Team:Austin Texas/kit
From 2014.igem.org
| Line 129: | Line 129: | ||
To use the kit properly, each culture must have two of the three plasmids from above. There must always be a pStG plasmid as well as either pFRYC or pFRY. However, additional controls can be conducted using only one of the three plasmids. | To use the kit properly, each culture must have two of the three plasmids from above. There must always be a pStG plasmid as well as either pFRYC or pFRY. However, additional controls can be conducted using only one of the three plasmids. | ||
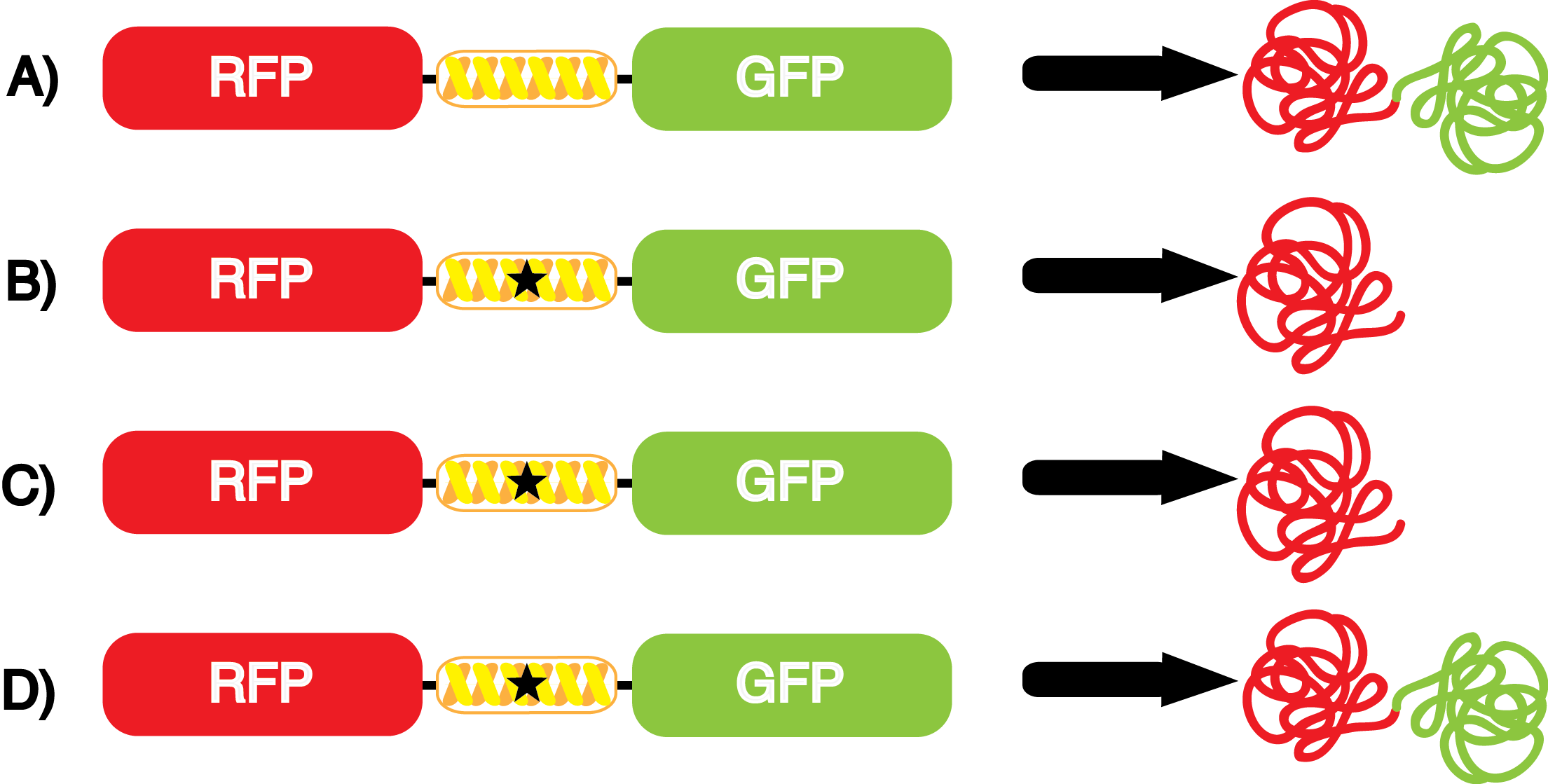
| - | [[File:NcAA kit incorpotation 10-14-14.png|450px|thumb|left|<b>Figure 3</b> Schematic demonstrating the gene expression of the kit plasmids under different growth conditions in the presence of IPTG.<br> | + | [[File:NcAA kit incorpotation 10-14-14.png|450px|thumb|left|<b>Figure 3.</b> Schematic demonstrating the gene expression of the kit plasmids under different growth conditions in the presence of IPTG.<br> |
A) pFRYC (+/-) pStG, (+/-) ncAA.<br> | A) pFRYC (+/-) pStG, (+/-) ncAA.<br> | ||
B) pFRY (-) pStG, (+/-) ncAA.<br> | B) pFRY (-) pStG, (+/-) ncAA.<br> | ||
| Line 191: | Line 191: | ||
<h2>Fidelity of Incorporation</h2> | <h2>Fidelity of Incorporation</h2> | ||
| - | [[File:UT_Austin_2014_Kit_Normalized_GFP_to_RFP_graph.png|thumb|600px|Figure 4. Graph showing the level of GFP fluorescence relative to RFP fluorescence for each condition. Each pStG plasmid is referred to based on the tRNA synthetase/tRNA pair present in the specific plasmid. Each of these plasmids was then paired with either pFRY or pFRYC and grown in the presence or absence of a specific ncAA. For example, the "3-AminoY-FRYC" and the "3-AminoY-FRY" samples both contain the 3-AminoY synthetase/tRNA pair and both samples were grown in the absence or presence of the ncAA "3-AminoY". Data are presented as the average of three independent cultures. Error bars denote standard deviation.]] | + | [[File:UT_Austin_2014_Kit_Normalized_GFP_to_RFP_graph.png|thumb|600px|'''Figure 4.''' Graph showing the level of GFP fluorescence relative to RFP fluorescence for each condition. Each pStG plasmid is referred to based on the tRNA synthetase/tRNA pair present in the specific plasmid. Each of these plasmids was then paired with either pFRY or pFRYC and grown in the presence or absence of a specific ncAA. For example, the "3-AminoY-FRYC" and the "3-AminoY-FRY" samples both contain the 3-AminoY synthetase/tRNA pair and both samples were grown in the absence or presence of the ncAA "3-AminoY". Data are presented as the average of three independent cultures. Error bars denote standard deviation.]] |
'''Set the experiment up - What do we want to test? How are we going to test it (keep it simple)? You may just need to rearrange the paragraph.''' | '''Set the experiment up - What do we want to test? How are we going to test it (keep it simple)? You may just need to rearrange the paragraph.''' | ||
| Line 208: | Line 208: | ||
<h2>Incorporation Value</h2> | <h2>Incorporation Value</h2> | ||
| - | [[File:UT_Austin_2014_Kit_Incorporation_Value_Graph.png|600px|thumb|Figure 5.<br> | + | [[File:UT_Austin_2014_Kit_Incorporation_Value_Graph.png|600px|thumb|'''Figure 5.'''<br> |
<i>May need to consider a different naming convention or the cultures. Possibly AY-C and AY-E for Control and Experimental?? </i> The fluorescence and OD600 readings of each culture were used to calculate a value for incorporation efficiency of each synthetase. <i>Need to add an explanation of how these values were calculated</i>]] | <i>May need to consider a different naming convention or the cultures. Possibly AY-C and AY-E for Control and Experimental?? </i> The fluorescence and OD600 readings of each culture were used to calculate a value for incorporation efficiency of each synthetase. <i>Need to add an explanation of how these values were calculated</i>]] | ||
Revision as of 16:54, 17 October 2014
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"