Team:Austin Texas/kit
From 2014.igem.org
(Difference between revisions)
Jordanmonk (Talk | contribs) (→Background) |
Jordanmonk (Talk | contribs) (→Experimental Design and Method) |
||
| Line 87: | Line 87: | ||
==Experimental Design and Method== | ==Experimental Design and Method== | ||
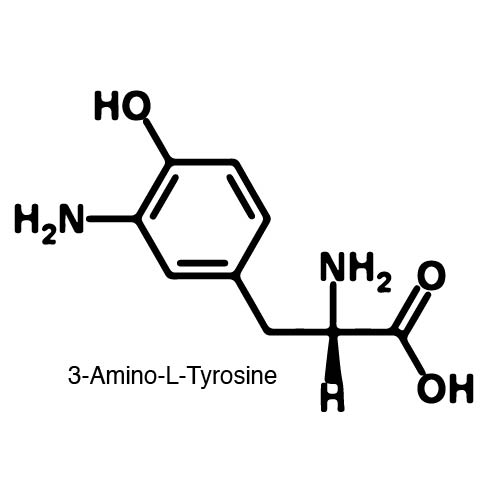
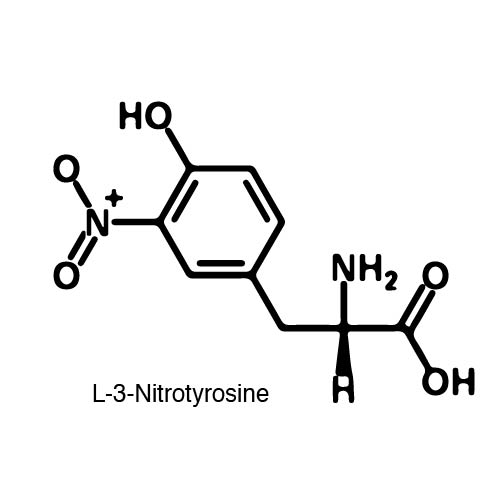
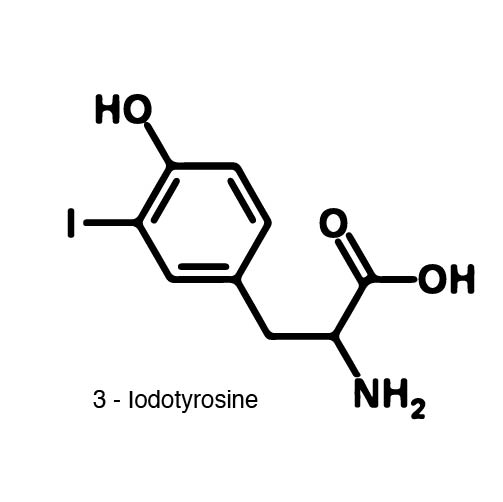
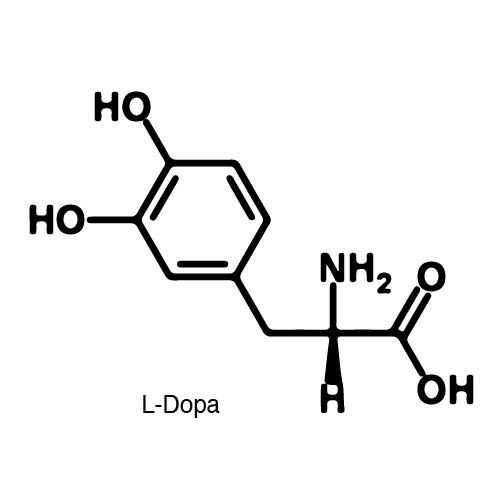
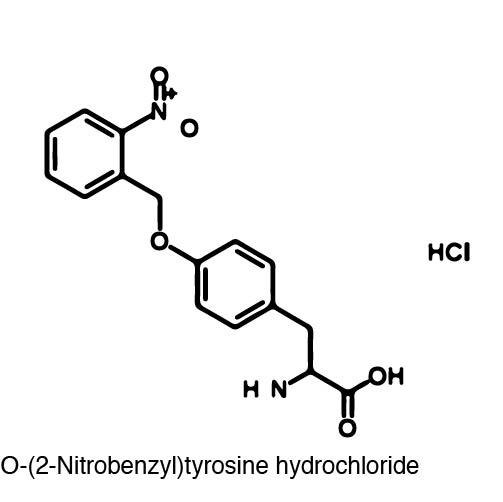
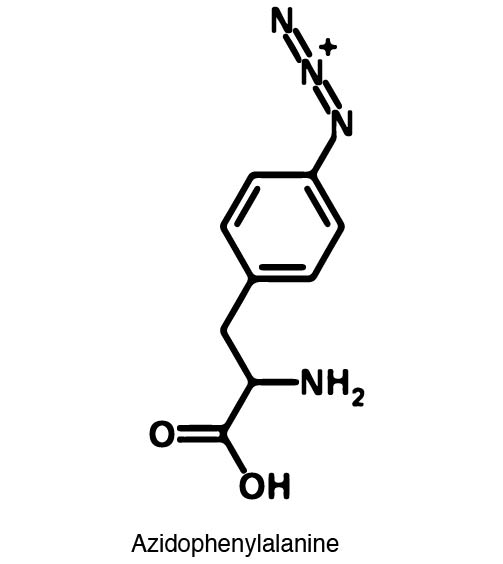
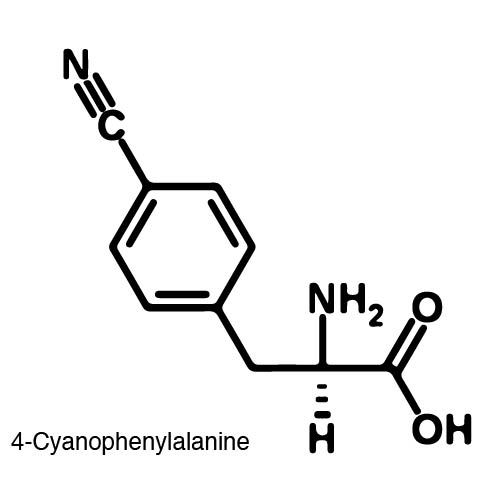
| - | In order to | + | In order to recode UAG, a synthetase/tRNA pair much be modified to effectively grab onto an ncAA. Various methods of directed evolution are typically used to modify a synthetase such that it can accept and charge a non-canonical amino acid. The library of ncAA synthetases available have ranging levels of reported efficiency and are not well characterized. This year the UT iGEM Team created a test kit designed to characterize the efficiency of any ncAA synthetase/tRNA pair. |
| Line 93: | Line 93: | ||
| - | The kit consists of a three plasmid system: pBLG, pFRYC, and pFRY. pBLG contains | + | The kit consists of a three plasmid system: pBLG, pFRYC, and pFRY. pBLG contains the ncAA synthetase/tRNA pair to be tested as well as a gentamicin resistance gene. pFRYC is the control plasmid and contains the IPTG-induced reporter system and a kanamycin resistance gene. The reporter system is composed of RFP and sfGFP fused via a linker sequence between the two. pFRY is the experimental plasmid and is nearly identical to pFRYC with the exception that its linker sequence contains an amber stop codon in the middle of the linker whereas pFRYC contains a codon for tyrosine in the same location. In a cell containing pFRYC, the ribosome will translate the RFP reporter, linker, and finally sfGFP, producing red and green fluorescent proteins that result in visible yellow fluorescence. In a cell containing pFRY, the ribosome will translate the sfGFP and terminate at the amber stop codon on the linker producing a green fluorescence. When pBLG and pFRY are present in the cell, the ribosome will incorporate an ncAA at the amber codon in the linker and continue translation producing both RFP and sfGFP reporters if the synthetase/tRNA pair encoded on pBLG effectively incorporate the ncAA. |
[[File:NcAA Kit Plasmid Alignment 10-11-14.png|500px|thumb|right|Figure 2]] | [[File:NcAA Kit Plasmid Alignment 10-11-14.png|500px|thumb|right|Figure 2]] | ||
| Line 99: | Line 99: | ||
| - | An ncAA synthetase/tRNA pair was cloned into pBLG and transformed into | + | An ncAA synthetase/tRNA pair was cloned into pBLG and transformed into pFRYC amberless ''E. coli''. and pFRY amberless ''E. coli''. Other necessary control strains include RFP amberless ''E. coli'' (RFP control), sfGFP amberless ''E. coli'' (GFP control), amberless ''E. coli'' (cell background control), and LB media supplemented with ncAA ('''was it supplemented?''') (media background control). An overnight culture of each strain was grown in LB with the appropriate antibiotics at 37ºC and 225rpm. 10 mL of media with the appropriate antibiotics was inoculated with 100 µL of overnight culture and allowed to grow in the same conditions until the culture density was ~0.2-0.3 OD, or ~3 hours. The 10 mL culture was split between 4 different sterile test-tubes, 2 mL of culture per tube. The conditions of test tubes A through D were as follows: A (-IPTG,-ncAA), B (-IPTG,+ncAA), C (+IPTG, -ncAA), and D (+IPTG, +ncAA). IPTG stock solution was made at 1000X concentration ('''?''') and the ncAA was added to yield a concentration of 1 mM. Sterile deionized water was added in the place of ncAA and IPTG as a control ('''?'''). Once the controls, IPTG, and the ncAA were added appropriately, the cultures were allowed to grow to ~0.5 OD. 70 µL of each culture condition and control culture was added to a separate wells in a transparent 96-well plate for fluorescence and OD readings in a microplate reader. |
Revision as of 06:01, 12 October 2014
|
 "
"