Team:Austin Texas/kit
From 2014.igem.org
(→Synthetase Efficiency) |
(→Synthetase Efficiency) |
||
| Line 199: | Line 199: | ||
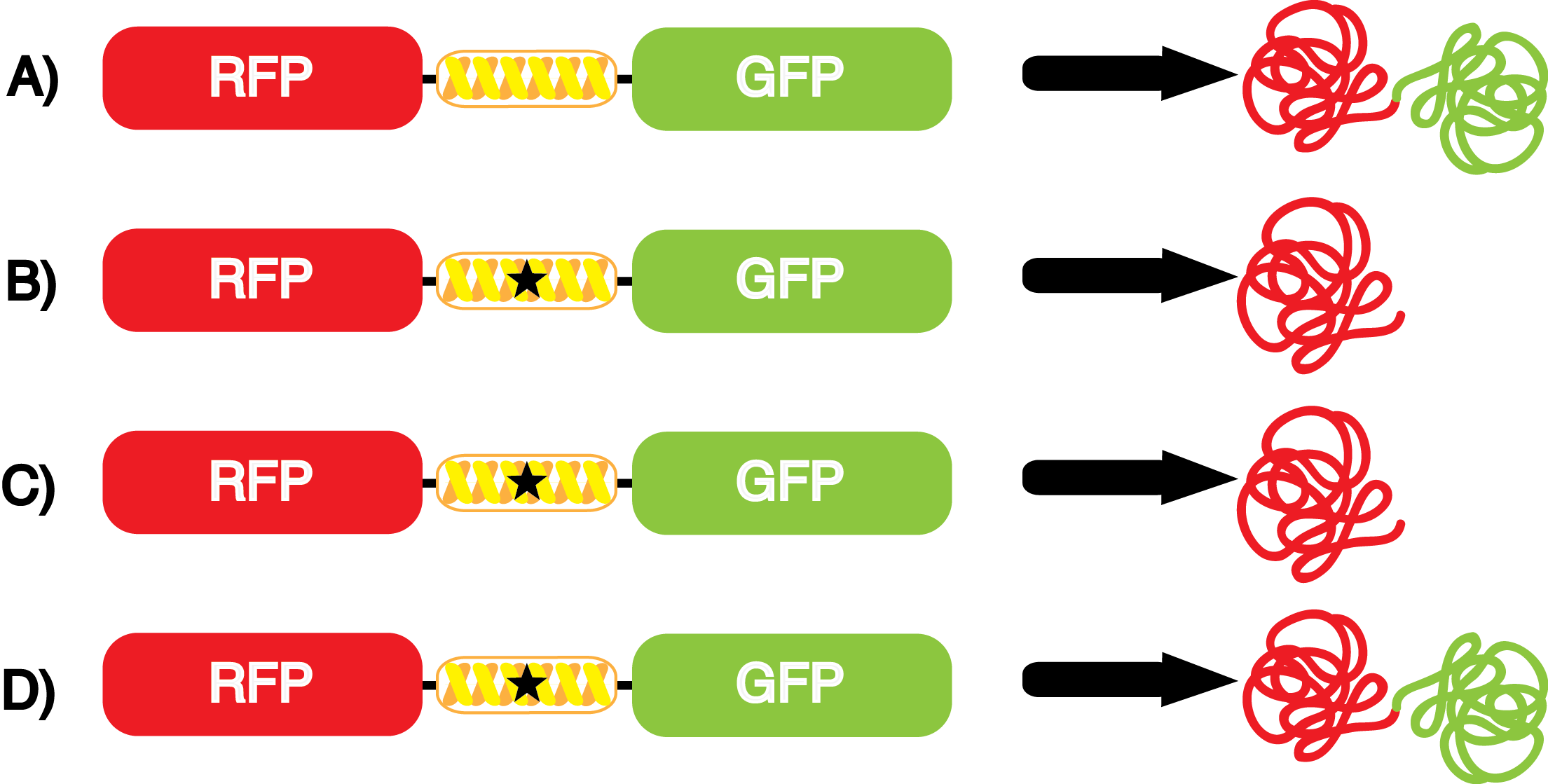
Another measure of quality for these ncAA synthetase/tRNA pairs is how efficiently they incorporate their ncAA. An inefficient synthetase/tRNA pair will only incorporate their ncAA a fraction of the time, even when their ncAA is present. Our system can be used to measure this level of efficiency, though it does not indicate why a particular synthetase is efficient or inefficient. By comparing the level of GFP fluorescence to the normalized level of RFP fluorescence when the amino acid is present, we can see how efficient the synthetase is. In essence, if the normalized fluorescence of GFP relative to RFP is close to 100% (if the GFP is expressed roughly 100% of the time that RFP is expressed), then the synthetase is very efficient. On the other side, if say the normalized fluorescence of GFP relative to RFP is closer to 10%, then the synthetase would not be very efficient, because even when the ncAA was there, it only incorporated it at the amber stop codon (UAG) about 10% of the time. | Another measure of quality for these ncAA synthetase/tRNA pairs is how efficiently they incorporate their ncAA. An inefficient synthetase/tRNA pair will only incorporate their ncAA a fraction of the time, even when their ncAA is present. Our system can be used to measure this level of efficiency, though it does not indicate why a particular synthetase is efficient or inefficient. By comparing the level of GFP fluorescence to the normalized level of RFP fluorescence when the amino acid is present, we can see how efficient the synthetase is. In essence, if the normalized fluorescence of GFP relative to RFP is close to 100% (if the GFP is expressed roughly 100% of the time that RFP is expressed), then the synthetase is very efficient. On the other side, if say the normalized fluorescence of GFP relative to RFP is closer to 10%, then the synthetase would not be very efficient, because even when the ncAA was there, it only incorporated it at the amber stop codon (UAG) about 10% of the time. | ||
| + | |||
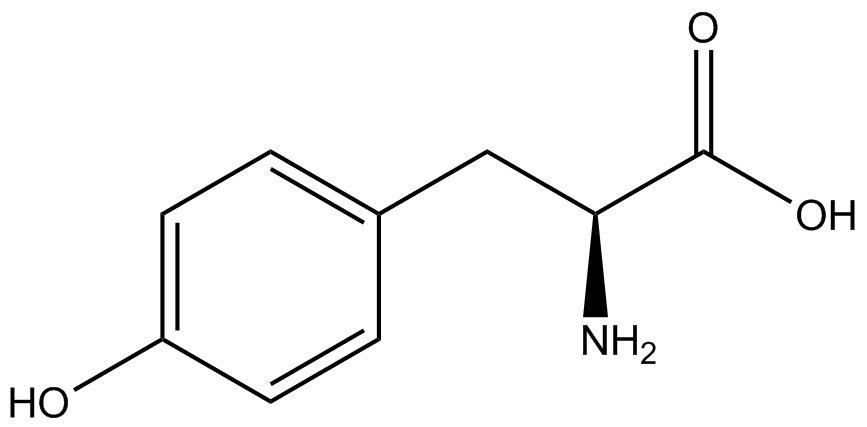
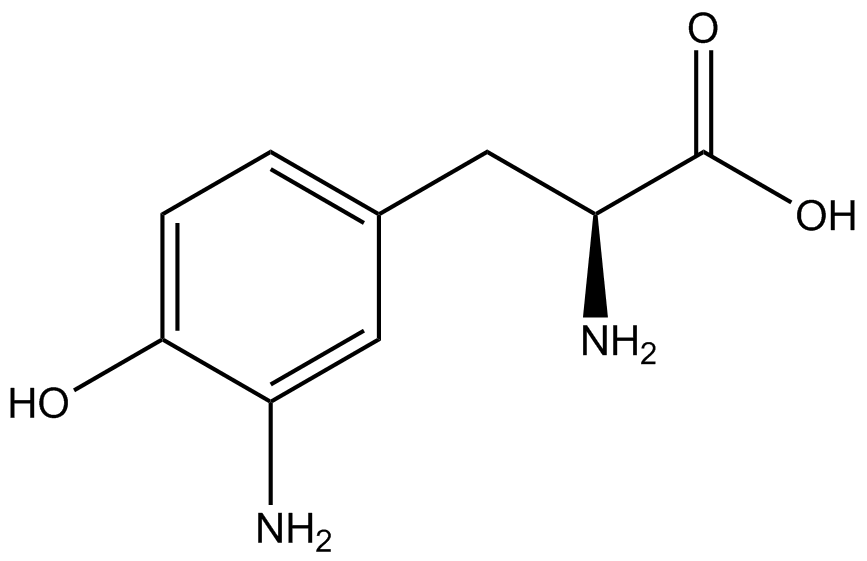
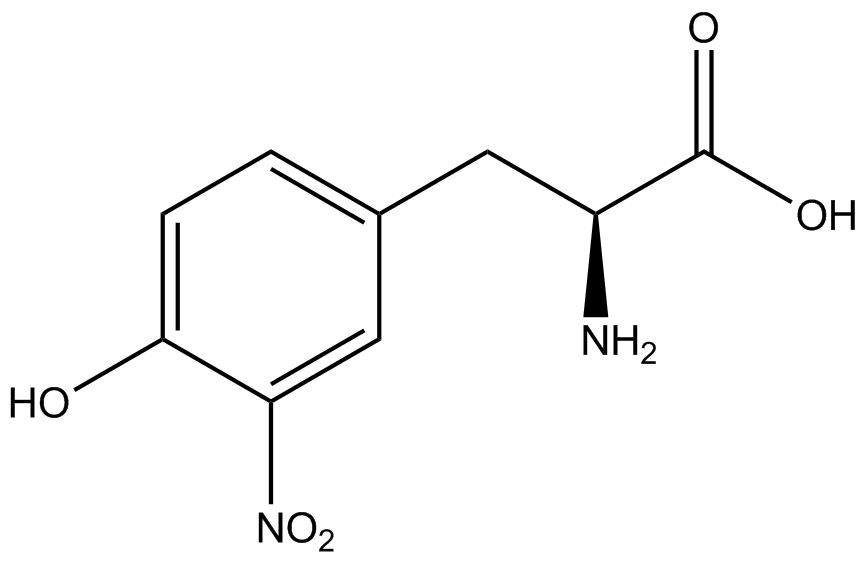
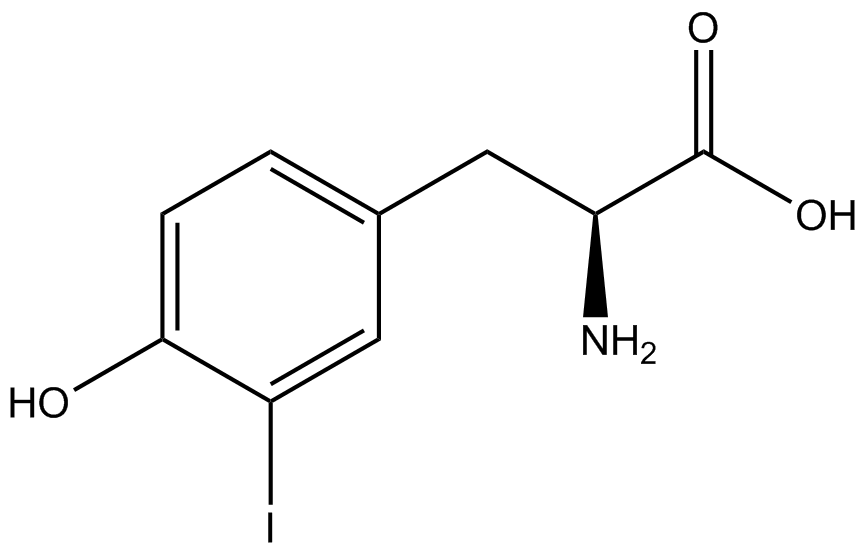
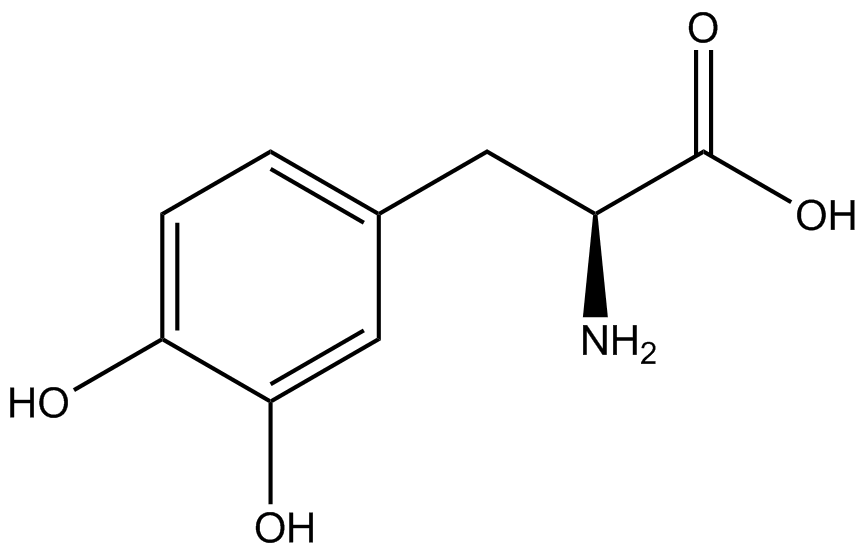
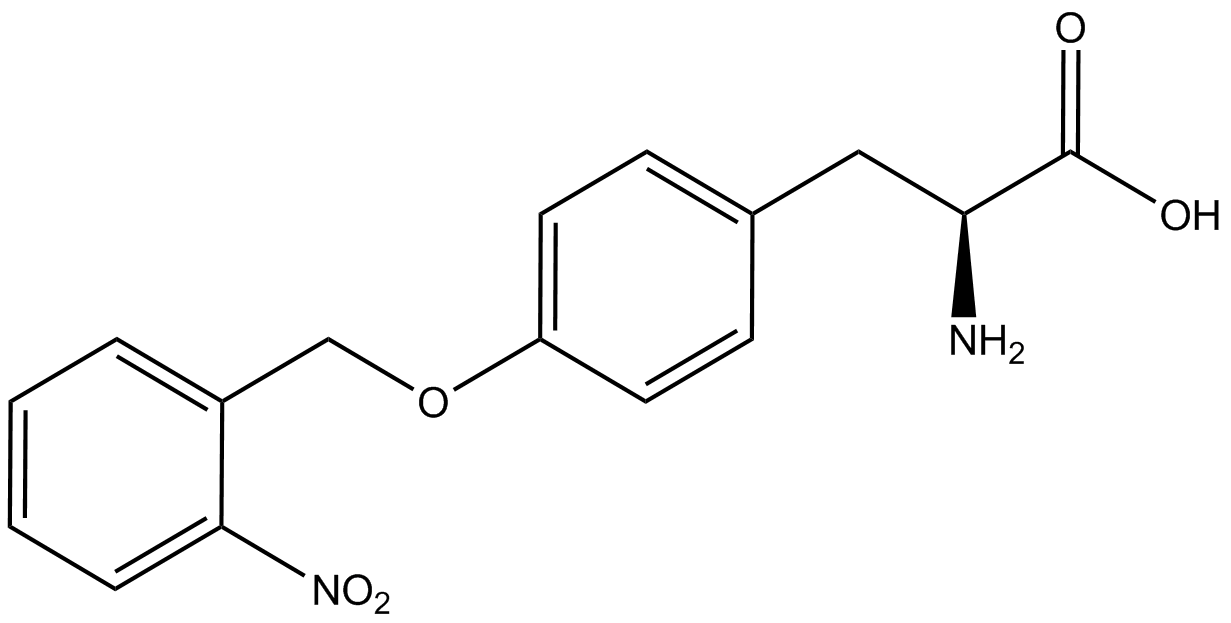
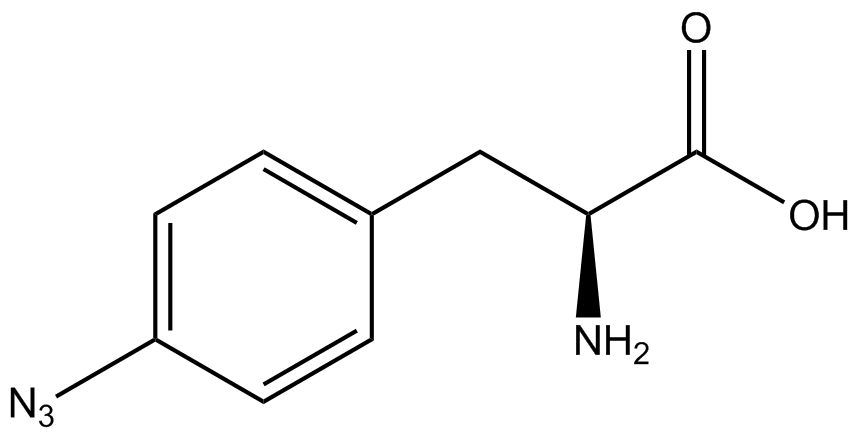
For our results (Figure 3), two synthesase/tRNA pairs stood out as relatively inefficient: 3-nitrotyrosine and ortho-nitrobenzyltyrosine. Both of these synthetase/tRNA pairs showed a significantly smaller normalized GFP to RFP fluorescence when the amino acid was present with the pFRY construct. While they both show a significant increase in normalized GFP to RFP fluorescence when the amino acid was present compared to when it was absent, which indicates a high fidelity, the actual GFP fluorescence relative to the RFP fluorescence was only around 50% for 3-nitrotyrosine and 20% for ortho-nitrobenzyltyrosine. These results suggest that for whatever reason, these synthetase/tRNA pairs do not always incorporate their ncAA at the amber stop codon. | For our results (Figure 3), two synthesase/tRNA pairs stood out as relatively inefficient: 3-nitrotyrosine and ortho-nitrobenzyltyrosine. Both of these synthetase/tRNA pairs showed a significantly smaller normalized GFP to RFP fluorescence when the amino acid was present with the pFRY construct. While they both show a significant increase in normalized GFP to RFP fluorescence when the amino acid was present compared to when it was absent, which indicates a high fidelity, the actual GFP fluorescence relative to the RFP fluorescence was only around 50% for 3-nitrotyrosine and 20% for ortho-nitrobenzyltyrosine. These results suggest that for whatever reason, these synthetase/tRNA pairs do not always incorporate their ncAA at the amber stop codon. | ||
Revision as of 01:32, 17 October 2014
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"