Team:Waterloo/Math Book
From 2014.igem.org
(Difference between revisions)
(sRNA moved out) |
|||
| Line 23: | Line 23: | ||
</ul> | </ul> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<!------------------- CONJUGATION SECTION ---------------------------------> | <!------------------- CONJUGATION SECTION ---------------------------------> | ||
Revision as of 00:02, 18 October 2014

Math Book
Our Math Book is meant to be the mathematical modelling equivelent of a lab book, where we store everything another team might need to recreate our models. You can access code related to the models can be accessed from this github page.
We modelled three main aspects of the Staphylocide system: CRISPR Interference, silencing RNA and Conjugation.
We hope you enjoy learning more about our model on the subpages!
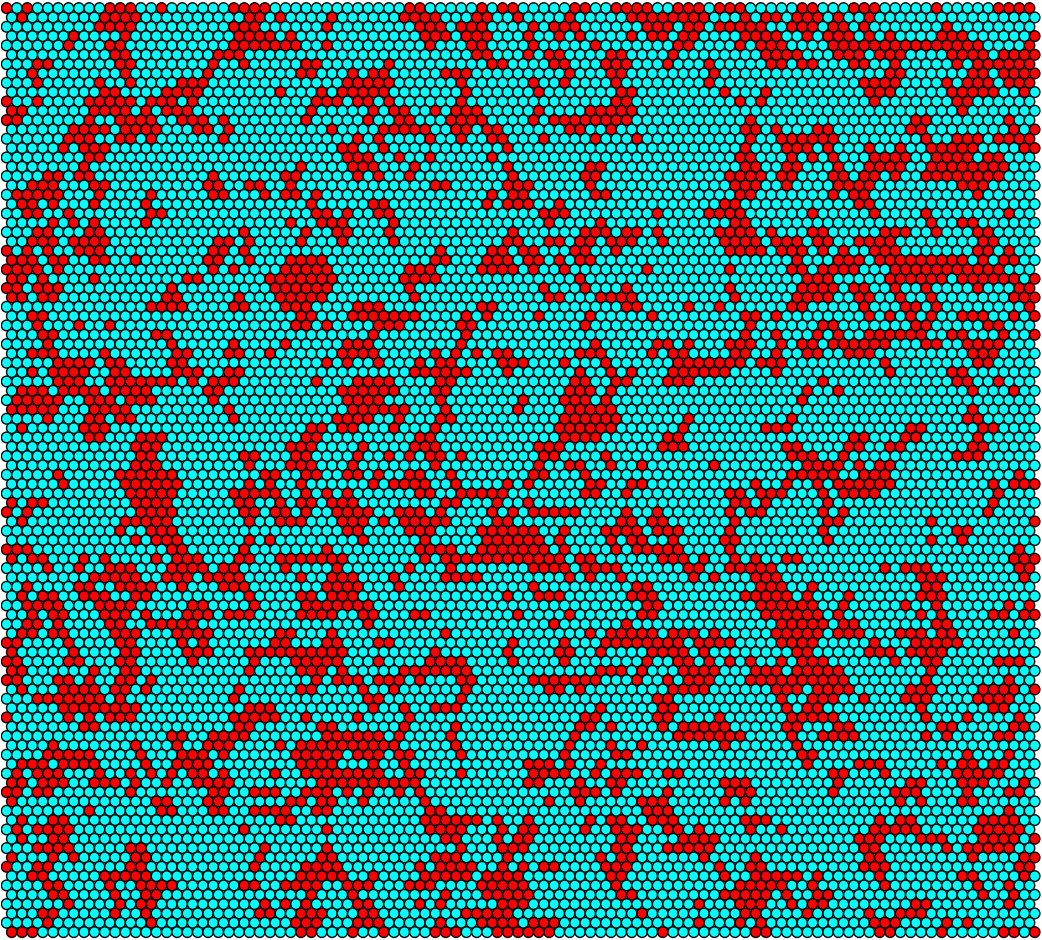
Conjugation
ABM






References
| [1]D. Bikard et al. “Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system”. In: Nucleic Acids Res. 41.15 (Aug. 2013), pp. 7429–7437. |
| [2]Florian Brandt et al. “The Native 3D Organization of Bacterial Polysomes”. In: Cell 136.2 (2009), pp. 261 –271. issn: 0092-8674. doi: 10.1016/j.cell.2008.11.016. |
| [3]A. G. Cheng, D. Missiakas, and O. Schneewind. “The giant protein Ebh is a determinant of Staphylococcus aureus cell size and complement resistance”. In: J. Bacteriol. 196.5 (2014), pp. 971–981. |
| [4]A. L. Cheung, K. Nishina, and A. C. Manna. “SarA of Staphylococcus aureus binds to the sarA promoter to regulate gene expression”. In: J. Bacteriol. 190.6 (Mar. 2008), pp. 2239–2243. |
| [5]G. Domingue, J. W. Costerton, and M. R. Brown. “Bacterial doubling time modulates the effects of opsonisation and available iron upon interactions between Staphylococcus aureus and human neutrophils”. In: FEMS Immunol. Med. Microbiol. 16.3-4 (Dec. 1996), pp. 223–228. |
| [6]S. Michalik et al. “Life and death of proteins: a case study of glucose-starved Staphylococcus aureus”. In: Mol. Cell Proteomics 11.9 (Sept. 2012), pp. 558–570. |
| [7]R. Milo et al. “BioNumbers-the database of key numbers in molecular and cell biology”. In: Nucleic Acids Res. 30 (Jan. 2010), pp. D750–D753. url: http://bionumbers.hms.harvard.edu/bionumber.aspx?id=107869}. |
| [8]L. S. Qi et al. “Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression”. In: Cell 152.5 (Feb. 2013), pp. 1173–1183. |
| [9]C. Roberts et al. “Characterizing the effect of the Staphylococcus aureus virulence factor regulator, SarA, on log-phase mRNA half-lives”. In: J. Bacteriol. 188.7 (Apr. 2006), pp. 2593–2603. doi: 10.1128/JB.188.7.2593-2603.2006 |
| [10]Marlena Siwiak and Piotr Zielenkiewicz. “Transimulation - Protein Biosynthesis Web Service”. In: PLoS ONE 8.9 (Sept. 2013), e73943. doi: 10.1371/journal.pone.0073943. |
| [11]S.H. Sternberg et al. “DNA interrogation by the CRISPR RNA-guided endonuclease Cas9”. In: Nature 7490 (2014), 6267. doi: 10.1038/nature13011. url: http://www.nature.com/nature/journal/v507/n7490/full/nature13011.html. |
| [12]Freiburg iGEM Team. dCas9. BBa K1150000 Standard Biological Part. 2013. url: http://parts.igem.org/Part:BBa_K1150000. |
| [13]UCSF iGEM Team. Operation CRISPR: Decision Making Circuit Model. 2013. url: https://2013.igem.org/Team:UCSF/Modeling. |
| [14]Jian-Qiu Wu and Thomas D. Pollard. “Counting Cytokinesis Proteins Globally and Locally in Fission Yeast”. In: Science 310.5746 (2005), pp. 310–314. doi: 10.1126/science.1113230. |
| [15]Jianfang Jia and Hong Yue. “Sensitivity Analysis and Parameter Estimation of Signal Transduction Pathways Model”. In: Proceedings of the 7th Asian Control Conference (Aug. 2009), pp. 1357–1362. |
| [16]Fi-John Chang and J. W. Delleur. “Systematic Parameter Estimation Of Watershed Acidification Model”. In: Hydrological Processes 6. (1992), pp. 29–44. doi: 10.1002/hyp.3360060104. |
| [17]Aiba, H. (2007). Mechanism of RNA silencing by Hfq-binding small RNAs. Current opinion in microbiology, 10 (2), 134-139. |
| [18]Horstmann, N., Orans, J., Valentin-Hansen, P., Shelburne, S. A., & Brennan, R. G. (2012). Structural mechanism of Staphylococcus aureus Hfq binding to an RNA A-tract. Nucleic acids research, gks809. |
| [19]Eyraud, A., Tattevin, P., Chabelskaya, S., & Felden, B. (2014). A small RNA controls a protein regulator involved in antibiotic resistance in Staphylococcus aureus. Nucleic acids research, gku149. |
| [20]Shimoni, Y., Friedlander, G., Hetzroni, G., Niv, G., Altuvia, S., Biham, O., & Margalit, H. (2007). Regulation of gene expression by small non‐coding RNAs: a quantitative view. Molecular Systems Biology, 3 (1) |
| [21]Fender, A., Elf, J., Hampel, K., Zimmermann, B., & Wagner, E. G. H. (2010). RNAs actively cycle on the Sm-like protein Hfq. Genes & Development, 24 (23),2621-2626. |
| [22] Swain, P. S. (2004). Efficient attenuation of stochasticity in gene expression through post-transcriptional control. Journal of molecular biology, 344 (4),965-976. |
| [23] Hussein, R., & Lim, H. N. (2012). Direct comparison of small RNA and transcription factor signaling. Nucleic acids research, 40 (15), 7269-7279. |
| [24] Levin, B.R., Stewart, F.M. and Rice, V.A. 1979. “The Kinetics of Conjugative Plasmid Transmission: Fit of a Simple Mass Action Model.” In: Plasmid. 2. pp. 247-260. |
| [25]Projan, S.J. and Archer, G.L. 1989. “Mobilization of the Relaxable Staphylococcus aureus Plasmid pC221 by the Conjugative Plasmid pGO1 Involves Three pC221 Loci.” In: Journal of Bacteriology. pp. 1841-1845. |
| [26]Phornphisutthimas, S., Thamchaipenet, A., and Panijpan, B. 2007. “Conjugation in Escherichia coli.” In: The International Union of Biochemistry and Molecular Biology. 35. 6. pp. 440-445. |
| [27]Phornphisutthimas, S., Thamchaipenet, A., and Panijpan, B. 2007. “Conjugation in Escherichia coli.” In: The International Union of Biochemistry and Molecular Biology. 35. 6. pp. 440-445. |
| [28]P Chung P., McNamara P.J., Campion J.J., Evans M.E. 2006. “Mechanism-based pharmacodynamic models of fluoroquinolone resistance in Staphylococcus aureus.” In: In: Antimicrobial Agents Chemotherapy. 50. pp. 2957-2965. |
| [29] Chang H., Wang L. “A Simple Proof of Thue's Theorem on Circle Packing” In: arXiv:1009.4322v1. |
 "
"




