Team:Austin Texas/kit
From 2014.igem.org
| Line 191: | Line 191: | ||
<h2>Fidelity of Incorporation</h2> | <h2>Fidelity of Incorporation</h2> | ||
| - | [[File:UT_Austin_2014_Kit_Normalized_GFP_to_RFP_graph.png|thumb|600px|Figure | + | [[File:UT_Austin_2014_Kit_Normalized_GFP_to_RFP_graph.png|thumb|600px|Figure 4. Graph showing the level of GFP fluorescence relative to RFP fluorescence for each condition. Each pStG plasmid is referred to based on the tRNA synthetase/tRNA pair present in the specific plasmid. Each of these plasmids was then paired with either pFRY or pFRYC and grown in the presence or absence of a specific ncAA. For example, the "3-AminoY-FRYC" and the "3-AminoY-FRY" samples both contain the 3-AminoY synthetase/tRNA pair and both samples were grown in the absence or presence of the ncAA "3-AminoY". Data are presented as the average of three independent cultures. Error bars denote standard deviation.]] |
'''Set the experiment up - What do we want to test? How are we going to test it (keep it simple)? You may just need to rearrange the paragraph.''' | '''Set the experiment up - What do we want to test? How are we going to test it (keep it simple)? You may just need to rearrange the paragraph.''' | ||
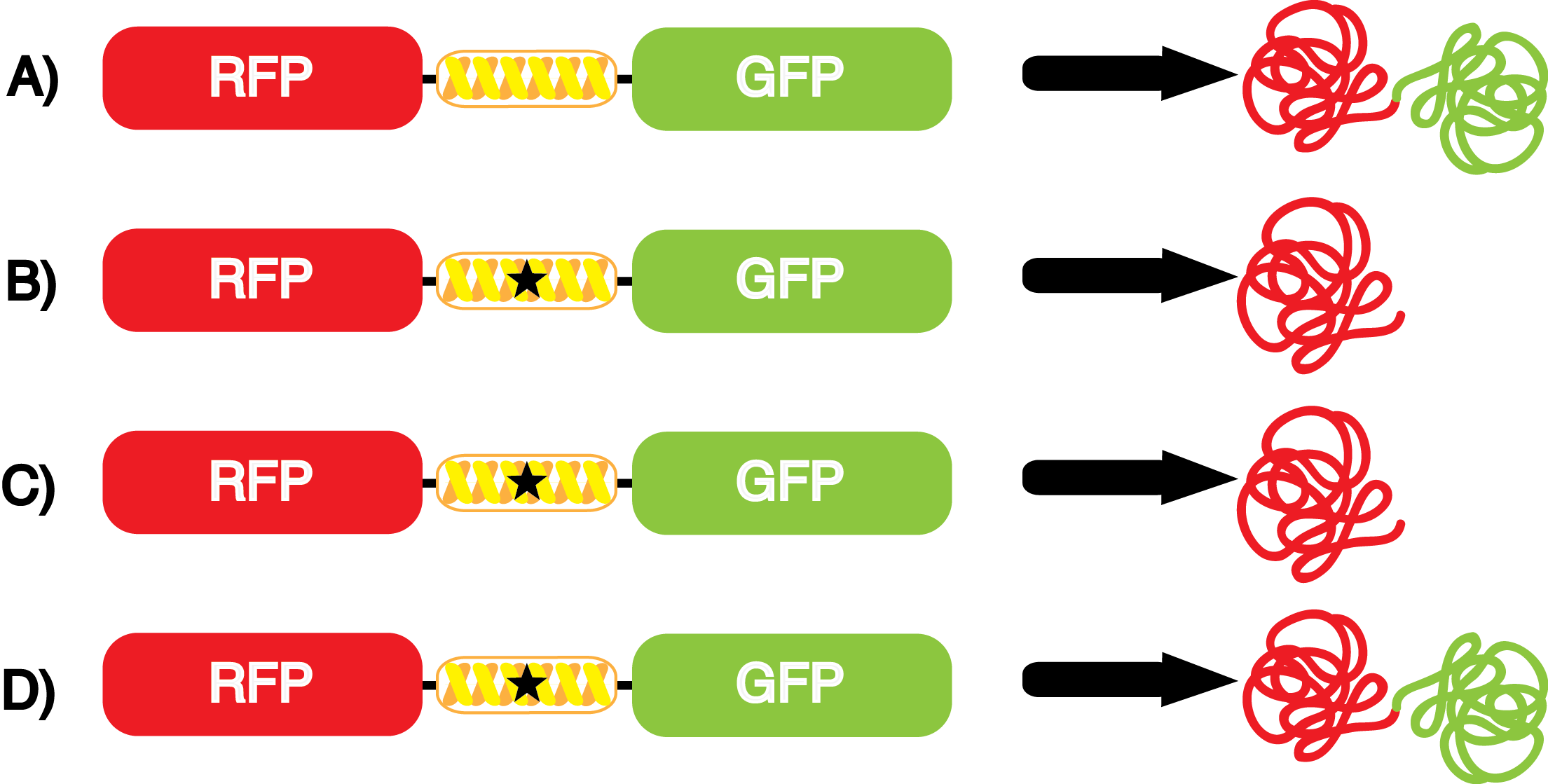
| - | We wanted to test whether the ncAA synthetase/tRNA pairs would incorporate anything besides their amino acid at the amber stop codon (UAG), and that they would in fact incorporate their specific ncAA if it was present. The fidelity of each ncAA sythetase/tRNA pair was measured by comparing the production of GFP in cultures containing pStG/pFRY with or without ncAAs. In the absence of an ncAA only RFP should be translated, as translation is expected to terminate between RFP and GFP at the amber stop codon on the linker sequence in pFRY. Alternatively, if the corresponding ncAA for pStG is present or if the synthetase/tRNA pair has low fidelity and can misincorporate a different amino acid, translation should continue through the UAG. In this case, RFP and GFP should both be translated. We also tested pStG/pFRYC strains in (+/-) ncAA conditions as a control for what effect the ncAA has on the fluorescence or growth of the cells. These strains should express RFP and GFP in all conditions since pFRYC does not have an amber stop codon in the linker (Figure | + | We wanted to test whether the ncAA synthetase/tRNA pairs would incorporate anything besides their amino acid at the amber stop codon (UAG), and that they would in fact incorporate their specific ncAA if it was present. The fidelity of each ncAA sythetase/tRNA pair was measured by comparing the production of GFP in cultures containing pStG/pFRY with or without ncAAs. In the absence of an ncAA only RFP should be translated, as translation is expected to terminate between RFP and GFP at the amber stop codon on the linker sequence in pFRY. Alternatively, if the corresponding ncAA for pStG is present or if the synthetase/tRNA pair has low fidelity and can misincorporate a different amino acid, translation should continue through the UAG. In this case, RFP and GFP should both be translated. We also tested pStG/pFRYC strains in (+/-) ncAA conditions as a control for what effect the ncAA has on the fluorescence or growth of the cells. These strains should express RFP and GFP in all conditions since pFRYC does not have an amber stop codon in the linker (Figure 2). However, if the presence of ncAA affects cell growth or fluorescence activity, we will need these controls to determine the extent of the effect. |
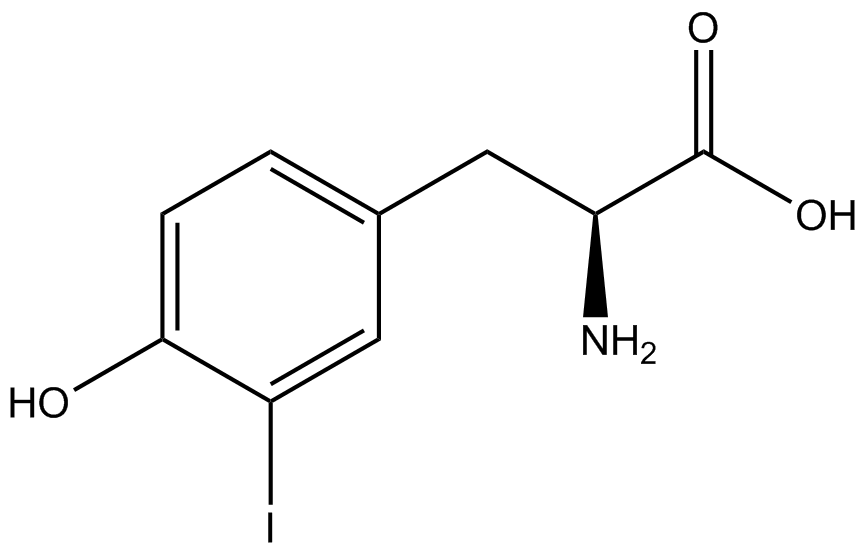
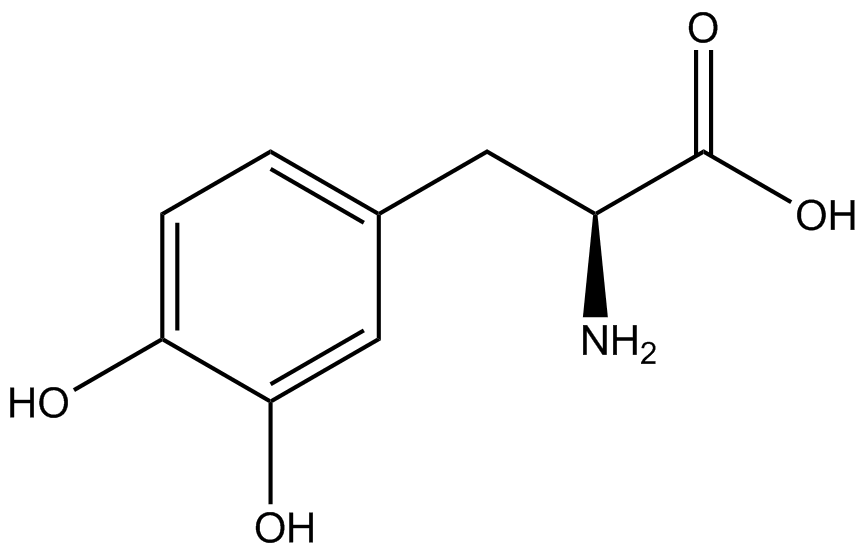
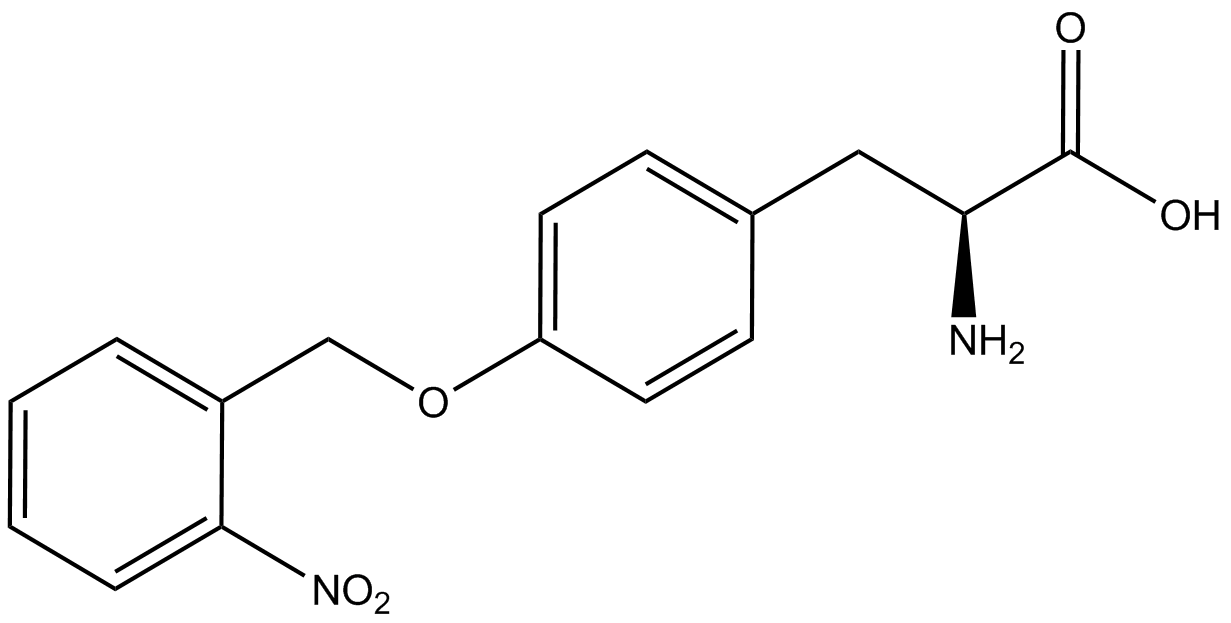
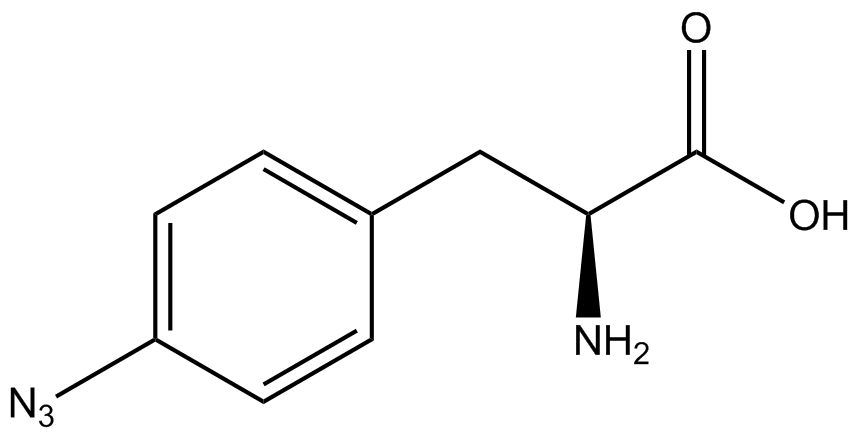
| - | To determine the change in GFP fluorescence when the ncAA was present, we first had to calculate how much GFP was expressed relative to the RFP, which would give an upper estimate of how much GFP could theoretically be expressed. We first divided both the GFP and RFP levels by the OD 600 of the culture in order to get the per cell fluorescence levels. We then normalized the GFP fluorescence for one culture to its RFP fluorescence so that we could compare the GFP fluorescence levels between cultures. The normalized GFP values were then compared between cultures grown in the presence of ncAA and cultures grown in the absence of ncAA, which would indicate how the level of GFP fluorescence changes when the ncAA is present. When these values were graphed (Figure | + | To determine the change in GFP fluorescence when the ncAA was present, we first had to calculate how much GFP was expressed relative to the RFP, which would give an upper estimate of how much GFP could theoretically be expressed. We first divided both the GFP and RFP levels by the OD 600 of the culture in order to get the per cell fluorescence levels. We then normalized the GFP fluorescence for one culture to its RFP fluorescence so that we could compare the GFP fluorescence levels between cultures. The normalized GFP values were then compared between cultures grown in the presence of ncAA and cultures grown in the absence of ncAA, which would indicate how the level of GFP fluorescence changes when the ncAA is present. When these values were graphed (Figure 4), some synthetase/tRNA pairs such as 4-azidophenylalanine, 3-nitrotyrosine, 3-iodotyrosine, and ortho-nitrobenzyltyrosine resulted in higher GFP fluorescence in the presence of ncAA than in the absence of ncAA, which suggests that those synthetases only incorporated an amino acid if their specific amino acid was present, meaning that they have a high fidelity. However, the other synthetase/tRNA pairs (3-aminotyrosine, L-DOPA, and cyanophenylalanine) did not show a significant increase in GFP fluorescence normalized to RFP fluorescence when the ncAA was present, indicating that they would incorporate other amino acids at the amber stop codon when their specific amino acid was not present (and perhaps even when it was), and thus have a low fidelity. |
<h2>Synthetase Efficiency</h2> | <h2>Synthetase Efficiency</h2> | ||
| Line 205: | Line 205: | ||
| - | For our results (Figure | + | For our results (Figure 4), two synthesase/tRNA pairs stood out as relatively inefficient: 3-nitrotyrosine and ortho-nitrobenzyltyrosine. Both of these synthetase/tRNA pairs showed a significantly smaller normalized GFP to RFP fluorescence when the amino acid was present with the pFRY construct. While they both show a significant increase in normalized GFP to RFP fluorescence when the amino acid was present compared to when it was absent, which indicates a high fidelity, the actual GFP fluorescence relative to the RFP fluorescence was only around 50% for 3-nitrotyrosine and 20% for ortho-nitrobenzyltyrosine. These results suggest that for whatever reason, these synthetase/tRNA pairs do not always incorporate their ncAA at the amber stop codon. |
<h2>Incorporation Value</h2> | <h2>Incorporation Value</h2> | ||
| - | [[File:UT_Austin_2014_Kit_Incorporation_Value_Graph.png|600px|thumb|Figure | + | [[File:UT_Austin_2014_Kit_Incorporation_Value_Graph.png|600px|thumb|Figure 5.<br> |
<i>May need to consider a different naming convention or the cultures. Possibly AY-C and AY-E for Control and Experimental?? </i> The fluorescence and OD600 readings of each culture were used to calculate a value for incorporation efficiency of each synthetase. <i>Need to add an explanation of how these values were calculated</i>]] | <i>May need to consider a different naming convention or the cultures. Possibly AY-C and AY-E for Control and Experimental?? </i> The fluorescence and OD600 readings of each culture were used to calculate a value for incorporation efficiency of each synthetase. <i>Need to add an explanation of how these values were calculated</i>]] | ||
Revision as of 03:31, 17 October 2014
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"