Team:LMU-Munich/Project/Bakillus
From 2014.igem.org
BaKillus - Engineering a pathogen-hunting microbe
Increasing bacterial resistance to classical antibiotics remains a serious threat and urges the development of novel pathogen-killing strategies. Exploiting bacterial communication mechanisms such as quorum sensing is a promising strategy to specifically target certain pathogens. The major aim of this project is the introduction of a genetic circuit enabling Bacillus subtilis to actively detect, attach to, and eventually kill Staphylococcus aureus and Streptococcus pneumoniae. Initially, we will introduce the autoinducer-sensing two-component systems of S. aureus and S. pneumoniae into B. subtilis to create a pathogen-detecting strain. By utilizing quorum sensing-dependent promoters, we will then trigger pathogen-killing strategies like the production of antimicrobial peptides or biofilm degradation. As a safety measure, a delayed suicide-switch guarantees non-persistence of genetically modified B. subtilis in the absence of pathogens. We envision the use of BaKillus as a smart, cheap and simple-to-use medical device for diagnostics and targeted treatment of multiresistant superbugs.
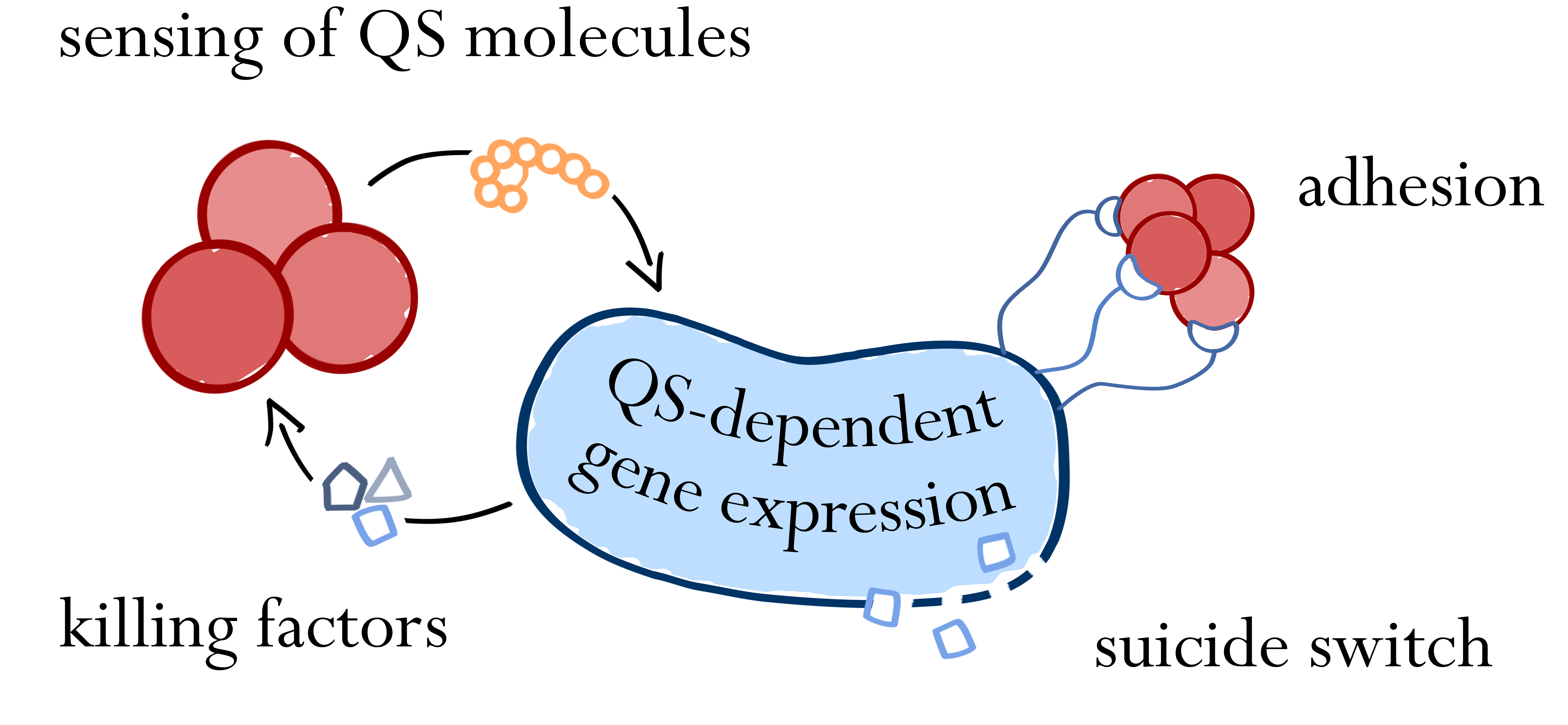

Sensing
Lorem ipsum dolor sit amet, consetetur sadipscing elitr, sed diam nonumy eirmod tempor invidunt ut labore et dolore magna aliquyam erat, sed diam voluptua. At vero eos et accusam et justo duo dolores et ea rebum. Stet clita kasd gubergren, no sea takimata sanctus est Lorem ipsum dolor sit amet. Lorem ipsum dolor sit amet, consetetur sadipscing elitr, sed diam nonumy eirmod tempor invidunt ut labore et dolore magna aliquyam erat, sed diam voluptua. At vero eos et accusam et justo duo dolores et ea rebum. Stet clita kasd gubergren, no sea takimata sanctus est Lorem ipsum dolor sit amet.

Adhesion
As part of the BaKillus strategy, we want to reduce the selective pressure on non-pathogenic bacteria to prevent resistance development. To this end, BaKillus is equipped with pathogen-binding peptides on its surface, thereby adhering specifically to the targeted bacteria. This measure ensures high local concentration of killing factors while minimizing harmful effects on untargeted species.
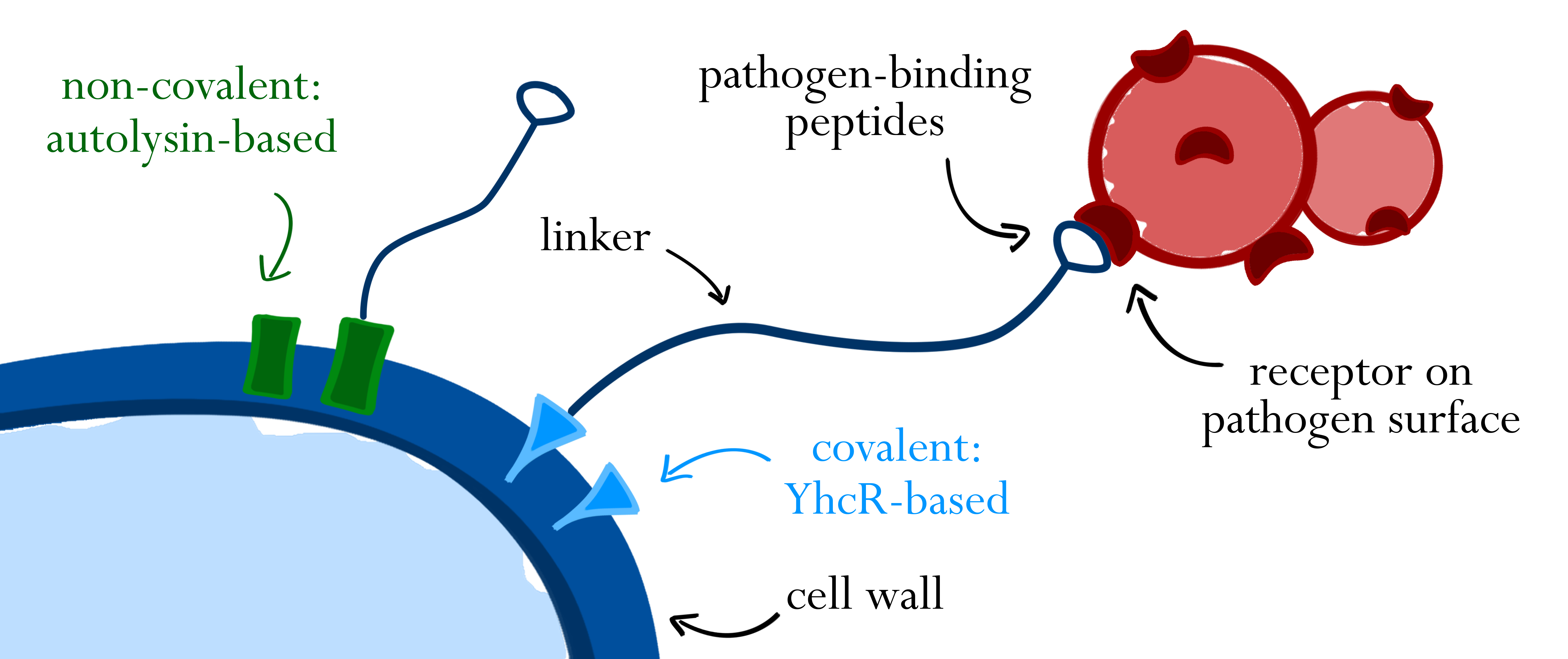
Surface display of the pathogen-binding peptides is achieved by translational fusions to naturally occurring surface proteins of B. subtilis. Cell-wall binding modules of autolysins are used for non-covalent interaction with the B. subtilis cell wall, whereas a sortase-based system functions via covalent attachment to the cell wall. In both cases, a linker between cell wall anchor and peptide allows flexibility of the fusion protein and increases its range. Peptides binding to different receptors on the surface of pathogens mediate specific binding.
Bacterial cell surfaces are covered with a wide range of proteins functioning as transporters, interacting with molecules and cells or fulfilling a variety of other functions. As they are naturally displayed on the cell’s surface, they can also be used for presenting heterologous proteins or peptides.
One class of proteins that have been used for surface display are B. subtilis autolysins, in particular LytC and LytE [1, 2]. Autolysins hydrolase peptidoglycan and thus play a role in cell wall turnover, vegetative growth and cell division. They consist of an N-terminal cell wall binding (CWB) domain with a signal peptide directing export via the Sec pathway and of a C-terminal catalytic domain [3, 4]. Fusions of the CWB domains of LytC and LytE to heterologous proteins have been shown to successfully facilitate surface display of their passenger protein in B. subtilis [1, 2].
Another promising system is based on the B. subtilis sortase YhcS and its substrate YhcR. Sortases represent a class of membrane-anchored proteins in gram-positive bacteria, which recognize their substrates via a conserved C-terminal pentapeptide sequence. They catalyze covalent binding of their substrates to lipid II, an intermediate of peptidoglycan synthesis [5]. Two putative sortase-substrate pairs have been identified in B. subtilis, one of them being YhcS/YhcR, which has already been used for successful surface display of recombinant proteins [5, 6].
Bacterial adhesins mediate binding of the cells to all kind of surfaces, thereby preventing removal by physical influences like sheer forces. Attachment of pathogens to their host is often crucial for infection, which is why many adhesins are listed as virulence factors [7]. Binding partners of adhesins are often components of the extracellular matrix, which would be difficult to express on the BaKillus surface. In contrast, proteins of human cells that get bound by adhesins are optimal candidates for translational fusions to surface display systems. BaKillus mimics human cells by expressing parts of the proteins on its surface, leading to binding of the pathogens to BaKillus. Barbu et. al. 2010 [8] and Muchnik et. al. 2013 [9] used phage display to identify the binding peptide interaction partners of Staphylococcus aureus virulence factor SdrC and Streptococcus pneumoniae adhesin NOX, respectively. Those peptides are the specific binding sequences of SdrC and NOX within surface proteins of human cells. Thus, expression of the peptides on the BaKillus surface should enable it to adhere to the targeted pathogens.
Constructs for non-covalent and covalent surface display were cloned similar to those described by Chen et. al., 2008 [1], and Liew et. al., 2012 [6], respectively.
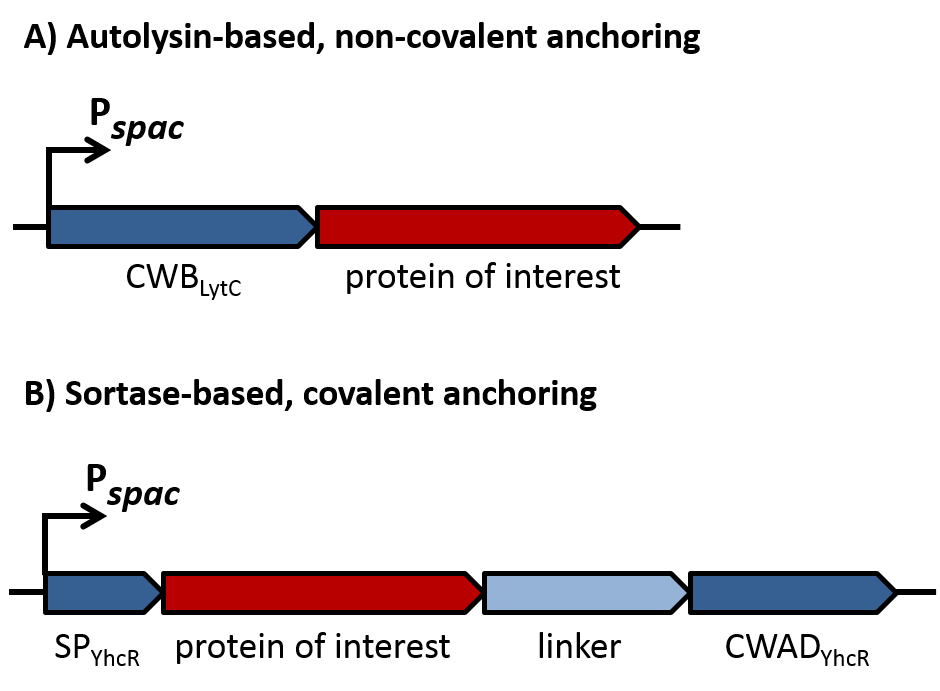
As depicted in figure 1A, cell-wall binding domains (CWB) of LytC or LytE (BBa_1351006 or BBa_1351007) were translationally fused to a protein of interest for non-covalent anchoring of the latter to the cell wall. These CWB domains also included the native signal peptide (SP) of the respective proteins to direct export via the Sec pathway. For covalent anchoring, signal peptide (BBa_1351008) and cell wall anchoring domain (CWAD, BBa_1351010) of the sortase substrate YhcR and a 56 amino acid linker (BBa_1351009) were assembled with the protein of interest as depicted in figure 1B to generate a YhcR-like fusion.
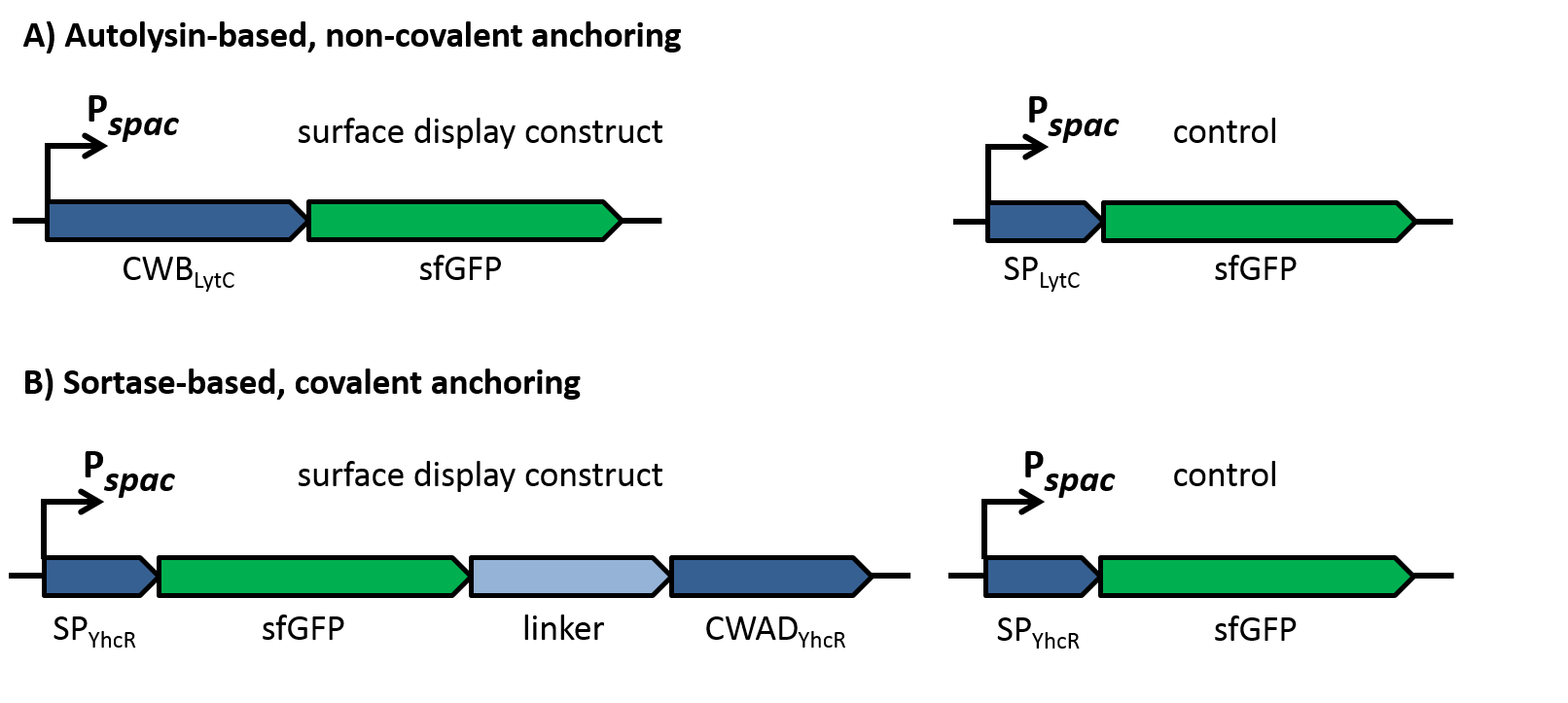
To evaluate the functionality of the constructs, the latter were cloned into the replicative B. subtilis vector pBS0K under the control of the inducible promoter Pspac. Two different proteins were used to test for successful display: Firstly, a superfolder GFP variant (sfGFP) was used for fluorescence microscopy to check if the constructs locate to the cell wall as shown in figure 2. This variant of GFP had been shown previously to fold in the periplasm of gram-negative bacteria [10] and could thus be expected to be fuctional also in the extracellular environment, in contrast to normal GFP that does not fold under reducing conditions. As controls for the sfGFP constructs, only the signal peptides without the respective cell wall binding or anchoring domains were fused to sfGFP, which should lead to secretion of the fusion into the growth medium.
Secondly, fusions to alkaline phosphatase (PhoA) were used to detect surface accessibility of the presented protein, as PhoA is only active when exported out of the cytoplasm [11]. Agar plates containing the PhoA substrate XP (5-bromo-4-chloro-3-indolyl-phosphate) indicate enzyme activity by turning blue where the substrate is hydrolyzed and further oxidized to an indigo dye [12]. PhoA was also fused to intracellular GFP as negative control and to an extracellular domain of LiaI as positive control. The PhoA-deficient B. subtilis strain MH3402 was used for expression of all PhoA constructs.
Binding of BaKillus to pathogens in general, and for this project to S. aureus and S. pneumoniae in particular, should be mediated by specific peptides that had been identified via phage display. Unfortunately, evaluation of the S. aureus-specific peptide Nrx1b (BBa1351000) was not possible due to the lack of an S2 laboratory, as the Nrx1b-binding partner SdrC is a virulence factor of S. aureus.
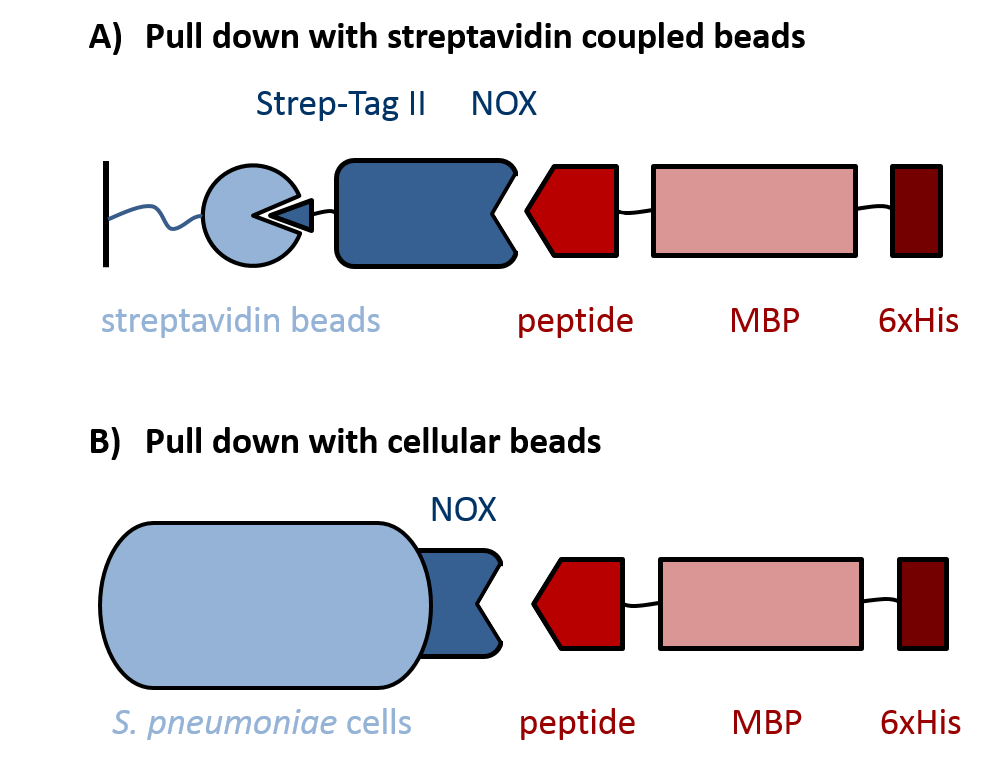
Evaluation of peptides binding to S. pneumoniae was done by a biochemical pull down assay with their binding partner NADH oxidase (NOX). As shown in figure 3, NOX was bound to streptavidin-coupled beads via an N-terminal fused Strep-Tag II, for which the pASK-IBA expression vector system was used. S. pneumoniae-specific peptides C4P, CSP and L5P (BBa1351001, BBa1351002 and BBa1351003, respectively) were fused C-terminally to a maltose-binding protein (MBP) and a 6xHis-tag. The MBP prevents degradation of the short peptides by proteases and was used for detection of the peptide fusions on an SDS gel. To increase specificity of the detection, the 6xHis-tag was used for Western Blot analysis of the pulled down proteins. Additionally, S. pneumoniae itself was used as cellular beads for a pull down of the same peptide-MBP-His constructs, again followed by analysis with SDS gel and Western Blot.
Surface display
Pathogen-binding peptides
Surface Display
Pathogen binding
Killing
A key aspect of our project is the killing module, which enables BaKillus to dispatch pathogens. Therefore, we implement a variety of killing strategies into our chassis.
One of our killing factors is subtilin, a lanthionine-containing antimicrobial peptide. Just like other lantibiotics, it is active against a wide-range of Gram-positive bacteria by inhibiting the cell wall synthesis and formation of membrane pores. The latter leads to the depolarisation of the membrane potential and thus, to rapid cell death. We decided to use subtilin, as it is natively produced by Bacillus subtilis ATCC6633, a close relative to our chassis and as lantibiotics are considered as alternative to classical antibiotics to which many bacteria already developed resistance.
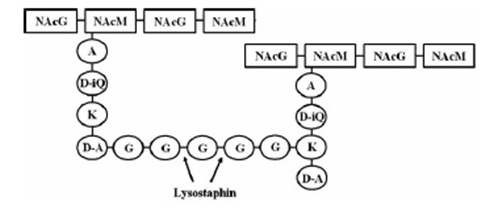
Another killing factor is lysostaphin, a metalloendopeptidase, which lysates the cellwall of some Staphylococcus species. Since this enzyme is highly specific, and S. aureus is especially sensitive towards lysostaphin, this is an ideal killing strategy for our BaKillus. In addition, lysostaphin shows no signs of toxicity and a very low potential for allergic reactions in first tests on animals and humans.
Since pathogens are often protected by biofilms and thus not very sensitive towards killing agents, dispersin was integrated as an auxiliary device. This glycosaminhydrolase destroys biofilms of many gram-positive and gram-negative bacteria, including S. aureus, making them more susceptible for antibiotics and other pathogen defeating agents.
Both, lysostaphin and dispersin where developed as BioBricks and tested in B. subtilis by the iGEM Team iGEM12_Lyon_INSA in 2012, and we aim to evaluate the effectiveness of this two S. aureus defeating agents.
Subtilin is a small antimicrobial peptide (AMP) which belongs to the class of lanthionine-containing antibiotics (lantibiotics). These are heat-stable, ribosomally synthesized molecules with a molecular weight below 4 kDa. Their main characteristic is the high proportion of unusual amino acids, synthesized by post-translational side-chain modifications of precursor peptides. Most prominent examples are the polycyclic thioether amino acids, lanthionine and –methyllanthionine and a number of dehydrated amino acids such as dehydroalanine (Dha) and dehydrobutyrine (Dhb).
Lantibiotics are encouraging candidates for future antimicrobials, as they inhibit the growth of many clinically relevant pathogens comprising even multidrug-resistant bacteria. Moreover, lantibiotics have only low tendency to generate resistance. Both features make them highly attractive for medical applications.
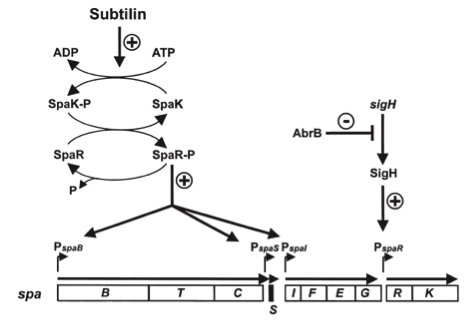
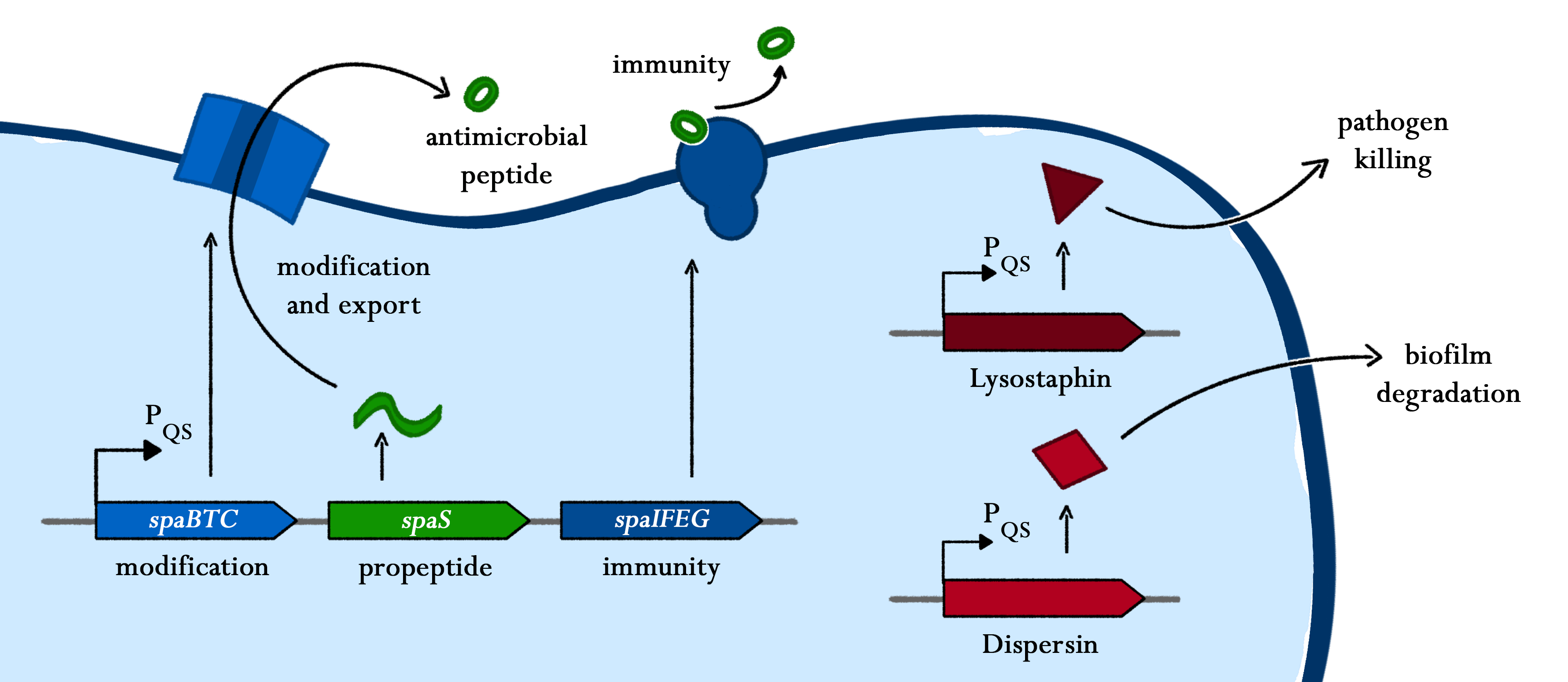
As true for many other lantibiotics, the genes for the subtilin biosynthesis are clustered. For subtilin, there is even a reasonable arrangement within the cluster. The genes spaBTC play a role for the posttranslational modifications and the transport of subtilin. They are regulated by their own promotor PspaB and polycistronically transcribed. spaS encodes the propeptide, which is derived from a short monocistronic mRNA. The next transcription entity, spaIFEG, are coding for the immunity, whereas spaRK encode an two-component-system that plays an important role for the regulation.. What is missing in comparison to other lantibiotic gene clusters is a specific protease (encoded by lanP) that cleaves off the leader sequence. In Bacillus subtilis, this task is accomplished by unspecific extracellular proteases.
TABLE MISSING
Of course, the producer needs to be immune against the antimicrobial agent it produces. B. subtilis ATCC6633 protects itself against subtilin by two mechanisms that both act independently and confer some level of resistance, however full resistance is only achieved when both mechanisms are active. SpaI is a membrane-anchored lipoprotein. It is suggested that it binds the antimicrobial peptide and by this keeps it away from the membrane. SpaFEG is a LanFEG-like ABC-transporter that works by exporting subtilin from the cytoplasmic membrane to the culture supernatant.
Subtilin biosynthesis and immunity in B. subtilis ATCC6633 are subject to a dual control mechanism.
To begin with, subtilin production is positively regulated by sigma factor H (SigH). sigH itself is repressed by the transition state regulator AbrB during exponential growth phase. Thus, subtilin production is highest at the transition to stationary phase. Moreover, when a threshold level of subtilin concentration is reached in the extracellular space, the peptide acts as a pheromone. Subtilin activates the two-component system SpaRK, consisting of a histidine kinase and a response regulator, which in turn induces expression of spaBTC, spaS and spaIFEG. This dual control mechanism allows the coordination of subtilin biosynthesis with the physiological state of the cell.
Lantibiotics are active against a wide range of Gram-positive bacteria including multidrug-resistant pathogens. Some lantibiotics have a dual mode of action, subtilin is proposed to be among them. Subtilin forms a complex with the cell wall precursor lipid II, whereby the pyrophosphate moiety may play a key role for target recognition. Thereby, the cell wall biosynthesis is inhibited. Furthermore, the complexes aggregate and form a pore in the bacterial membrane by using lipid II as a docking molecule. This leads to a depolarisation of the membrane potential, the efflux of cytoplasma and in turn to rapid cell death. Just like nisin, which is highly similar, subtilin is still active in the nanomolar range and thus serves excellently as alternative killing strategy against multidrug-resistant pathogens.
Lysostaphin is a murein-hydrolase, which is naturally produced by Staphylococcus simulans biovar staphylolyticus in order to defeat other, competing Staphylococcus species. It cleaves specifically the pentaglycinbridge between the tetrapeptides of the peptidoglycan in the cell wall and in addition to kill the cells itself, it also dissolves the biofilms of lysostaphin-sensitive bacteria. [A1]
This targeted pentaglycinbridge is a feature of some Staphylococcus species, leading to the high specificity of lysostaphin against S. carnosus, S. epidermidis and Staphylococcus aureus, which is particularly sensitive towards lysostaphin. [A2]
The immunity of S. simulans is accomplished by substitution of the glycine-residues in the cell wall by serine and mediated by the immunity factor Lif [A1]
The preprolysostaphin consists of a signal peptide to mediate the export, 15 tandem repeats of 13 aminoacids length and two protein domains: the peptidase-domain and the C-terminal targeting domain. The signal peptide is cleaved during the export, the tandem repeats are cleaved by an additionally secreted cysteine protease. This repeats are not necessary for correct protein folding or maturation of the protein, they just inhibit protein activity. Once cleaved, the maturated protein is 4.5 fold more active. [A1]
Because of its promising attributes, the usage of lysostaphin as coating for implants [A3] and treatment of S. aureus in the nasal area via cream [A4] is currently in the focus of medical research.
[A1] M.C.F. Bastos, H. Ceotto1, M.L.V. Coelho and J.S. Nascimento: Staphylococcal Antimicrobial Peptides: Relevant Properties and Potential Biotechnological Applications in Current Pharmaceutical Biotechnology (2009) S. 38-61
[A2] Jaspal K. Kumar: Lysostaphin: an antistaphylococcal agent in Appl Microbiol Biotechnol (2008) S.555-561 )
[A3] Rohan Satishkumar, Sriram Sankar, Yuliya Yurko, Amy Lincourt, John Shipp, B. Todd Heniford and Alexey Vertegel: Evaluation of the Antimicrobial Activity of Lysostaphin-Coated Hernia Repair Meshes in Antimicrobial Agents and Chemotherapy (2011)
[A4] John F. Kokai-Kun, Scott M. Walsh, Tanya Chanturiya, and James J. Mond: Lysostaphin Cream Eradicates Staphylococcus aureus Nasal Colonization in a Cotton Rat Model in Antimicrobial Agents and Chemotherapy, 47 (2003)
DispersinB (DspB) is a biofilm-dissolving enzyme, specifically a 1,6-beta-N-acetyl-glucosaminidase, produced by Aggregatibacter actinomycetemcomitans, a pathogen, which causes aggressive forms of Parodontitis. [A5]
In its natural habitat, A. actinomycetecombitans uses this enzyme to release single cells from the biofilm in order to colonize new habitats. [A6]
DspB is specific towards its substrate, PNAG, (PGA in E. coli), which is an important component of the extracellular matrix and which can make up to 90% of the dry weight of a biofilm. [A7]
The process of dissolving biofilms is accomplished by cleaving monosaccharide-residues from the polysaccharide, beginning at the non-reducing end of the PNAG. [A6]
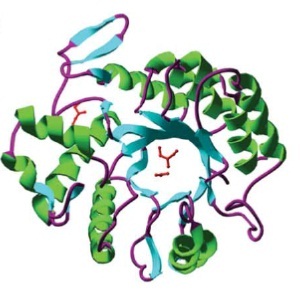
This glycosamin-hydrolase has a typical TIM-barrel-structure, consisting of 8α- and 8β-barrels, with a large cavity in the middle, which is considered to be the substrate-binding pocket. [A5]
[A5] Erika Fazekas, Lili Kandra, Gyongyi Gyemant: Model for b-1,6-N-acetylglucosamine oligomer hydrolysis catalysed by DispersinB, a biofilm degrading enzyme, in Carbohydrate Research 363 (2012), S. 7-13
[A6] N. RamasubbuL. M. Thomas, C. Ragunath and J. B. Kaplan: Structural Analysis of Dispersin B, a Biofilm-releasing Glycoside Hydrolase from the Periodontopathogen Actinobacillus actinomycetemcomitans, in Journal of Molecular Biology 340 (2005) S. 475-486
[7] Jeffrey B. Kaplan, Kabilan Velliyagounder, Chandran Ragunath, Holger Rohde,
Dietrich Mack, Johannes K.-M. Knobloch, and Narayanan Ramasubbu: Genes Involved in the Synthesis and Degradation of Matrix Polysaccharide in Actinobacillus actinomycetemcomitans and Actinobacillus pleuropneumoniae Biofilms, in JOURNAL OF BACTERIOLOGY, 186 (2004) S.8213-8220
[A8] Suba G. A. Manuel, Chandran Ragunath, Hameetha B. R. Sait, Era A. Izano, Jeffrey B. Kaplan and Narayanan Ramasubbu: Role of active-site residues of dispersin B, a biofilm-releasing b-hexosaminidase from a periodontal pathogen, in substrate hydrolysis, in The FEBS journal (2007)
[A9] Darouiche RO, Mansouri MD, Gawande PV, et al. Antimicrobial and antibiofilm efficacy of triclosan and DispersinB combination. J Antimicrob Chemother 2009; 64: 88-93
The Biobrick K802000, built by the iGEM Team iGEM12_Lyon_INSA, was kindly sent by the team Insa-Lyon. Via PCR-amplification, the actual lyosostaphin-gene was amplified, adding the restriction sites required for the RFC 25.
The coding sequence was fused with the promoter Pxyl and into the plasmid pBS1C. After transformation into B. subtilis, the iGEM-team Groningen will evaluate the effectiveness of lysostaphin, produced by B. subtilis against S. aureus via spot on lawn test. This test will be conducted with living cells as well as with cell lysate in order to have a clue, whether the lysostaphin is secreted into the substrate by B.subtilis or not.
The Biobrick K802001 built by the iGEM Team iGEM12_Lyon_INSA, was kindly sent by the Team Insa-Lyon. The original DspB contains an AgeI restriction site, wich was deleted via site directed mutagenesis by overlap extension PCR in order to make the gene compatible with the Freiburgstandard.
Overhangs with RFC 25 compatible restriction sites where added during this PCR and the PCR-product was cloned into the TOPO-vector.
After sequence confirmation, the DspB is cloned into pSB1C3 in order to be send in as a BioBrick. A His-Tag is added C- and N-terminal, and a copy of each of the Histag-constructs, and one copy without Histag is fused with the promoter Pxyl and cloned into pBS1C. The transformation into B. subtilis will follow.
In order to test the biofilm dissolving abilities of this recombinant B. subtilis, biofilms of E. coli Nissle 1917, an excellent biofilmformer with GRAS-status, are grown in 96 well plates. The biofilms will be dyed with crystalviolett and living B. subtilis cells as well as cell lysate will be transferred into the wells.
If biofilm-destruction takes places, the crystalviolett will vanish by carefully washing the plates.
Additionally, an expression control will be conducted via Western Blot.
A first sequencing, prior to the PCR-Amplification revealed an error in the DNA-sequence provided by the Registry. The actual sequence of the lysostaphin of the BioBrick [http://parts.igem.org/Part:BBa_K802000 K802000] is codon-optimized for B. subtilis.
The correct sequence was posted as a user review in the experience section of the Registry.
After cloning the lysostaphin into pBS1C with the promoter Pxyl, the construct was verified by sequencing and transformed into B. subtilis.
A culture in DSM was incubated at 37°C and 220 rpm and 5µl of this culture where spotted on a sterile filterpaper and sent to the iGEM-team Groningen.
The sample of the BioBrick K802001, which was originally sent by the Registry contained a wrong gene.
Fortunately, the sample sent by the iGEM-Team Insa_Lyon, was the matching plasmid to the given sequence. Thank you very much for providing us with this part!
The mutated gene was cloned into the TOPO-Vektor and transformed into E. coli DH5α.
Subtilin

The spaBTCSIFEGRK gene cluster
Self-protection from subtilin
Dual regulation of subtilin biosynthesis and immunity
Mode of action

Lysostaphin
Sources:
Dispersin
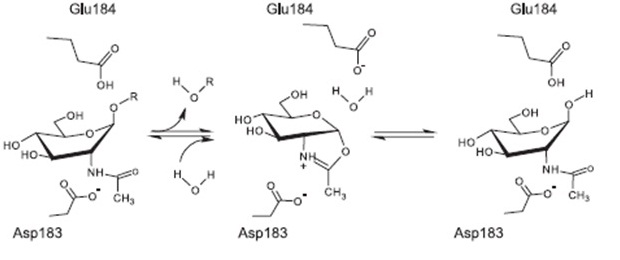
The currently proposed mechanism of reaction suggests a substrate assisted nucleophilic attack (see chart below. [A8]
DspB is active against many bacteria, both gram-positive and gram-negative, and shows some very promising features for its use in medical devices.
It can prevent the growth of biofilms in coated catheters if used in combination with triclosan, and can be used to clear medical devices of biofilms. [A9]
Sources:

Lyostaphin:
Dispersin:
Lysostaphin
Dispersin
Suicide Switch
Bacillus subtilis has the ability to differentiate into several sub-populations during stationary phase, influenced by the regulation of different gene-clusters. As one of the options, a sub-population can become a so called "Cannibalism" toxin producer - a process which is driven by the master regulator Spo0A. Those cannibalistic cells kill their non-cannibalistic mates and feed on the nutrients set free upon lysis.

Hi there!
Welcome to our Wiki! I'm BaKillus, the pathogen-hunting microbe, and I'll guide you on this tour through our project. If you want to learn more about a specific step, you can simply close the tour and come back to it anytime you like. So let's start!

What's the problem?
First of all, what am I doing here? The problem is, pathogenic bacteria all around the world are becoming more and more resistant against antimicrobial drugs. One major reason for the trend is the inappropriate use of drugs. With my BaKillus super powers, I want to reduce this misuse and thus do my part to save global health.

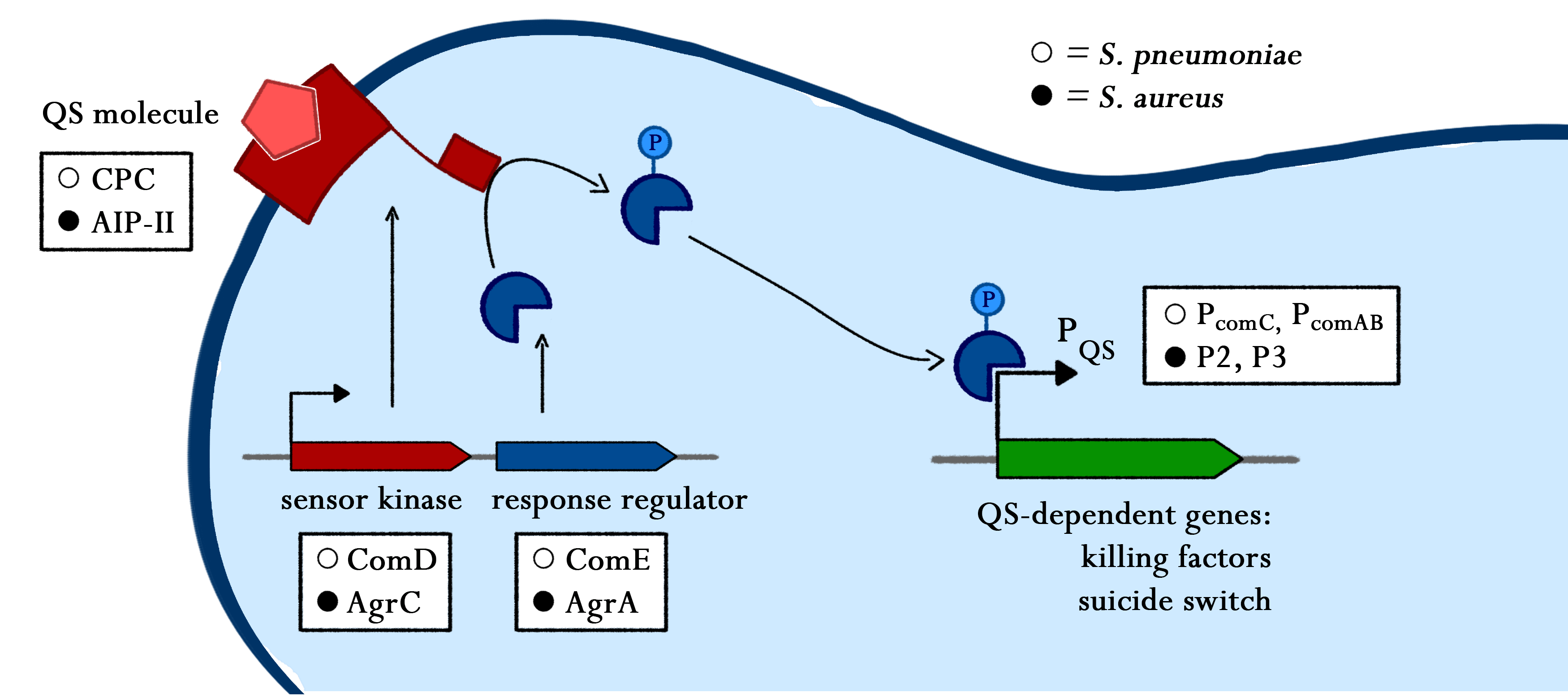
Sensing of pathogens
To combat the pathogenic bacteria, I simply eavesdrop on their communication. Bacteria talk with each other via quorum sensing systems, which I use to detect them and trigger my responses.

Adhesion
The more specific and effective I can use my powers, the lower the danger is of provoking new resistance development. So I catch pathogens whenever I get hold of them and stick to them until my work is done.

Killing
Talking about my work - killing pathogens is finally what I am made for. In response to quorum sensing molecules of the pathogens, I export a range of antimicrobial substances leading to dissipation of biofilms and the killing of the targeted bacteria.

Suicide switch
When the job is done and all the bad guys are finished, you don't need a super hero anymore. So after fulfilling my work I say goodbye to the world by activating my suicide switch.

Application
Of course I'm not only a fictional hero, but a very real one. In two different prototypes, I could be used for diagnosis or treatment of pathogen-caused diseases. However, there is still a whole lot of regulational and economical questions that have to be answered before.

See you!
So now you know my short story - and it is time for me to return to my fight for a safer world. Feel free to take a closer look on my super powers, the process of my development or the plans for a medical application.
 "
"