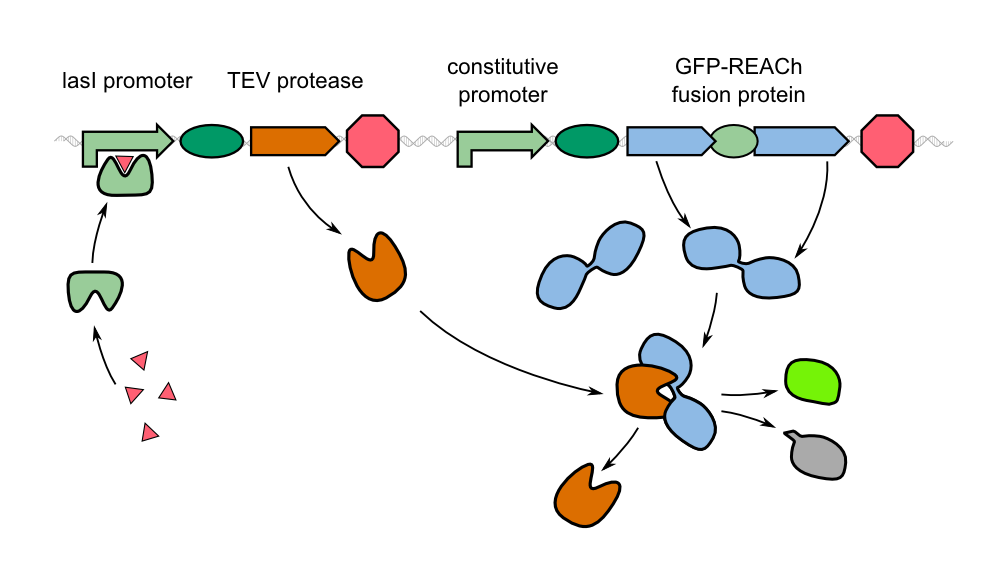
| - | ''Cellock Holmes'' is devised based upon a SynBio approach comprised of a '''two-dimensional biosensor and a measurement device'''. The two-dimensional biosensor is designed to recognize quorum sensing molecules secreted by the pathogen cells and to generate a distinct fluorescence signal, while the measurement device is designed to recognize and analyse the produced signal. In general different detection mechanisms could be incorporated into the sensing cells, e.g. our fluorescence based [https://2014.igem.org/Team:Aachen/Project/FRET_Reporter REACh construct] or our [https://2014.igem.org/Team:Aachen/Project/Gal3 Galectin-3 construct]. | + | ''Cellock Holmes'' is devised based upon a SynBio approach comprised of a '''two-dimensional biosensor and a measurement device'''. The two-dimensional biosensor is designed to recognize quorum sensing molecules secreted by the pathogen cells and to generate a distinct fluorescence signal, while the measurement device is designed to recognize and analyse the produced signal. To overcome the limitation of our fluorescence based [https://2014.igem.org/Team:Aachen/Project/FRET_Reporter REACh construct] to bacteria that secrete autoinducers, we developed an [https://2014.igem.org/Team:Aachen/Project/Gal3 Galectin-3 construct] alternative detection method, which is based on tagging cells with a fluorescent reporter. |
| + | |||
| + | In general different detection mechanisms could be incorporated into the sensing cells, e.g. our fluorescence based [https://2014.igem.org/Team:Aachen/Project/FRET_Reporter REACh construct] or our [https://2014.igem.org/Team:Aachen/Project/Gal3 Galectin-3 construct]. | ||
[graph quorum senising] | [graph quorum senising] | ||
Revision as of 11:20, 15 October 2014
|
|
|
|
 "
"