Team:Austin Texas/kit
From 2014.igem.org
(→Fidelity of Incorporation) |
(→Fidelity of Incorporation) |
||
| Line 185: | Line 185: | ||
[[File:UT_Austin_2014_Kit_Normalized_GFP_to_RFP_graph.png|thumb|600px|Figure 3. Graph showing the level of GFP fluorescence relative to RFP fluorescence. Each pStG plasmid is referred to based upon the tRNA synthetase/tRNA pair present in the specific plasmid. Each of these plasmids was then paired with either pFRY or pFRYC and grown in the presence or absence of a specific ncAA. For example, the "3-AminoY-FRYC" and the "3-AminoY-FRY" samples both contain the 3-AminoY synthetase/tRNA pair and both samples were grown in the absence or presence of the ncAA "3-AminoY". Data are presented as the average of three independent cultures. Error bars denote standard deviation.]] | [[File:UT_Austin_2014_Kit_Normalized_GFP_to_RFP_graph.png|thumb|600px|Figure 3. Graph showing the level of GFP fluorescence relative to RFP fluorescence. Each pStG plasmid is referred to based upon the tRNA synthetase/tRNA pair present in the specific plasmid. Each of these plasmids was then paired with either pFRY or pFRYC and grown in the presence or absence of a specific ncAA. For example, the "3-AminoY-FRYC" and the "3-AminoY-FRY" samples both contain the 3-AminoY synthetase/tRNA pair and both samples were grown in the absence or presence of the ncAA "3-AminoY". Data are presented as the average of three independent cultures. Error bars denote standard deviation.]] | ||
| - | '''Set the experiment up - What do we want to test? How are we going to test it (keep it simple)?''' | + | '''Set the experiment up - What do we want to test? How are we going to test it (keep it simple)? You may just need to rearrange the paragraph.''' |
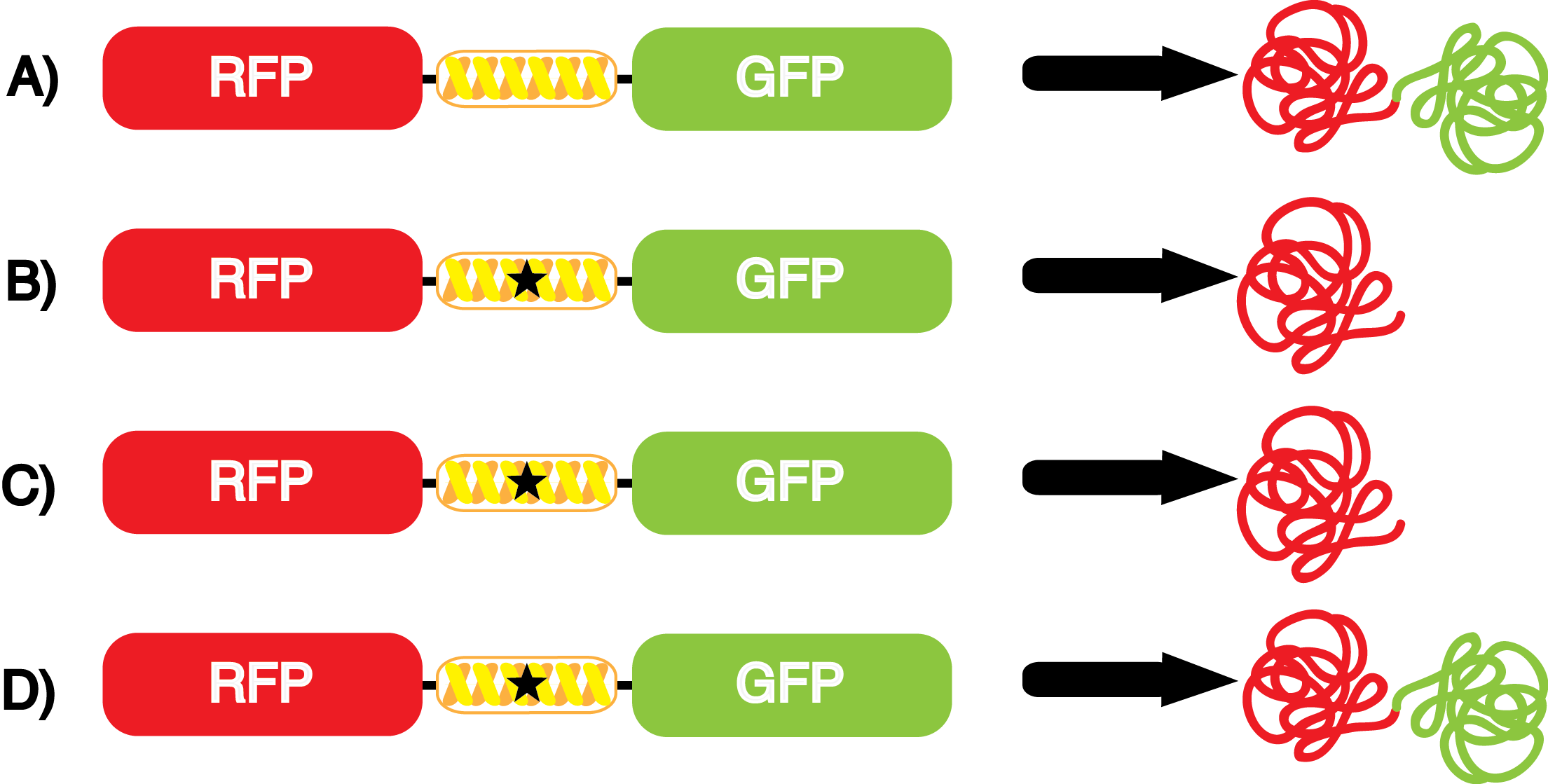
| - | The fidelity of each ncAA sythetase/tRNA pair was measured by comparing the | + | The fidelity of each ncAA sythetase/tRNA pair was measured by comparing the production of GFP in cultures containing pStG/pFRY with or without ncAAs. In the absence of an ncAA only RFP should be translated, as translation is expected to terminate between RFP and GFP at the amber stop codon (UAG) on the linker sequence in pFRY. Alternatively, if the corresponding ncAA for pStG is present or if the synthetase/tRNA pair has low fidelity and can misincorporate a different amino acid, translation should continue through the UAG. In this case, RFP and GFP should both be translated. The amount of fluorescence from RFP and GFP is detectable by fluorometer.'''I don't think you need to mention the fluorometer here... methods''' We also tested pStG/pFRYC strains in (+/-) ncAA conditions as a control, '''control for what? for how fluorescence/growth changes with ncAA''' which should express RFP and GFP in all conditions since pFRYC does not have an amber stop codon in the linker (Figure 1). However, if the presence of ncAA affects cell growth or fluorescence activity, we will need these controls to determine the extent of the effect. |
Revision as of 20:53, 16 October 2014
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"