Team:Paris Saclay/Notebook/August/11
From 2014.igem.org
(→Transformation of E.coli DH5 α by pJBEI+ApraR 2) |
(→Art & Design) |
||
| Line 92: | Line 92: | ||
''by Terry'' | ''by Terry'' | ||
| - | Incubation of a preculture of E. Coli with plasmid FNR RBS AmylCP (BB used in iGEM 2013 Paris-Saclay) coding for a blue chromoprotein. Will be used tomorrow for a test of colored bacteria density in agar. | + | Incubation of a preculture of ''E. Coli'' with plasmid FNR RBS AmylCP (BB used in iGEM 2013 Paris-Saclay) coding for a blue chromoprotein. Will be used tomorrow for a test of colored bacteria density in agar. |
{{Team:Paris_Saclay/notebook_footer}} | {{Team:Paris_Saclay/notebook_footer}} | ||
Revision as of 08:02, 19 August 2014
Contents |
Monday 11th August
Lab work
A - The chassis coli Odor free
by Melanie
Preculture odor free
C - Salicylate Inducible Suppressing System
Liquid Culture
by Fabio
4 liquid cultures of BBa_K1372001 with 20ml of LB and 20µl of antibiotic Cm (at 5pm - 150 rpm - 37 °C), from the stock made the 8th August.
D - Lemon scent
PCR Clean-up of BBa_K762100
by Sean
PCR prepared on the 7th August.
Clean-up performed with the following protocol.
NB: in the present case we have 90 µl of PCR result and we use 20 µl of elution buffer.
Digestion
by Laetitia and Hoang Vu
Check the replacement of Limonen synthase gene by Apramycin resistant gene in the pJBEI plasmid.
1)3 µL plasmid pJBEI (or plasmid pJBEI+ApraR 2) 0.5 µ BglII 1 µL fast digest buffer 10 µL H2O QSP
37°C - 1 hour Migration on agarose gel
Check if the restriction enzymes PacI from Sylvie's and from Alice's stock work.
1) 3 µL plasmid pJBEI (or plasmid pJBEI+ApraR 2) 0.5 µ PacI(Alice) 1 µL PacI buffer 10 µL H2O QSP
2)3 µL plasmid pJBEI (or plasmid pJBEI+ApreR 2) 0.5 µ PacI(Sylvie) 1 µL PacI buffer 10 µL H2O QSP
37°C - 1 hour Migration on agarose gel
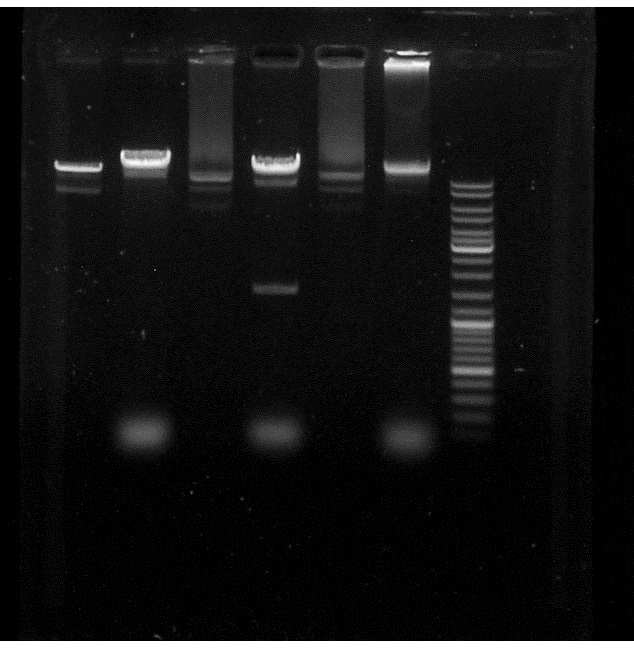
Conclusion: Alice's PacI works because we can see a supplementary band in the pJBEI+ApraR 2 compared to the control but not the Sylvie's. So, we do have the ApraR gene in our plasmid.
Transformation of E.coli DH5 α by pJBEI+ApraR 2
100µL competent bacteria 2 µL pJBEI+Apra 2
20 at 4°C 230' at 42°C 2 at 4°C
Then, we add 900µL of lysogeny broth and put it
1H at 37°C
In parallel, we did a control. (Without adding plasmid)
Then, we spread our E.coli on the petri dish containing Solid LB+Apra (1/2000) 100 µL control 100 µL transformed E.coli 200 µL transformed E.coli 100 µL concentrated transformed E.coli
Results: Nothing grew up. We think that the problem comes from the competent DH5 α that were not really competent...
Human Practices
Art & Design
by Terry
Incubation of a preculture of E. Coli with plasmid FNR RBS AmylCP (BB used in iGEM 2013 Paris-Saclay) coding for a blue chromoprotein. Will be used tomorrow for a test of colored bacteria density in agar.
 "
"