Team:Nagahama project
From 2014.igem.org
| ||||||||||||||
Contents |
Our Project
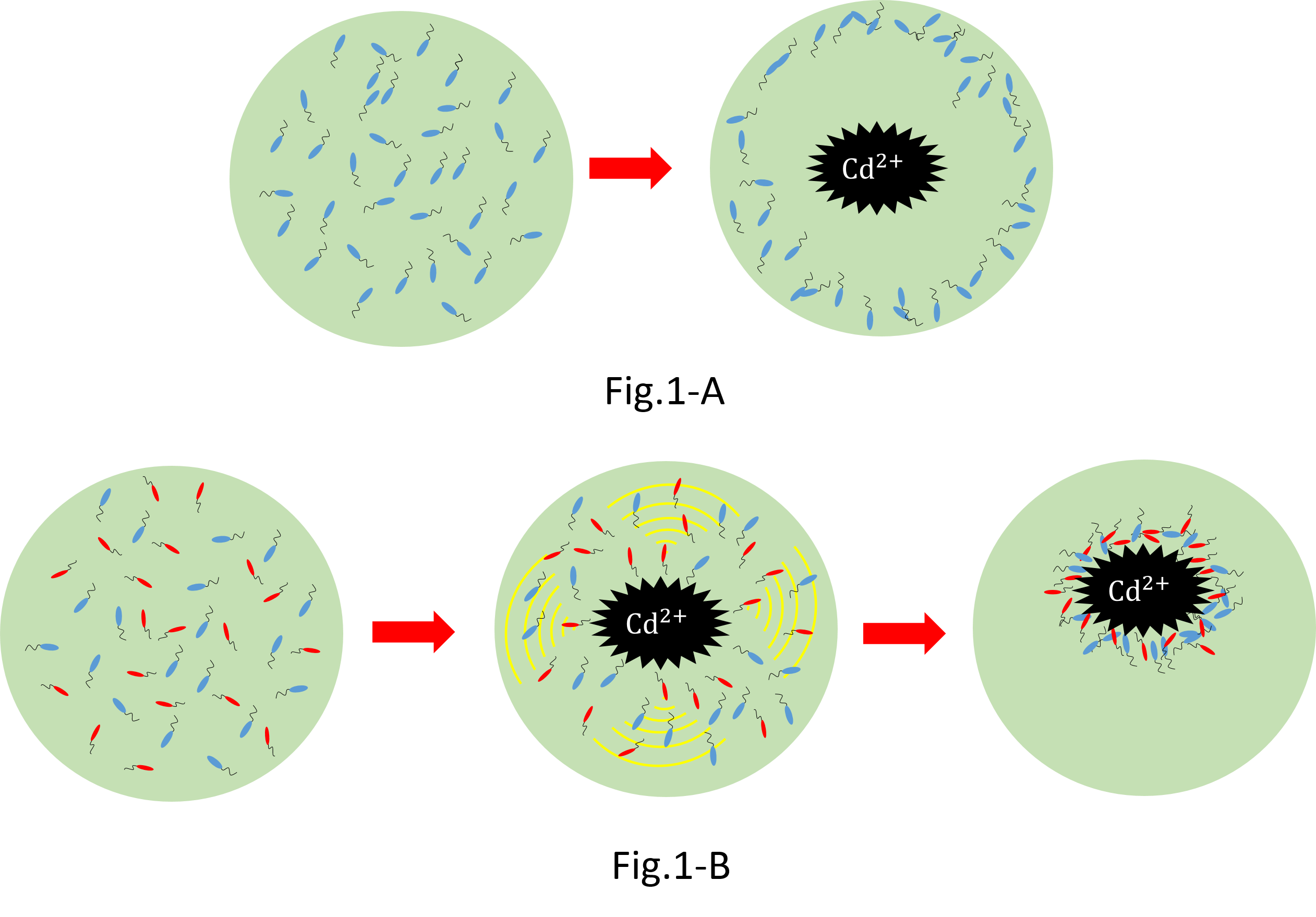
We make various systems by interaction of cell-cell communication. We keep one function in one E. coli. This means to make simple plasmid. The following is one example. We’d like to collect cadmium in water. Therefore we use two kinds of E.coli. One catches Cadmium. The other attracts all E.coli by using chemoattractant. Catches E.coli displays metallothionein a protein combines a heavy metal. Cadmium is a kind of heavy metal. The other synthesizes aspartic acid (Asp) one kind of chemoattractant. All E.coli gather in the E. coli synthesizes Asp. To use these E.coli, finally cadmium will be caught.
When we collect cadmium in the water, if use only catching E. coli(Blue),they go away from cadmium. Because they have negative chemotaxisin contrast with cadmium. Consequently, they don't catch cadmium very much.[fig1-A] In constract, if we use two kind of recombinant Escherichia coli(E. coli);gathering E. coli(Red) and catching E. coli(Blue), we can build the creature system which collects cadmium while sitting at cadmium by using two kinds of Escherichia coli.[fig1-B]
It is this theoretical merit that we can suggest two kinds of possibility that meeting condition and collection quantity of Escherichia coli are changed by changing the ratio of Escherichia coli in addition to cadmium solution.
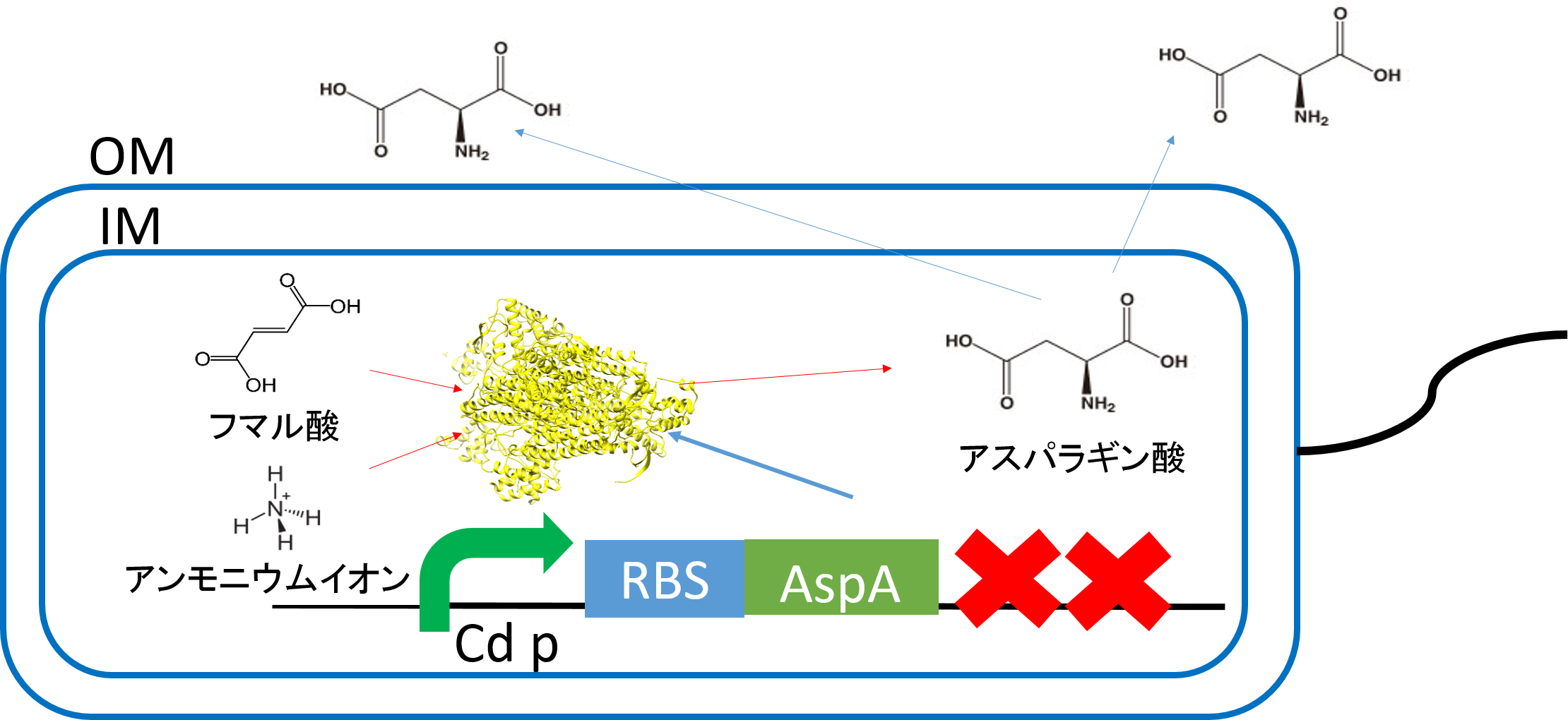
The function of our parts
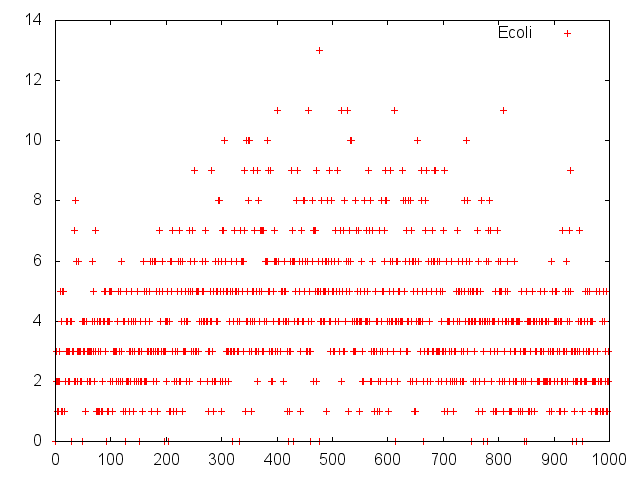
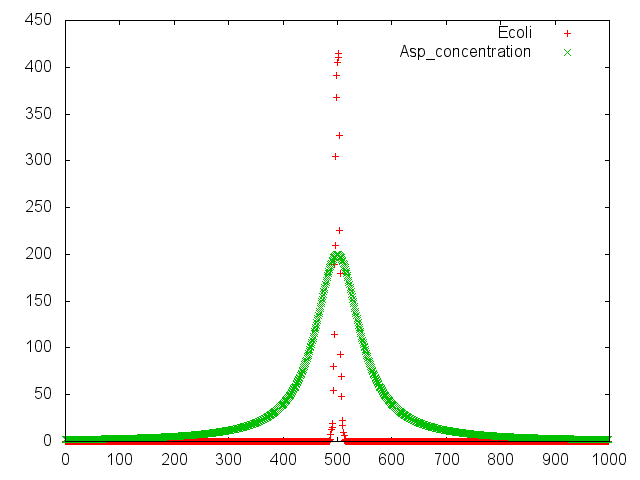
Modeling
We consigned modeling of our project to
UT-Tokyo.They readily compiled with our requests. Their model is ideal.We really appreciate their jobs We wish a friendly relationship with UT-Tokyo.Thank you so much!! The detail of their jobs are lists below!!
Method
Medium
Trypton Broth
Trypton 10g/L
NaCl 10g/L
H2O 1L
Wash medium
Potassium phosphate buffer (pH 7.0), 10⁻²M
MgSO₄, 10⁻³M
potassium ethylenediaminetetraacetate (EDTA), 10⁻⁴M
Chemotaxis medium
Potassium phosphate buffer (pH 7.0), 10⁻²M
potassium EDTA, 10⁻⁴M
L-methionine 10⁻⁶M
Synthesis medium
Sodium hydrogen fumarate 46g/L
Ammonium chloride 17.8g/L
Magnesium sulfate 7 hydrate 0.25g/L
H₂O 1L
Ph8.5with sodium hydrogen
LB medium
Tryptone 10g/L
Yeast extract 5g/L
NaCl 10g/L
(agar 15g/L)
H₂O 1L
2×YT medium
Tripton 16g/L
Yeast extract 10g/L
NaCl 5g/L
(agar 15g/L)
H₂O 1L
M9 swarming Agar
M9 salt
1.25 % (v/v) glycerol
0.3 % (w/v) agar
0.1 % (w/v) CaCl2 solution (after autoclave)
0.1 % (w/v) MgSO4 solution (after autoclave)
0.03 % (w/v) thiamine (after autoclave)
5 × M9 stock solution
6.4 % (w/v) Na2HPO4
1.5 % (w/v) KH2PO4
0.25 % (w/v) NaCl
0.5 % (w/v) NH4Cl
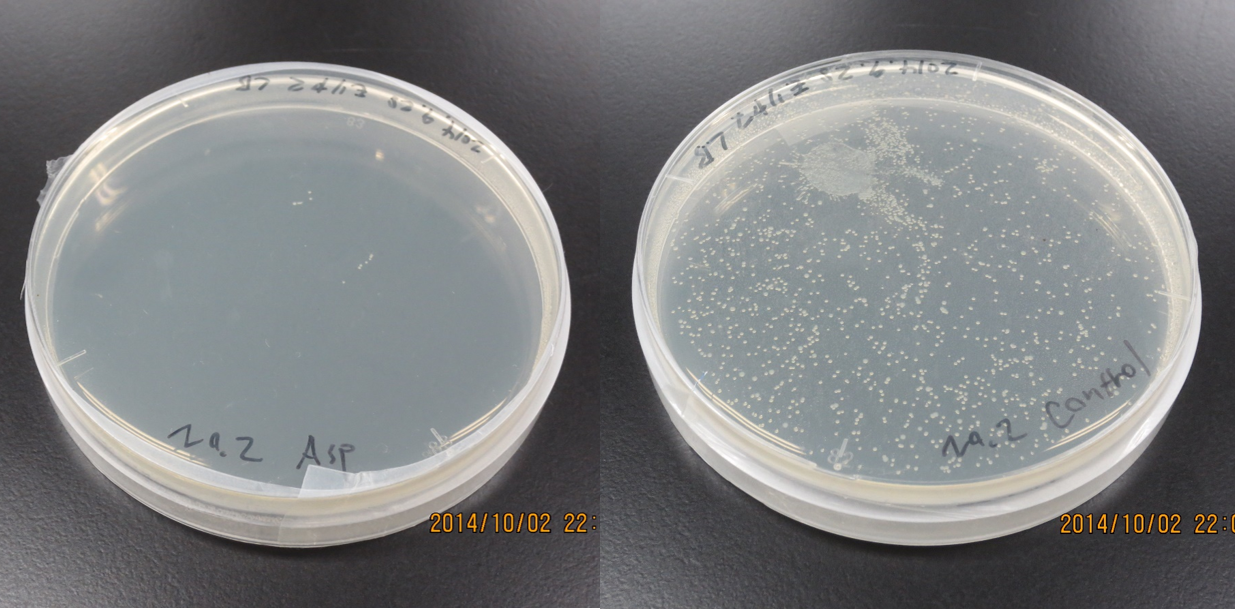
Aspartate synthesis
E.coli K12 transformed with CdP-R.B.S-AspA-d.Ter (BBa_K1342001) previous cultured with cadmium in LB medium (250μM) in 37℃ for 12hr at 120rpm. Adjust Cell mass (OD1.0) and therefor centrifuged 4000rpm for 20 min. Cell pellets ware activated in synthesis medium in 37℃ for 2hr at 120rpm/min.
SDS PAGE
・Preculture E.coli holding a plasmid containing a target gene or nomal E.coli.
・Measure OD600 0.6-1.0
・the expression of fusion protein by isopropylthio-β-D-galactoside (IPTG) and Cd2+ soln.
・Transfer a sample a 200 µl in a microcentrifuge tube
・Centrifuge at 13,000rpm for a minute at 4℃
・Discard supernatant quantitative
・Store pellet at -20 °C
・Thaw pellet and resupend in Sample Buffer (100 µL 1xSample Buffer per samples)
・Heat for 5 minutes at 98 °C
・Centrifuge at 13,000rpm for 10 minutes at 4℃
・Transfer supernatant to a new microcentrifuge tube
・Analyze samples by SDS-PAGE.(Use 20 µL per samples)
TLC assay
We analyzed L-aspartate by thin-layer chromatography (TLC). The equal volume of the supernatant of synthesis medium was added with 7 mg/mL 5-Dimethylamino naphthalene-1-sulfonyl Chloride (dissolved in acetone) and was incubated for more than 30 min at room temperature. Two microliters of the reactant was spotted on a TLC silica plate, and was developed in a mixture of ethyl acetate, pyridine, water, and acetic acid(162:21:11:6 v/v).
Chemotaxis Assay
Introduction
Chemotaxis of E. coli against asparate was assayed by two methods. One was swarming assay, and the other was capillary assay. In swarming assay, we have used "sawaming plate", a solid culture plate containing lower concentration of agar, on that plate, E. coli can swim. We have used capillary containing aspartate. The capillary was set in chemotaxis medium containing motile E. coli. The number of E. coli attracted to asparate in capillary was determined by colony formation on LB plate.
Swarming Assay
E. coli JM109 was cultured at 30℃ for 12 hours with shaking (50 rpm).
We assayed chemotaxis of E. coli against Cadmium and aspartic acid on soft agar, containing M9 synthetic medium, on that plate E. coli can swim.
Protocol
1. Aliquot of the culture was spotted on the center of agar plate.
2. 10 mM L-Aspartic acid (40 μl) or 100 mM Cadmium chloride (4 μl) was spotted on 25 mm distant from the center of the agar plate.
3. The plate was standed for 5 min at RT.
4. The plate was incubated at 30℃.
Capillary Assay
Object
Assay the chemotaxis of E. coli against aspartarte with capillary.
Plotocol
1. Preculture E. coli in tripton broth at 30 ℃ for 12 hr with shaking (50 rpm).
2. Check the motility of E. coli by microscope (× 800).
3. Dilute the E. coli culture (200 μL) with tripton broth (20 mL).
4. Incubate at 30 ℃ with shaking (50 rpm) until log phase (OD600 = ~ 0.2).
5. Aliquote (500 μL) of the culture was centrifuged at 25℃ for 10 min at 3400 x G.
6. Dicard the supernatant.
7. Resuspend the pellets gently with wash medium (50 μL).
8. Repeat 5~7 step.
9. Resuspend the pellets gently with chemotaxis medium (500 μL).
10. Check the motility of E. coli by microscope (× 800).
11. Aliquote (100 μL) of E. coli suspended in chemotaxis medium was set into chamber apparatus.
12. Prepare asparate capillary and negative control capillary that contain 1 μL of chemotaxis medium with and without 10 mM aspartate, respectively.
13. Both capillaries (asparate capillary and negative control capillary) were set into chamber apparatus.
14. Incubate at 30 ℃ for 90 mim.
15. The chemotaxis medium in capillary was collected in 100 μL of fresh chemotaxis medium.
16. All chemotaxis medium containing attracted E. coli was plated on LB plate.
17. Incubate at 37 ℃ for 12 hr.
18. Count the number of colony.
Result & Discussion
TLC Assay

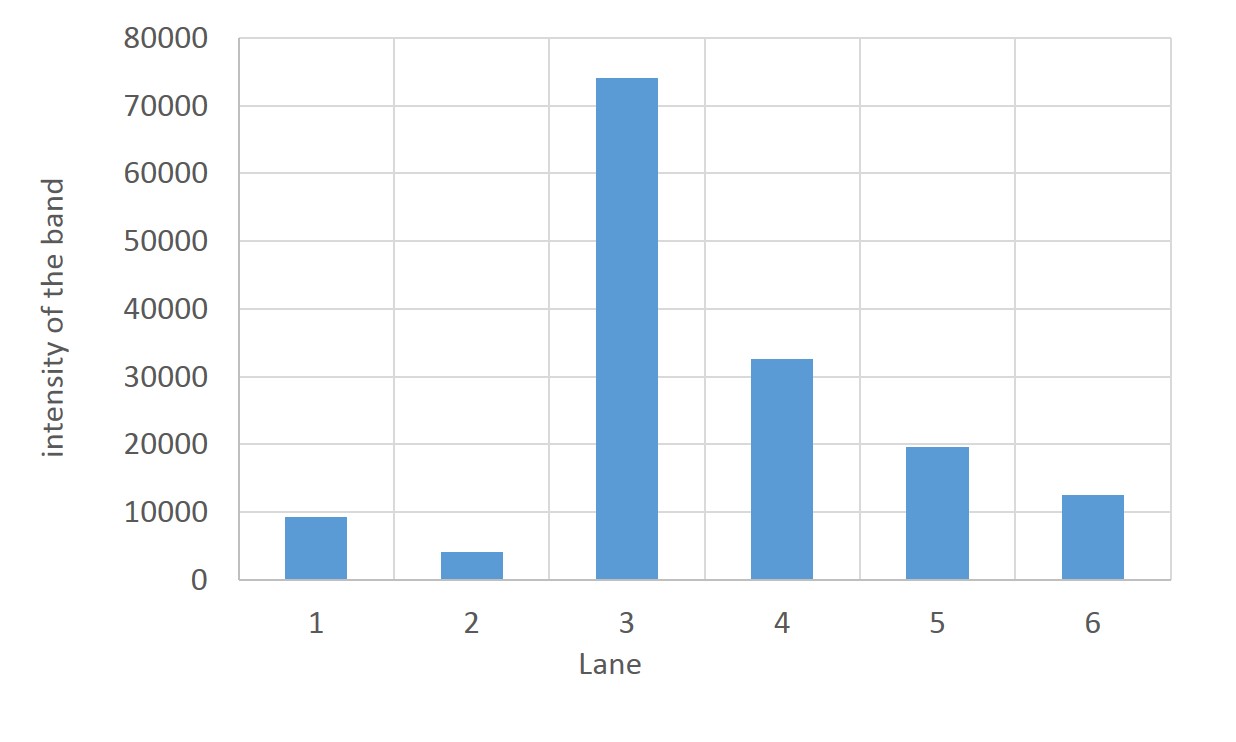
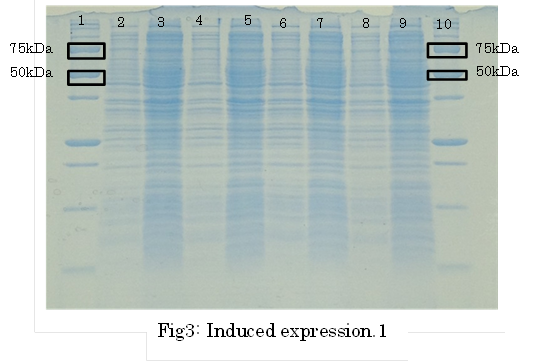
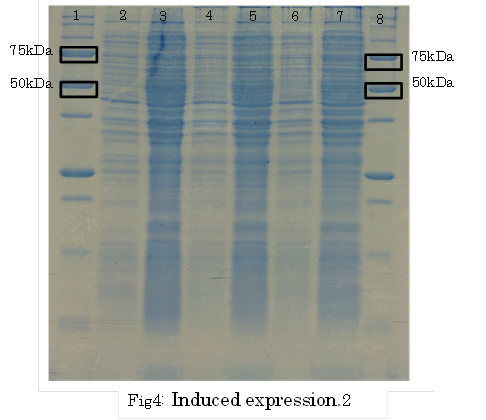
SDS PAGE
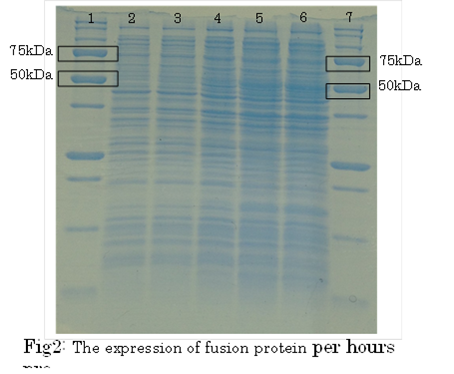 1. marker 2. 0hr 3. 0.5hr 4. 2hr 5. 6hr 6. 24hr 7. Marker
marker:Precision Plus Protein Standards (BIO-RAD)
1. marker 2. 0hr 3. 0.5hr 4. 2hr 5. 6hr 6. 24hr 7. Marker
marker:Precision Plus Protein Standards (BIO-RAD)
OD600=0.74
CdCl2 100μM/IPTG 1mM
Swarming Assay
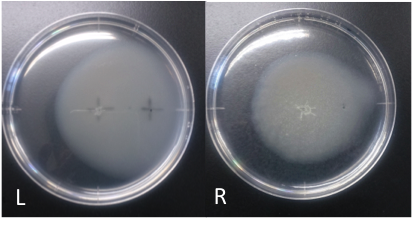
Incubation: 108 hours
E. coli cultured was spotted on the center of agar plate, and chemotaxis was spotted on 25 mm distant from the center.
The swarming circle of aspartic acid (L) was larger than that of ddH2O (R), and the swarming area around aspartate acid (L) was more larger than that of ddH2O (R), suggesting that E. coli swarming to aspartate acid.
This experiment indicate that E. coli may have positive chemotaxis for aspartic acid.
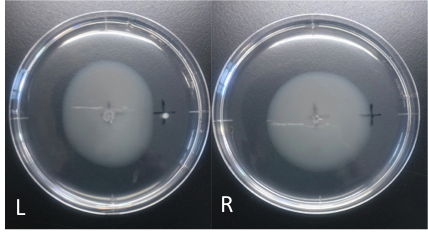
Culture time: 109 hours 50 minutes (4days and 13 hours 50 minutes)
It is 4.3cm from the edge of plate to the center
It is 2.5cm from the center of the plate to the place that I spotted.
Circle is smaller the medium which I spotted Cdim into than the medium which I spotted ddH20 into.
It is that E. coli swims against Cdim to understand from photographs of L and R.
In the side where I spotted Cd in, circle becomes dented.
I think that E. coli may have negative chemotaxis for Cd than these two (Cd and ddH2O) control experiment.
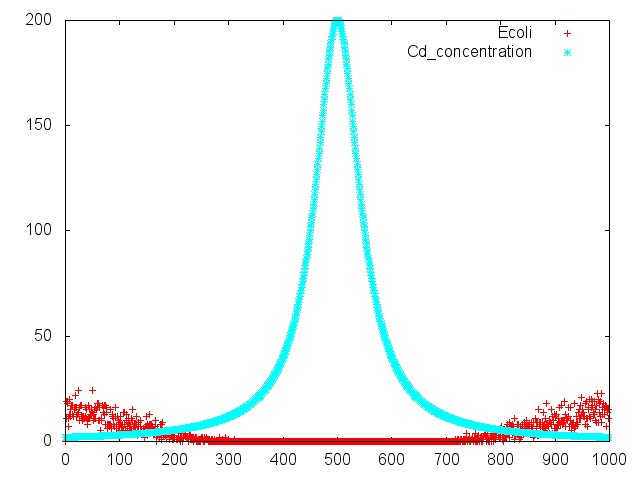
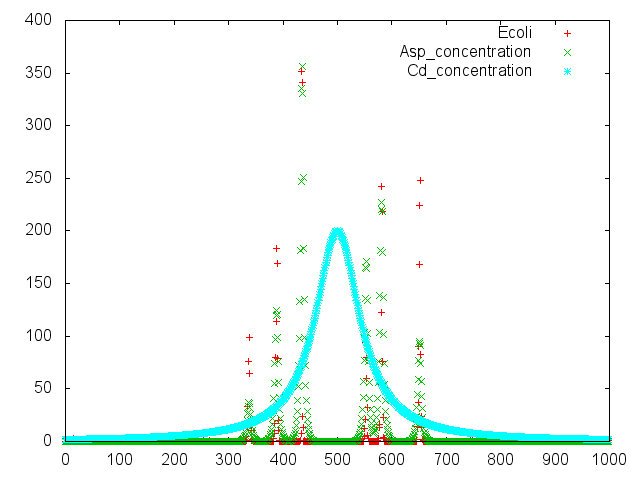
Capillary Assay
Result
Discussion
Future work
Reference
[1][http://www.ncbi.nlm.nih.gov/pubmed/12480884 Ferianc, P, et al, Regulation of yodA encoding a novel cadmium-induced protein in Escherichia coli, Microbiology, vol.148, 3801–3811, (2002)]
[2]iGEM Goettingen, Homing Coli, https://2012.igem.org/Team:Goettingen/Notebook/Results, (2012)
[3][http://www.ccbi.cam.ac.uk/iGEM2006/index.php/Protocols#Autoclaving_stock_solutions_or_glassware_.28sterilisation.29 iGEM University Of Cambridge, Making swimming Agar, http://www.ccbi.cam.ac.uk/iGEM2006/index.php/Protocols#Autoclaving_stock_solutions_or_glassware_.28sterilisation.29, (2006)]
[4]iGEM UT-Tokyo, L-aspartate chemotaxis assay, https://2011.igem.org/Team:UT-Tokyo/Data/Method, (2011)
[5][http://ecocyc.org/ Keseler et al, EcoCyc, http://ecocyc.org/, (2013)]
[6][http://www.ncbi.nlm.nih.gov/pubmed/17650676 Chmiel, A, Selection and activation of Escherichia coli strains for L-aspartic acid biosynthesis, Polish J. Microbiol, Vol.56, 71-76, (2007)]
[7][http://www.ncbi.nlm.nih.gov/pubmed/4632978 ADLER, J, A method for measuring chemotaxis and use of the method to determine optimum conditions for chemotaxis by Escherichia coli, Polish J. General Microbiol, Vol.74, 77-91, (1973)]
[8][http://www.ncbi.nlm.nih.gov/pmc/articles/PMC106306/ Sternberg, C, et al, New Unstable Variants of Green Fluorescent Protein for Studies of Transient Gene Expression in Bacteria, Polish J. Microbiol, Vol.64, 2240–2246, (1998)]
 "
"