Team:Aachen/Project/2D Biosensor
From 2014.igem.org
(→Principle of Operation) |
(→Principle of Operation) |
||
| Line 57: | Line 57: | ||
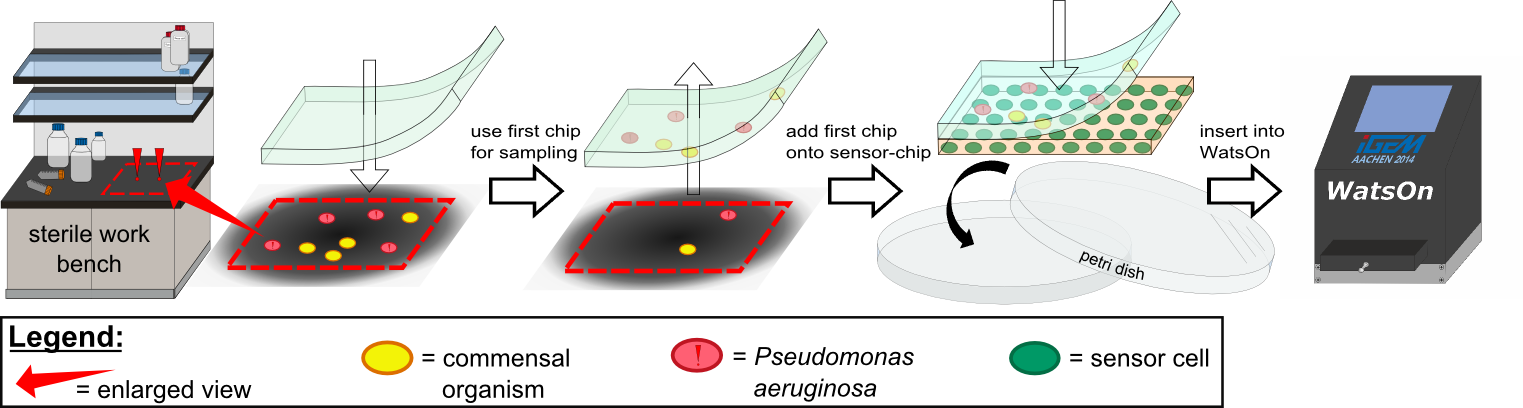
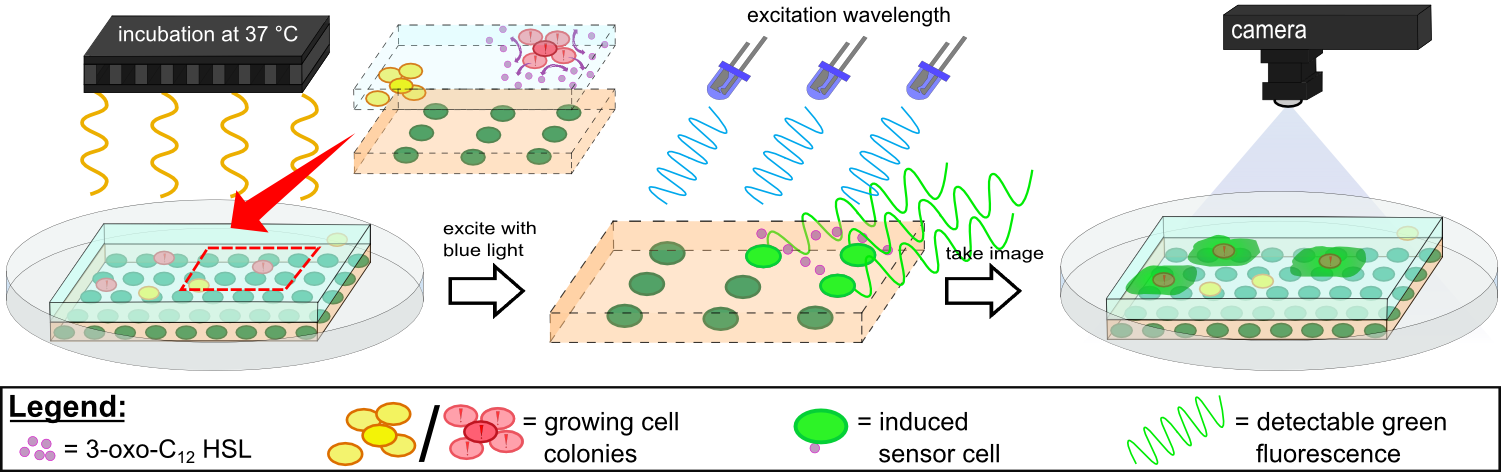
''Cellock Holmes'' is devised based upon a SynBio approach comprised of a '''two-dimensional biosensor and a measurement device'''. The two-dimensional biosensor is designed to recognize quorum sensing molecules secreted by the pathogen cells and to generate a distinct fluorescence signal, while the measurement device is designed to recognize and analyse the produced signal. | ''Cellock Holmes'' is devised based upon a SynBio approach comprised of a '''two-dimensional biosensor and a measurement device'''. The two-dimensional biosensor is designed to recognize quorum sensing molecules secreted by the pathogen cells and to generate a distinct fluorescence signal, while the measurement device is designed to recognize and analyse the produced signal. | ||
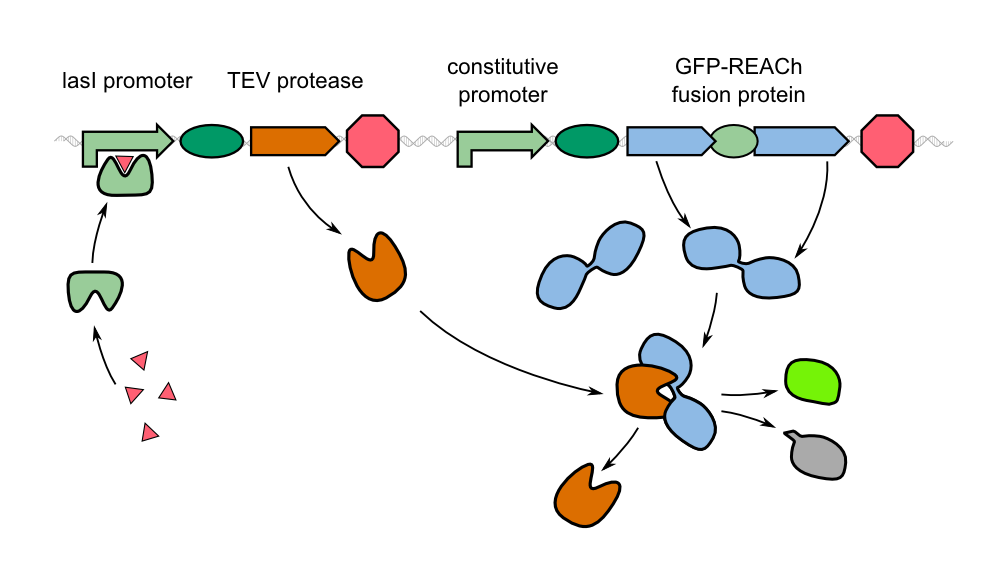
| - | Our '''sensor cells are immobilized in agar chips'''. To make the chips, we mix the sensing cells, also known as ''Cellocks'', with liquid LB agar. Different detection mechanisms can be | + | Our '''sensor cells are immobilized in agar chips'''. To make the chips, we mix the sensing cells, also known as ''Cellocks'', with liquid LB agar. Different detection mechanisms can be incorporated into the sensing cells, e.g. a detection mechanism based on a fluorescent response to autoinducers, which is explanined below in this section. |
In the course of our project, we designed a casting mold specifically for the production of our agar chips. When the agar has cooled down, the chips are cut out of the mold and ready to use . If you want to know exactly how our chips are manufactures, you can read up on more details in our [https://2014.igem.org/Team:Aachen/Notebook/Protocols/detection Protocols]. | In the course of our project, we designed a casting mold specifically for the production of our agar chips. When the agar has cooled down, the chips are cut out of the mold and ready to use . If you want to know exactly how our chips are manufactures, you can read up on more details in our [https://2014.igem.org/Team:Aachen/Notebook/Protocols/detection Protocols]. | ||
<center> | <center> | ||
Revision as of 10:46, 15 October 2014
|
|
|
|
 "
"