Team:NCTU Formosa/results
From 2014.igem.org
Contents |
Magic Power of Our Pyramidal Device
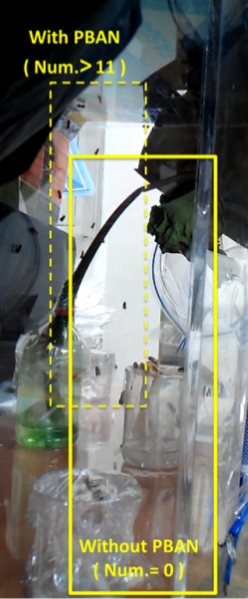
Our device combines blue light and PBAN to achieve a powerful and specific insect attraction. In this video, we do a test to see how this combination creates an effect greater than either blue light or PBAN alone. Firstly, we feed PBAN to a female moth by placing the moth in a small beaker that contains PBAN. We covered the beaker with plastic wrap in order to keep the moth inside. Soon, we can see that the female moth starts to flap its wings frantically. This is a sign of sexual stimulation, and from this point on, the female moth starts to release pheromones.
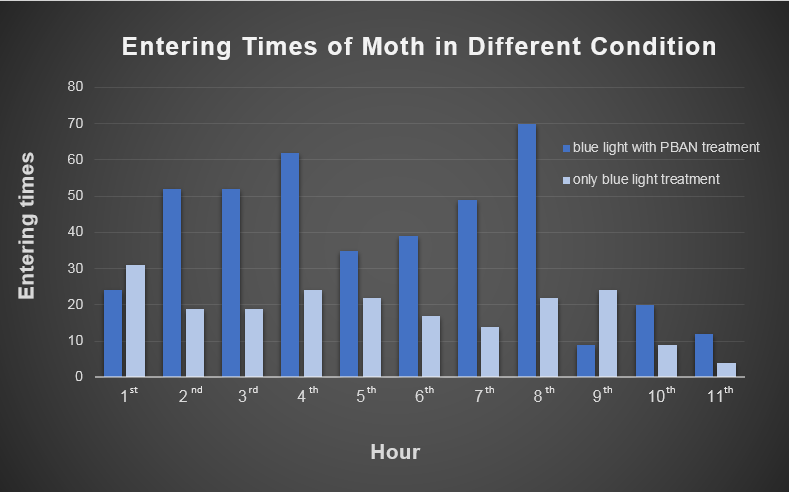
Secondly, we transfer the beaker into our device. Then we position the device in an acrylic chamber to begin our test. We keep the chamber dark so that blue light would be the only light source inside. We did a long-time observation to record the number of insects per hour entering the device . In Fig.2-0-1, we can clearly see the magic power of our device in attracting insects.
Our experiment can be divided into two categories.
1. PBAN Biobricks Tests: gene recombination and protein expression.
2. Insect Tests: PBAN effect test, insect behavior test and device test.
PBAN Biobricks Test
PBAN Gene Synthesis (Full Gene Sequence Design Process)
To capture the harmful insects causing damage in agriculture, we first found 9 different kinds of PBAN peptide of harmful insects common in many places of the world from our long literature review. Next, we obtain the DNA sequences by reversely translating the peptide sequences of these PBANs from NCBI (EX: PBAN Spodoptera litura:http://www.ncbi.nlm.nih.gov/protein/AAK84160.1 ) Finally, we modified every codon on the DNA sequence and designed the DNA sequence for E.coli to express a certain PBAN.
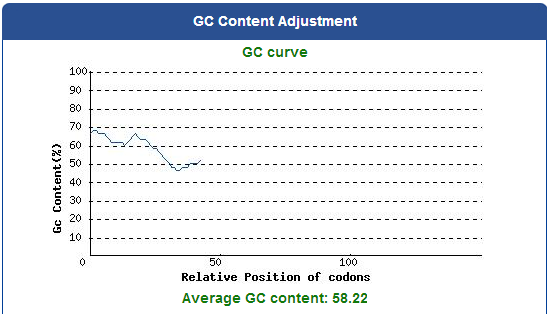
DNA Modification Process:
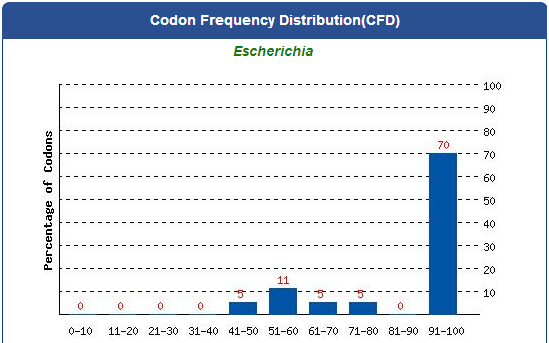
1. Avoid the rare codons of E.coli, and choose high frequency codons.
( Frequency Table Tool:http://www.genscript.com/cgi-bin/tools/codon_freq_table )
Use [http://www.genscript.com/cgi-bin/tools/rare_codon_analysis Rare Codon Analysis Tool] to inspect if there are any problems to express our gene for E.coli.
2. Avoid choosing the same codon when modifying our designed gene sequence to prevent the E.coli from running out of nucleotides due to repeated use.
3. Avoid the start codon ATG in the middle of the coding sequence.
Take the PBAN of Spodoptera litura for example:
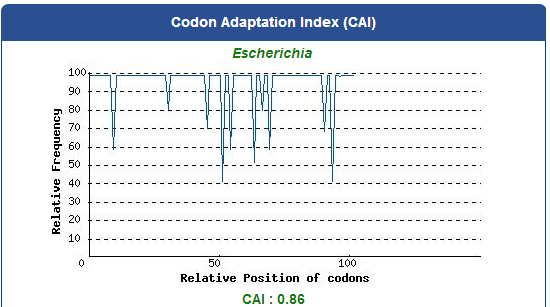
5. Add iGEM standard sequence in front of and at the back of our modified DNA sequence.
6. Synthesize the modified DNA sequence of PBANs in a gene synthesis company.
PCR experiment of PBAN

After receiving the DNA sequences from the gene synthesis company, we recombined each PBAN gene to PSB1C3 backbones and conducted a PCR experiment to check the size of each of the PBANs.
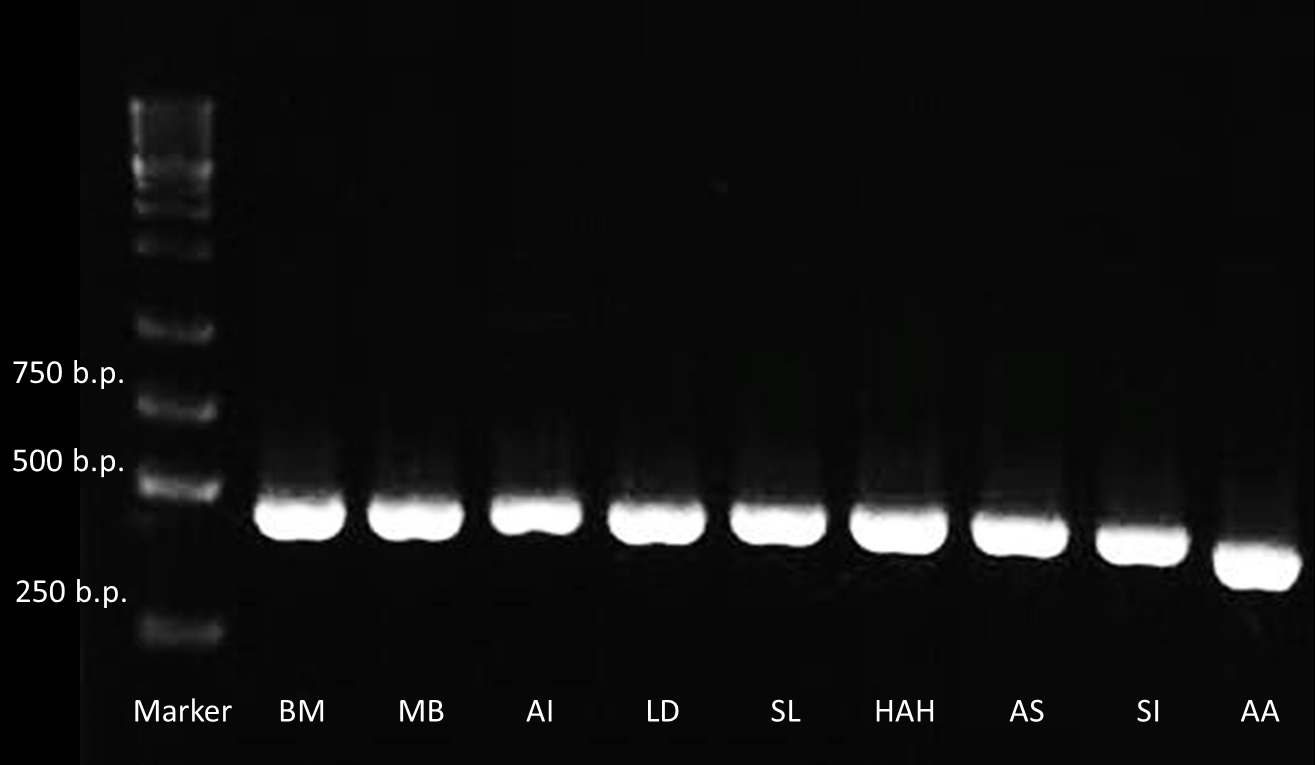
Below are biobrick serial numbers of PBAN abbrevation:
BM: BBa_K1415001 MB: BBa_K1415002 AI: BBa_K1415003 LD: BBa_K1415004 SL: BBa_K1415005 HAH:BBa_K1415006 AS: BBa_K1415007 SI: BBa_K1415008 AA: BBa_K1415009The DNA sequence length of the PBAN are around 100~150 bp. In this PCR experiment, the PBAN products size should be around 415~515 bp. Fig.2-1-3 showed the correct size of the PBAN, and proved that we successful ligated the PBAN DNA sequence onto an ideal backbone.
PBAN Peptide Check by SDS Protein Electrophoresis

Moreover, to verify that all 9 kinds of PBAN can be expressed by the E.coli, we conducted a SDS protein electrophoresis experiment. We first smashed the E.coli containing the PBAN with a sonicator and then took the supernatant divided from the bacterial pellet by centrifugation. Finally, we used the supernatant to run a SDS protein electrophoresis in a 20 % SDS gel.

Below are biobrick serial numbers of PBAN abbrevation:
BM: BBa_K1415001 AA: BBa_K1415009 LD: BBa_K1415104 AS: BBa_K1415007 SL: BBa_K1415005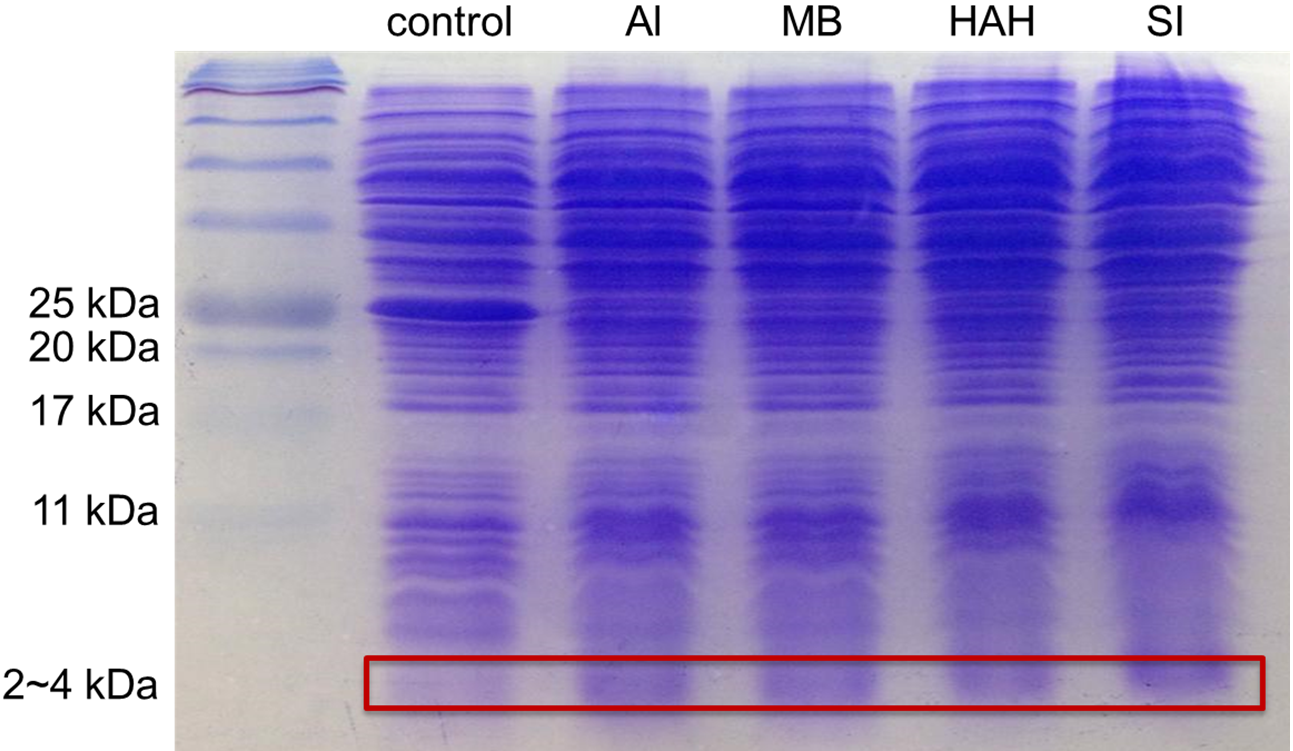
Below are biobrick serial numbers of PBAN abbrevation:
AI: BBa_K1415003 MB: BBa_K1415002 HAH:BBa_K1415006 SI: BBa_K1415008These SDS PAGE results in Fig.2-1-6 showed that the bands are at 2~4 kDa for each of the PBANs, while the plasmid of Pcons+RBS weren't there (the PBAN peptide is around 30 amino acids long). This result proves that the E.coli can produce the PBAN we chosen.
Blue Light Fluorescence / Bacteria Growth Test

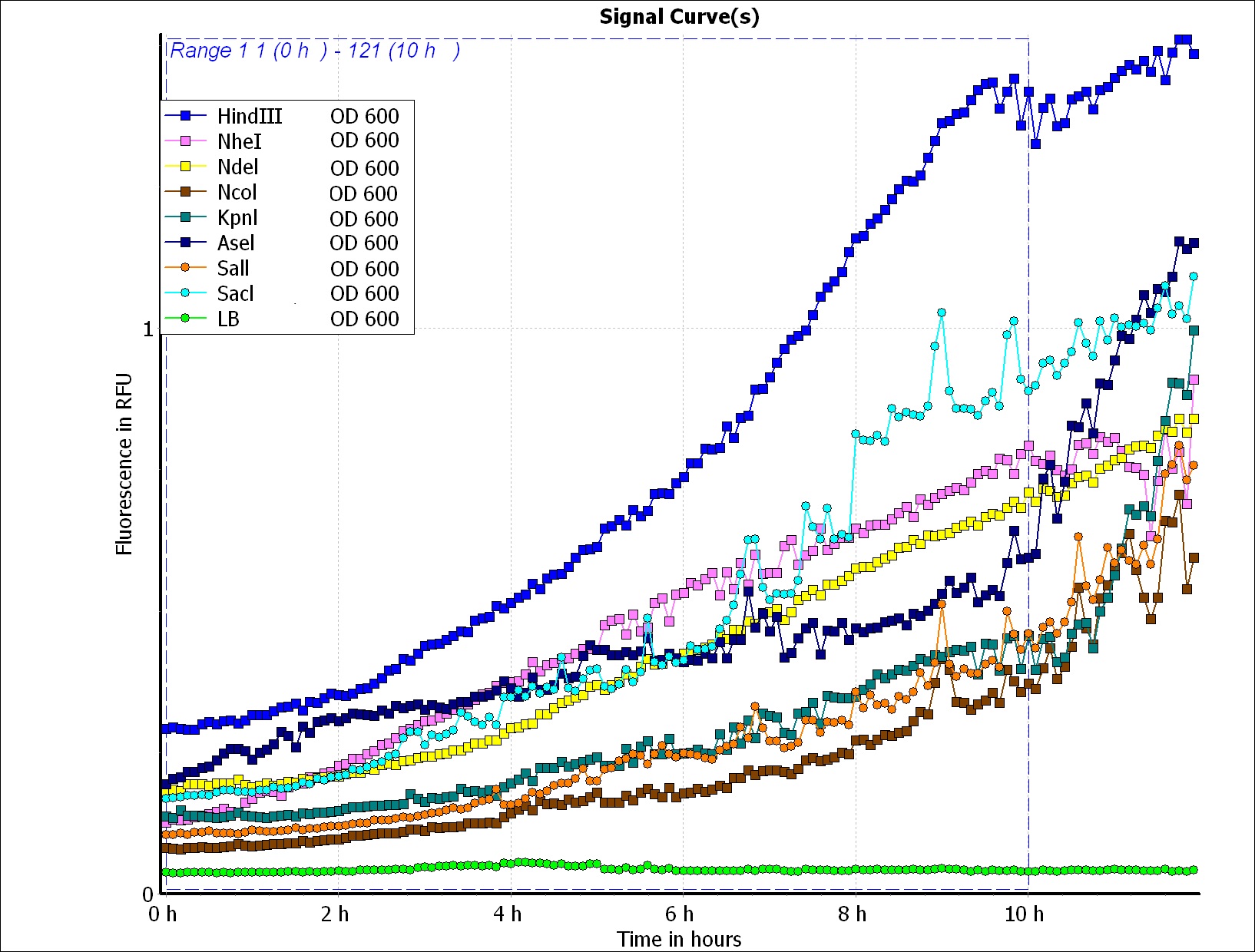
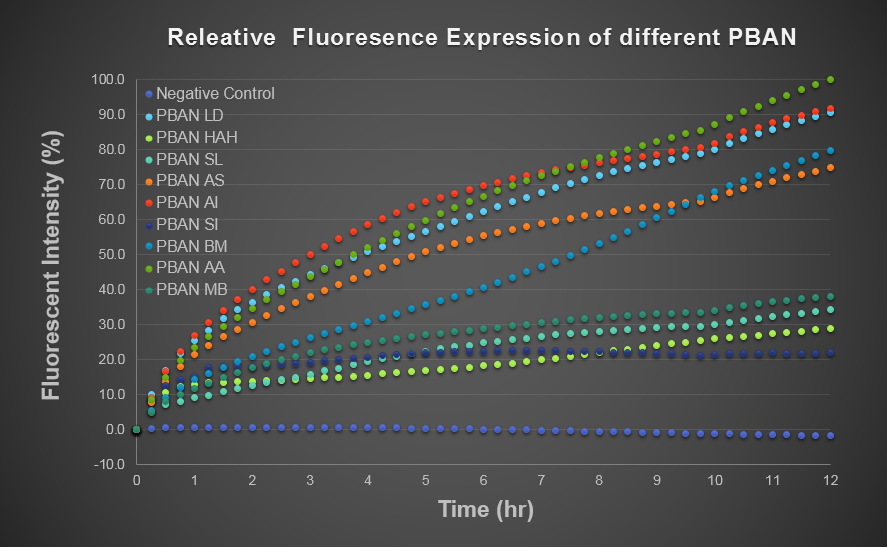
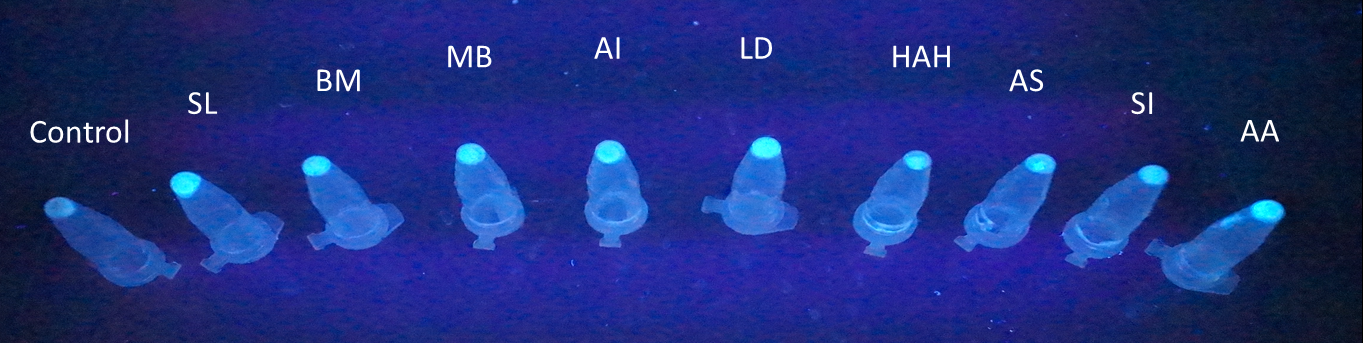
To predict the PBAN expression in E.coli by computer modeling, we next tested PBAN BFP biobricks. We obtained the average expressive value of the blue fluorescence in the biobrick part (above) and also the control part of Pcons + RBS + BFP + Ter. Therefore, we can use the average value to generate predictions of the PBAN expression in E.coli. (See more details in our Modeling Page). Below is the blue fluorescence expression curve and bacterial growth curve (OD 600) in a long period of time. We used these data to predict the PBAN expression in E.coli.
In Fig.2-1-5, we can clearly see that the blue fluorescence expressed by the E.coli is different from the control without BFP expressed.
Obtaining PBAN from E.coli
In order to obtain PBAN from our E.coli, we first culture the E.coli that contains our constructed plasmid of Pcons + RBS + One Kind of PBAN for 12 hr. Then, we smash the E.coli with a sonicator and centrifuged the solution to divide the PBAN from the cell debris pellets. Finally, we autoclave the supernatant to avoid biosafety issues. As we know, PBAN is a very simple and short peptide so it will not be degraded after the autoclave treatment. We also purified PBAN with HPLC from the supernatant ( after the autoclave treatment ) and diluted the pure PBAN powder with 1 liter of pure water for our PBAN effect test. A very small amount( pmol ) of PBAN can stimulate the maximum production of pheromone, therefore, we don't have to worry that our PBAN concentration will be inadequate after diluting with 1 liter pure water ( the actual concentration is 1.5mg/L ). This step was done for experimental purposes so that we could quantify the PBAN production and the effective concentration needed. After this experiment, in real-life application, we can simply use the autoclaved solution before the purification, following our recommended culturing condtitions.
Insect Tests
Behavior of Target Insects After PBAN Treatment
To investigate what behavior the female moth would show after ingesting PBAN, we put one female moth into a beaker for observation. The beaker is divided into two parts by plastic wrap. The bottom part contains the PBAN solution we prepared, and the upper part is the space for the moth to stay. We soaked cotton that spans the entire length of the beaker with the PBAN solution and sprinkle it with sugar. This way, the moth can suck on the PBAN without drowning in PBAN solution. After all the equipment is set, we put the female moth into the upper part of the beaker. At the time, we started filming as soon as we observed the female moth showing obvious behaviors of sexual stimulation such as flapping their wings. In this observation, the sample moths including Spodoptera litura, Mamestra brassicae and Helicoverpa armigera Hubner were caught in Sunny Morning organic farm.
We observed that the moth could absorb the PBAN in the solution through ingestion, and that the PBAN could stimulate the moth's pheromone gland to produce pheromone. As soon as the moth is sexually excited, it would flap its wings rapidly and move its tail slightly upward .
These movies show the behaviors of 3 different kinds of female moths after ingesting their separate PBANs. Each of the moths clearly became excited and all flapped their wings rapidly.
Effective Attraction after PBAN Treatment
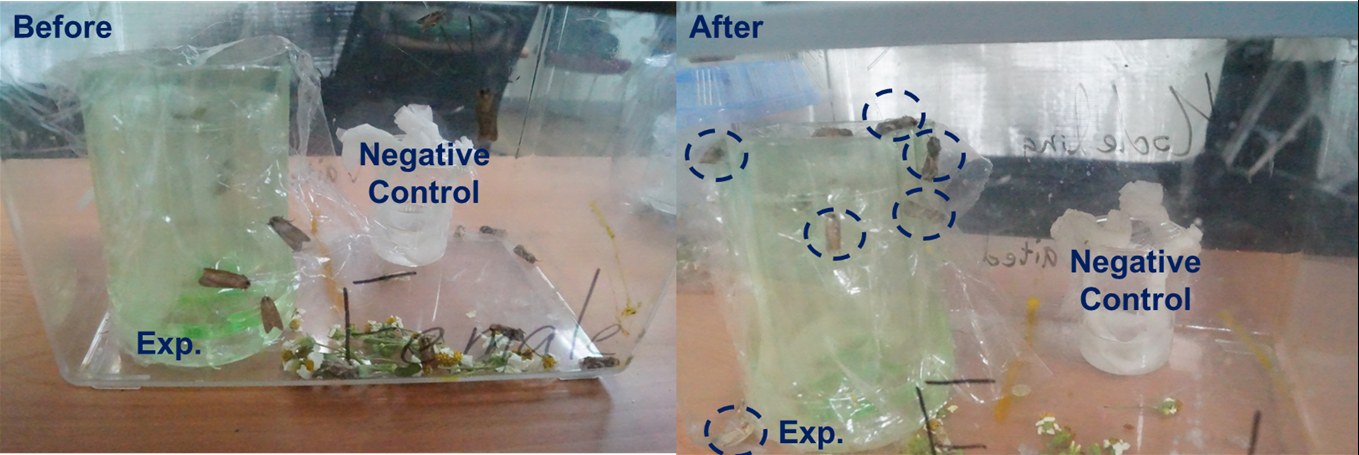
After observing the behaviors of female moth showed in PBAN treatment, we want to check the attractive effect of the moth. We expected that the female moth would not only become excited, flap its wings but also actually attract male moths to aggregate together after eating the PBAN. We used two beakers which are the same as what we used in the former experiment. One contained PBAN solution and the other contained only sucrose solution as control. We first put one beaker at one edge and the other at the opposite edge in a moth box (show in Fig.2-3-1). Then we put two female moths in each beaker and at least 100 male moths in the moth box. This time, we did a long time observation and took a picture with our camera. In Fig.2-3-1, the female moth ate the PBAN then attracted more male moths than the one eating sucrose solution. Thus, Fig.2-3-1 can prove the fact that the female moth ate our PBAN then release much sex pheromone to attract many male moths. In addition, we also conducted a simple test to compare the luring effect of female moths eating PBAN solution with the luring effect of female moths eating sucrose solution (the moth favorite food). Also, we can see the conspicuous effect again.
Spodoptera Litura Hobby for Temperature and Light
Light can be used to attract many kinds of harmful insects.
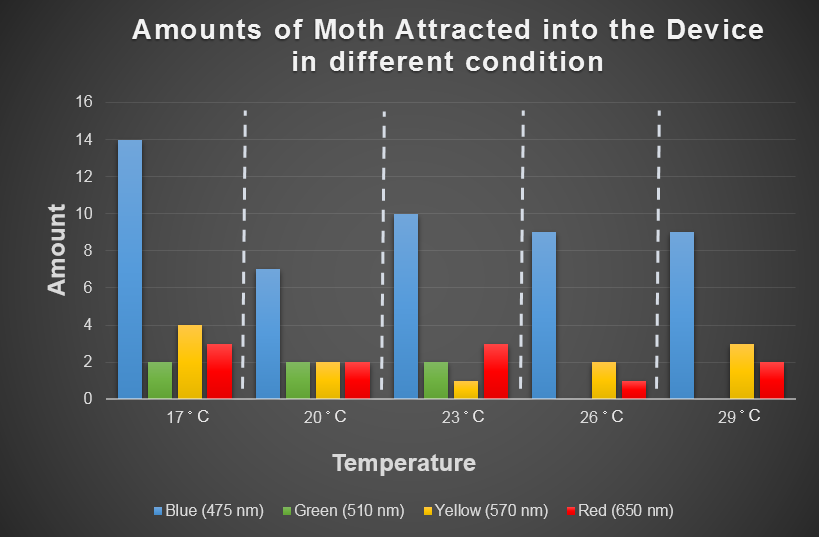
Temperature is the environmental factor which the farmer can not change practically. We want to use the computer modeling to deeply explore the relationship among light, temperature and the moths' hobby. In the future, we hope that farmers can choose the appropriate light according to temperature condition and even the kind of moths when using our device. For this, we chose the average temperature range in Taiwan in a year, and most common harmful insect, Spodoptera Litura to conduct this test (Fig.2-3-3 below), which we wanted to use to model the relationship among light, temperature and the moths' hobby with ANFIS (See detail in the device modeling page).
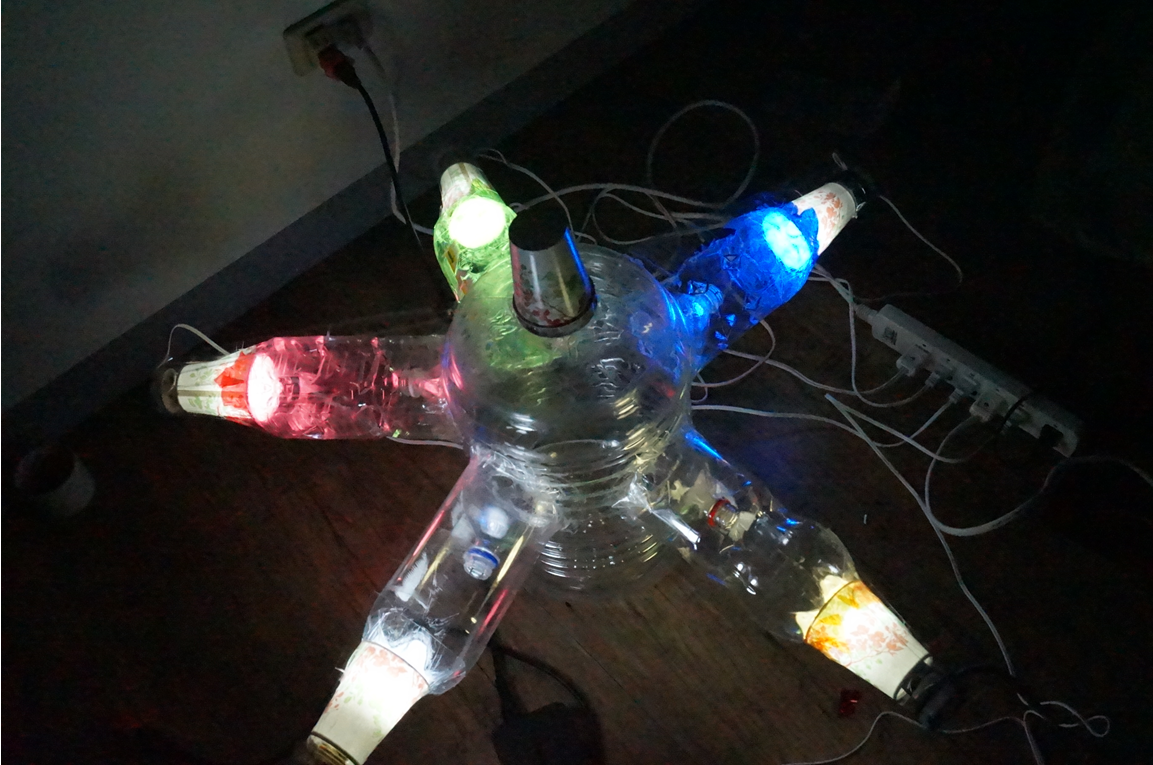

Fig.2-3-3 shows blue light have steady attraction to our target harmful moths, Spodoptera Litura, in any temperature condition. Thus, we decided to use blue LED light in our device design.
 "
"