Team:Oxford/notebook
From 2014.igem.org
| Line 90: | Line 90: | ||
Following an overnight 8hr Gibson Assembly the reaction volumes were treated with Dpn1 restriction enzyme that cuts bacterial (methylated) DNA. | Following an overnight 8hr Gibson Assembly the reaction volumes were treated with Dpn1 restriction enzyme that cuts bacterial (methylated) DNA. | ||
We transformed the Gibson products into chemically competent DH5-alpha cells as well as into NEB alpha-5 cells. Unfortunately no colonies grew on a KanR plate! | We transformed the Gibson products into chemically competent DH5-alpha cells as well as into NEB alpha-5 cells. Unfortunately no colonies grew on a KanR plate! | ||
| - | We know that the PCR products are correct so we think an issue may have arisen during the Gibson Assembly stage so we will re-do this part next week. | + | We know that the PCR products are correct so we think an issue may have arisen during the Gibson Assembly stage so we will re-do this part next week.<BR><BR> |
<h2> Week 2 </h2> | <h2> Week 2 </h2> | ||
| Line 102: | Line 102: | ||
<font style="font-weight:bold">Notebook:</font> | <font style="font-weight:bold">Notebook:</font> | ||
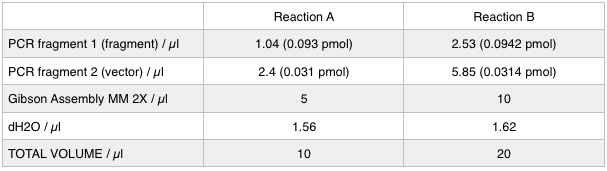
| - | To run equimolar amounts of insert to vector the following table was calculated: < | + | <p>To run equimolar amounts of insert to vector the following table was calculated: </p> |
| - | [[File:Oxford_iGEM_B_2_1.jpg]] | + | </html>[[File:Oxford_iGEM_B_2_1.jpg|centre]]<BR><html> |
| + | <p>These gibson assemblies were then transformed into E.coli cells using electroporation with the voltage at 1.8kV and the exposure time ranging from 4.8ms to 5.2ms. The cells were then recovered at 37ºC for 1.5 hrs before 100 µl was plated onto KanR plates. These were left overnight at 37ºC.</p> | ||
| + | <p>Unfortunately, there were no colonies grown and as we have no product from our gibson assemblies remaining we will have to repeat the process from Week 1 in order to trouble-shoot the method and get the desired plasmid. It is possible that we accidentally swapped the primers or PCR products around at some point and thus we had 1A with 2B and 1B with 2A in the Gibson assembly. These combinations are not complementary and thus the Gibson would fail meaning that none of our bacteria received intact Kanamycin resistance in the transformation. However there are many other reasons why the process may have failed therefore we went through the entire method trying to cover all instances where we may have made an error.</p> | ||
| + | <p>First we confirmed that the primer sequences on the delivery tubes matched what we had ordered. We ran the PCR as before; we re-calculated the annealing temperatures and extension times with the same result as last week.</p> | ||
| + | <p>In week one, we did the <a href="https://2014.igem.org/Team:Oxford/protocols/DpnI digest">DpnI digest</a> after the Gibson assembly and not directly after the PCR. This time we did the digest immediately after the PCR to ensure that all template DNA was destroyed before running on the gel, ensuring cleaner bands on our gel and less contamination from non-PCR products.</p> | ||
| + | <p>We loaded 20 μl of each PCR product onto a 0.8% agarose gel as before. The resulting bands were visualized after staining as shown below:</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_2.jpg|centre]]<BR><html> | ||
| + | <p>The bands are not as bright as in the first attempt probably due to less time spent in the ethidium bromide staining tank. The bands correspond with the correct sized fragments as before. This suggests there was no error in the PCR reaction with the exception of again mixing up the primers (the PCR would be successful with any combination of the primers for a 1 and 2 reaction). The bands are cleaner allowing us to try an alternative to the gel extraction in case that caused our assembly and transformation to fail.</p> | ||
| + | <p>Next we excised and extracted our bands from the gel with the same <a href="https://2014.igem.org/Team:Oxford/protocols/QIAquick_Gel_Extraction">QIAquick Gel Extraction Protocol</a> protocol as before but with two changes. First we allowed additional drying time after the step involving ethanol supplemented buffer in case ethanol contamination inhibited the enzymes used in the Gibson assembly later in the process. We also replaced the 20 μl EB buffer with 20μl of MilliQ water. We did our second elution with the kit EB buffer. We then measured the concentration of DNA in each eluted sample using the nanodrop. This still gives us a reasonable idea (despite the interference from the remaining QG buffer residue) as to the amount of DNA in each sample from which to calculate our dilutions for the Gibson assembly reaction.</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_3.jpg|centre]]<BR><html> | ||
| + | <p>We used the remaining 10μl of our PCR products to do a Promega PCR clean-up and eluted using water. We gathered the following Nanodrop data:</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_4.jpg|centre]]<BR><html> | ||
| + | <p>Next we ran the following Gibson assembly reactions:</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_5.jpg|centre]]<BR><html> | ||
| + | <p>Next we transformed the Gibson assembly products into chemically competent E.coli cells. We decided to run a positive control for each kind of cell used using the pUC19 plasmid provided with the cells. Unfortunately we were limited with the amount of cells available and had to use two different cell strains; this resulted in some variation in the length of the first incubation with the DNA on ice as we began the transformation in the morning but then decided to pause the process so that the cells would be plated out later in the day in order to not have too much growth overnight. The NEB 5-alpha cells were also on ice slightly longer before the DNA was added.</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_6.jpg|centre]]<BR><html> | ||
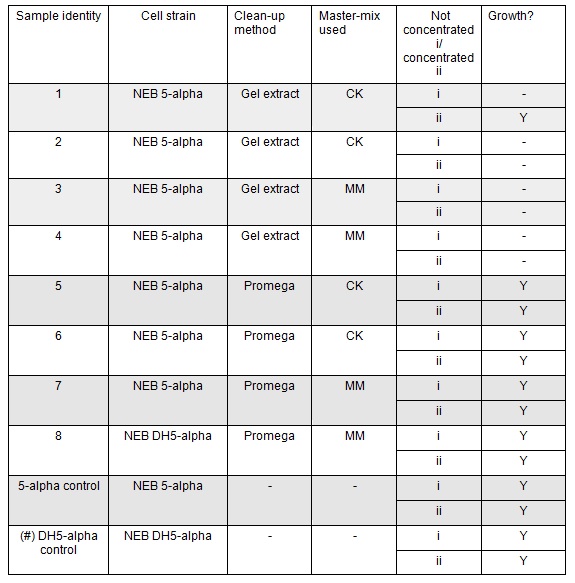
| + | <p>The cells were plated out after 1.5 hour incubation. It was noted that one of the sample appeared to have leaked from the Eppendorf into the petri dish containing all the samples in the incubator. We think it was the 5-alpha control as this was the only sample that appeared to have lost volume. Samples 1-8 were plated on Kanamycin plates (i) that we made up during the final incubation period. The remaining sample was re-suspended in ~100μl of SOC to create a concentrated solution of cells that were also plated out on kanamycin plates (ii). The two controls were spread on Ampicillin plates made up by Nick. All plates were placed in the 37⁰c incubator overnight. The results in the morning were as follows:</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_7.jpg|centre]]<BR><html> | ||
| + | <p>Apart from the concentrated sample 1ii, none of the samples prepared using the gel extraction protocol grew. Sample 1ii also had limited growth compared to the other plates with colonies:</p> | ||
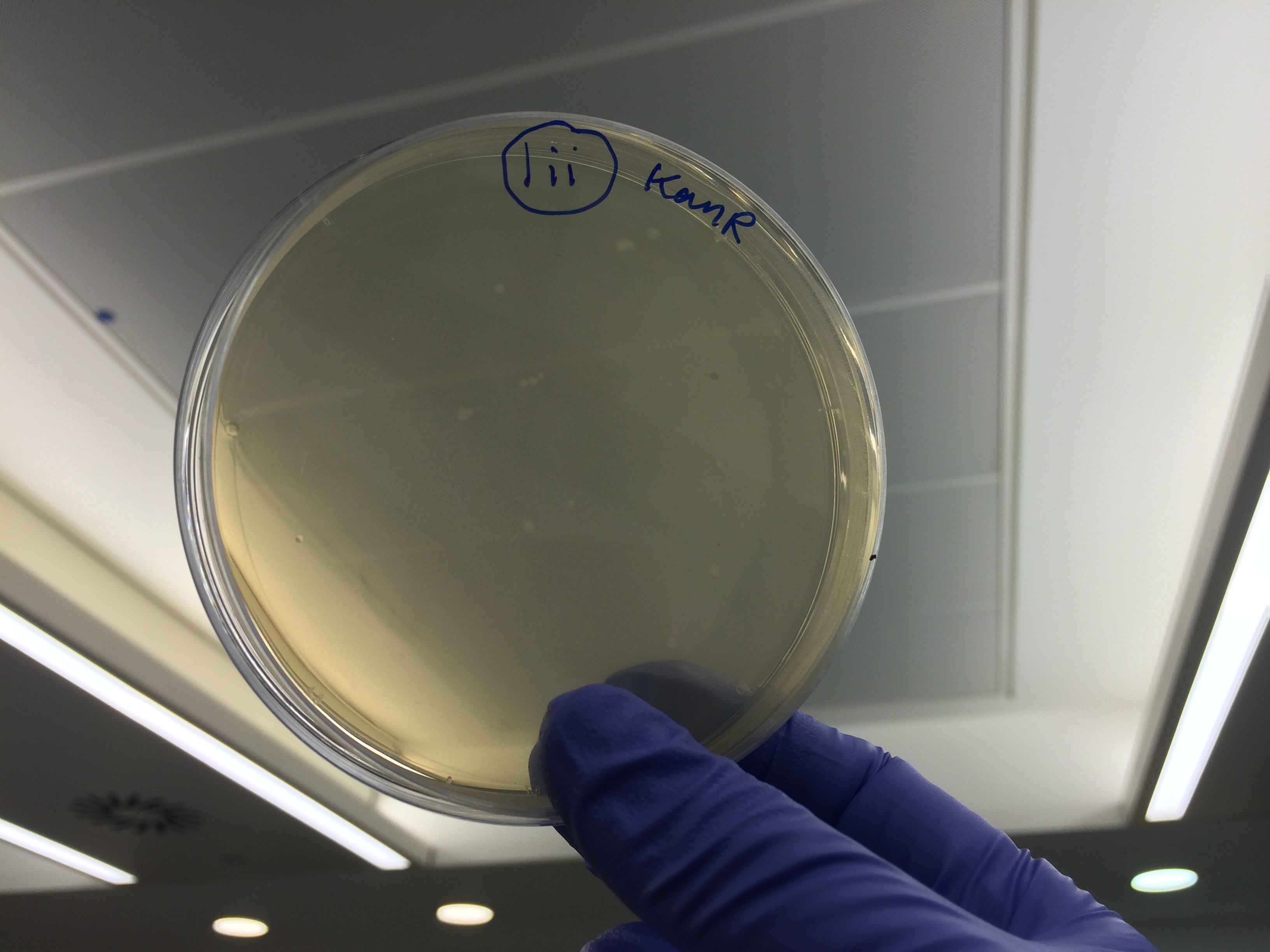
| + | </html>[[File:Oxford_iGEM_B_2_8.jpg|500px|centre]]<BR><html> | ||
| + | <p>Samples 1i and 5i differed only in their clean-up method; 5i has lots of growth and 1i has none. This comparison holds true for Samples 2+6, 3+7 and 4+8 with the same result:</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_9.jpg|500px|centre]]<BR><html> | ||
| + | <p>This suggests that we should use the Promega PCR clean-up kit and not the QIAquick Gel extraction. We will still run our samples on a gel to check the quality of our PCR reactions. We may investigate the Promega Gel extraction kit as this will enable us to pick our desired band out of a PCR that has also generated non-specific fragments which we cannot do with the Promega PCR clean-up protocol. </p> | ||
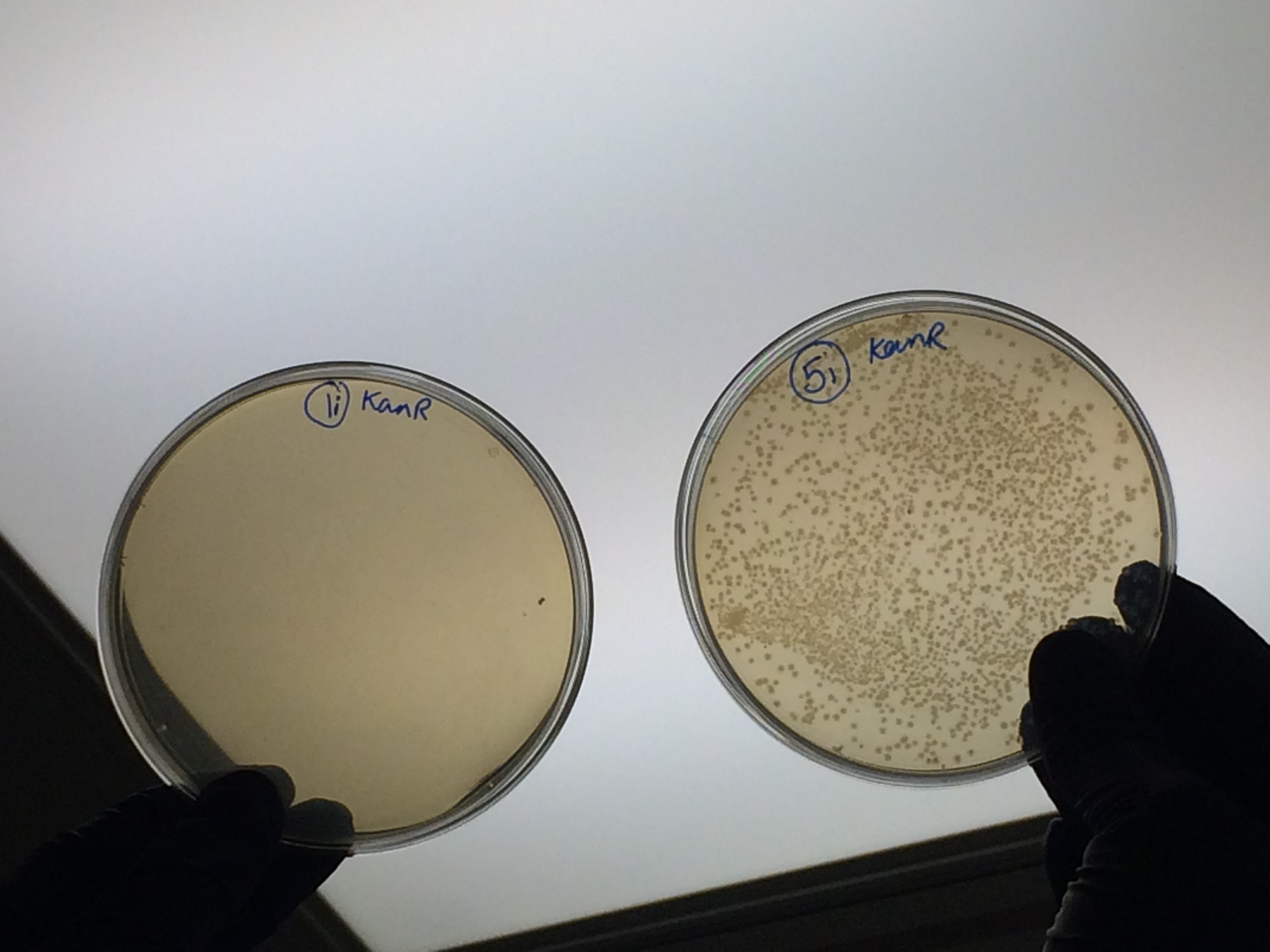
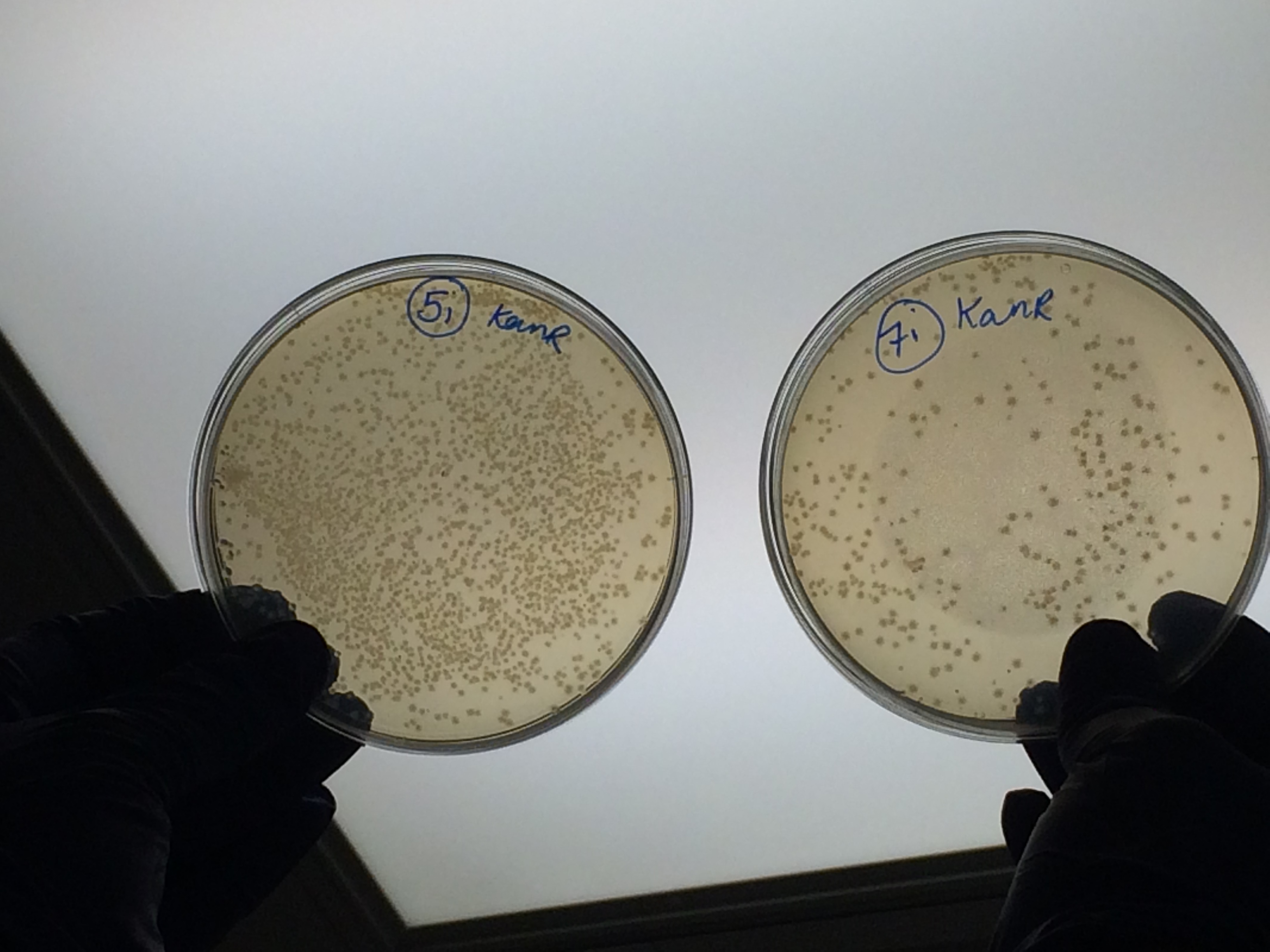
| + | <p>The samples that were treated with the master-mix and ligase supplied by Ciaran had more colonies than those that were treated with the NEB supplied master-mix as shown by the comparison between samples 5i and 7i:</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_10.jpg|500px|centre]]<BR><html> | ||
| + | <p>This suggests that in future we should use the master-mix supplied by Ciaran.</p> | ||
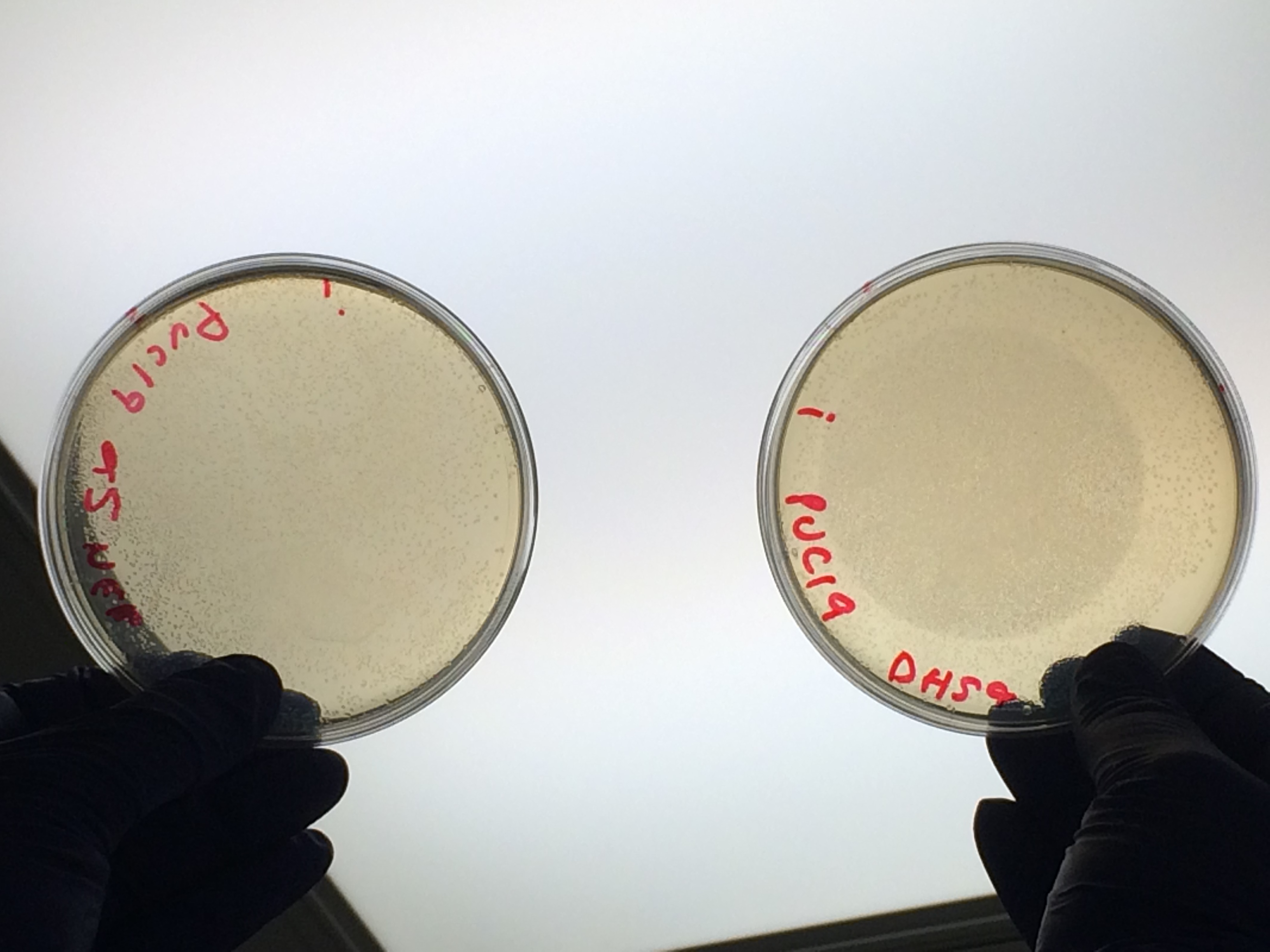
| + | <p>All the controls grew showing that the lack of growth of samples 1- 4 was not due to inefficient transformation:</p> | ||
| + | </html>[[File:Oxford_iGEM_B_2_11.jpg|500px|centre]]<BR><html> | ||
| + | <p>There was no observable difference between sample 8 and sample 7 which were treated the same but which used different chemically competent cells. Thus we can continue to use wither the NEB 5-alpha or DH5-alpha chemically competent cells. </p> | ||
| + | <p>There was no observable difference between the samples that correspond to A Gibson assembly reactions (template for PCR primers and therefore fragment = pCM66) and those that correspond to B Gibson assembly reactions (PCR primers and therefore fragment include Pamp).</p> | ||
| + | <p>We will confirm that the plasmid in each plate of transformed cells is the desired product by growing up the cultures overnight and extracting the plasmid for sequencing. We are fairly confident that it is the correct product as the fragment we were inserting was the kanamycin resistance itself which is necessary for growth on the kanamycin plates.</p> | ||
| + | |||
Revision as of 14:02, 24 July 2014
Lab Book
Part A
Week 1
Tasks completed:
testing the resistance of the plastic ware and the tolerance of E. coli, Pseudomonas fluorescens and Pseudomonas putida to DCM.
Methods:
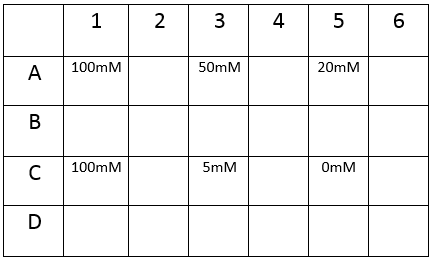
First, we tested the resilience of the plastic ware, specifically the 24 well plates, to find out what concentrations of DCM it could cope with.
The plate was set up with the following concentrations of DCM in each well.
After leaving the plate overnight covered in foil and the lid to prevent the evaporation of DCM, all wells were intact. This means we can use the plates in our further experiments with DCM.
To test the tolerance of the bacteria we grew up liquid culture strains using the
Growing Liquid Cell Cultures protocol. The strains we used were KT2240 (P. putida), SBW25 (P. fluorescens) and MGG155 (E. coli). Initially we incubated 100µl of each liquid culture with 5ml LB broth and DCM to make concentrations of 0, 5, 10 and 20mM. Overnight incubations of each strain at an appropriate temperature (30°C for ''Pseudomonas'' and 37°C for'' E. coli'') whilst shaking showed that all three strains could tolerate these concentrations, indicating that it’s the metabolic intermediates and not the DCM itself that causes the toxicity.
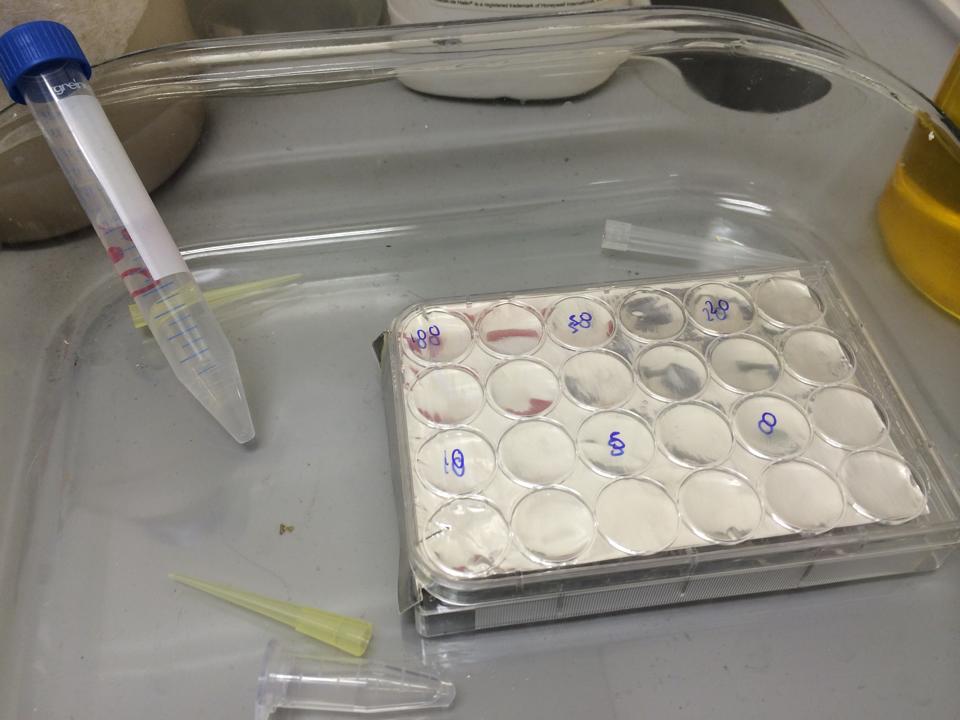
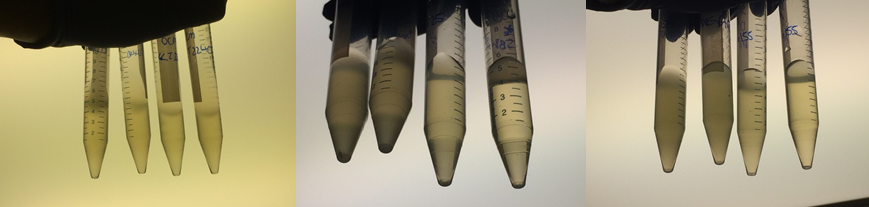
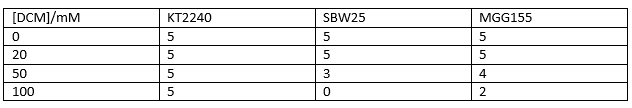
Therefore we repeated the experiment using 0, 20, 50 and 100mM concentrations of DCM. Before doing this we streaked out fresh plates of our cultures using the Agar Plate Preparation and Streaking Plates protocols before growing liquid cultures from these. The results of P. putida, P. fluorescens and E. coli overnight incubation are below:
We ranked the growth in each tube by eye where 0 was no growth and 5 was unhindered growth.
P. putida could tolerate even 100mM, so this strain will be useful when it comes to characterising the dcmA regulatory system. As we now have an idea of what concentrations each strain can tolerate, we can design our experiments for next week where we will obtain growth curves for each strain at various concentrations of DCM.
Part B
Week 1
Tasks completed:
First attempt at swapping resistance gene for plasmid pME6010 from tetracycline to kanamycin.
Methods:
First, we designed forward and reverse primers for KanR (with native RSB and promoter) which we isolated from plasmid pCM66. The 5’ ends were complementary to the insert region of pME6010 (reaction 1).
We then designed forward and reverse primers to amplify the pME6010 backbone (reaction 2).
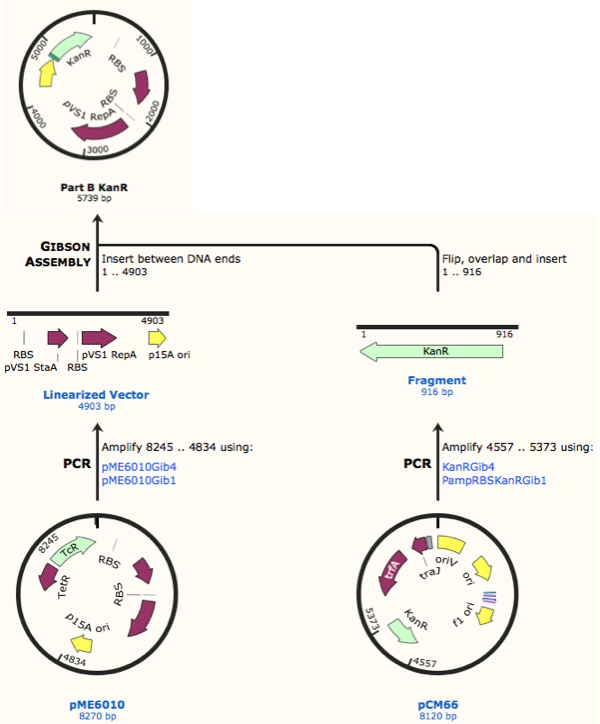
Further to this we redesigned each set of primers to incorporate an ampicillin promoter and optimised RBS (https://salis.psu.edu/software/) (B reactions) instead of the native promoter region (A reactions). This process is described in the following map:
The designed plasmids were as follows:
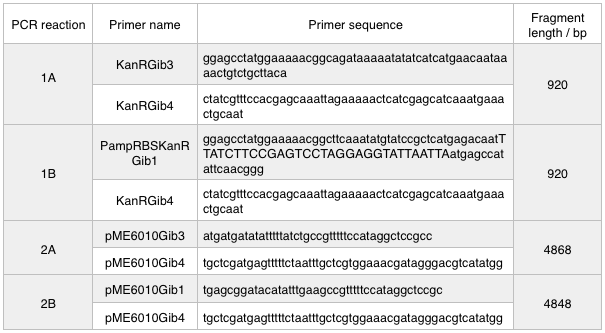
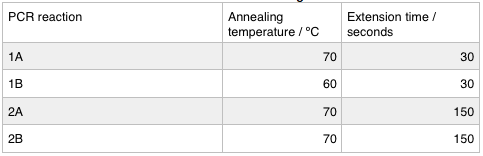
We ran these 4 PCR reactions as follows using the NEB Q5 PCR protocol:
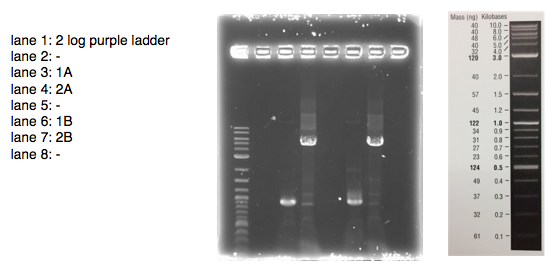
A 0.8% agarose gel was used for the extraction as this offered good separation around 1kb and 5kb. This gel was run and the bands extracted according to our QIAquick Gel Extraction Protocol using NEB purple loading dye and 2-log purple ladder and QIAquick extraction kit. Gel obtained:
As we later found out these NanoDrop readings are false since the QG buffer in the QIAGEN gel extraction kit interferes with the UV/Vis readings. Following extraction of our PCR products from the gel we used the NEB Gibson Assembly protocol to run an 8hr reaction over night. We ran an ‘A’ reaction which will insert the KanR gene into the pME6010 plasmid with the native promoter and a ‘B’ reaction that will insert the KanR gene with a pamp promoter and optimised RBS.
The volumes were chosen to satisfy 100ng vector with a 3-fold increase in the amount of insert. The insert amount must lie between 0.02 and 0.5 pmol and the total volume of total fragments cannot exceed 10µl. Following an overnight 8hr Gibson Assembly the reaction volumes were treated with Dpn1 restriction enzyme that cuts bacterial (methylated) DNA. We transformed the Gibson products into chemically competent DH5-alpha cells as well as into NEB alpha-5 cells. Unfortunately no colonies grew on a KanR plate! We know that the PCR products are correct so we think an issue may have arisen during the Gibson Assembly stage so we will re-do this part next week.
Week 2
Tasks completed:
Notebook:
To run equimolar amounts of insert to vector the following table was calculated:
These gibson assemblies were then transformed into E.coli cells using electroporation with the voltage at 1.8kV and the exposure time ranging from 4.8ms to 5.2ms. The cells were then recovered at 37ºC for 1.5 hrs before 100 µl was plated onto KanR plates. These were left overnight at 37ºC.
Unfortunately, there were no colonies grown and as we have no product from our gibson assemblies remaining we will have to repeat the process from Week 1 in order to trouble-shoot the method and get the desired plasmid. It is possible that we accidentally swapped the primers or PCR products around at some point and thus we had 1A with 2B and 1B with 2A in the Gibson assembly. These combinations are not complementary and thus the Gibson would fail meaning that none of our bacteria received intact Kanamycin resistance in the transformation. However there are many other reasons why the process may have failed therefore we went through the entire method trying to cover all instances where we may have made an error.
First we confirmed that the primer sequences on the delivery tubes matched what we had ordered. We ran the PCR as before; we re-calculated the annealing temperatures and extension times with the same result as last week.
In week one, we did the DpnI digest after the Gibson assembly and not directly after the PCR. This time we did the digest immediately after the PCR to ensure that all template DNA was destroyed before running on the gel, ensuring cleaner bands on our gel and less contamination from non-PCR products.
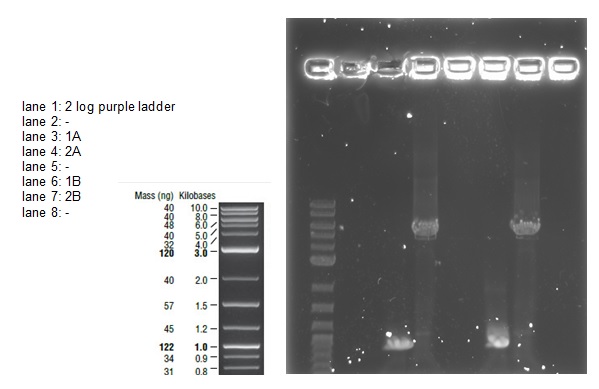
We loaded 20 μl of each PCR product onto a 0.8% agarose gel as before. The resulting bands were visualized after staining as shown below:
The bands are not as bright as in the first attempt probably due to less time spent in the ethidium bromide staining tank. The bands correspond with the correct sized fragments as before. This suggests there was no error in the PCR reaction with the exception of again mixing up the primers (the PCR would be successful with any combination of the primers for a 1 and 2 reaction). The bands are cleaner allowing us to try an alternative to the gel extraction in case that caused our assembly and transformation to fail.
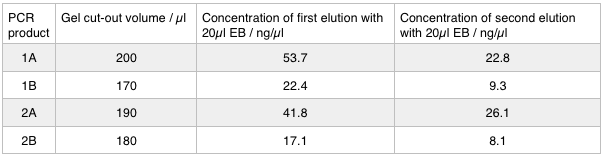
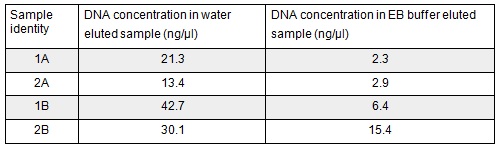
Next we excised and extracted our bands from the gel with the same QIAquick Gel Extraction Protocol protocol as before but with two changes. First we allowed additional drying time after the step involving ethanol supplemented buffer in case ethanol contamination inhibited the enzymes used in the Gibson assembly later in the process. We also replaced the 20 μl EB buffer with 20μl of MilliQ water. We did our second elution with the kit EB buffer. We then measured the concentration of DNA in each eluted sample using the nanodrop. This still gives us a reasonable idea (despite the interference from the remaining QG buffer residue) as to the amount of DNA in each sample from which to calculate our dilutions for the Gibson assembly reaction.
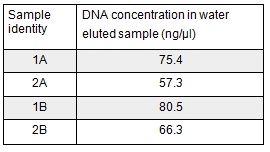
We used the remaining 10μl of our PCR products to do a Promega PCR clean-up and eluted using water. We gathered the following Nanodrop data:
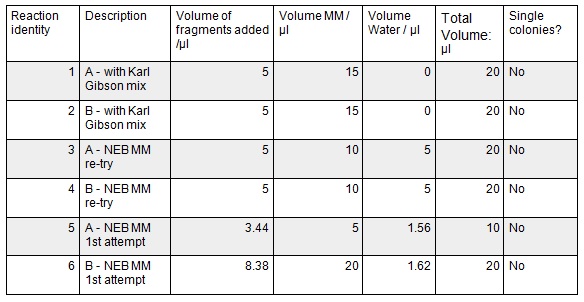
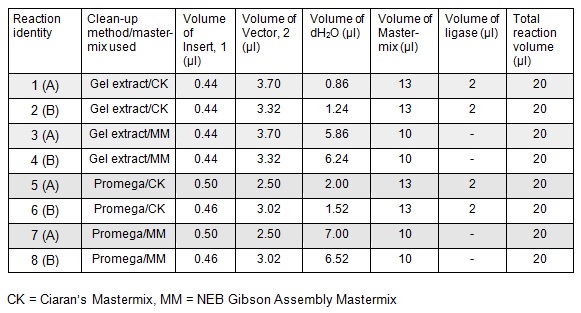
Next we ran the following Gibson assembly reactions:
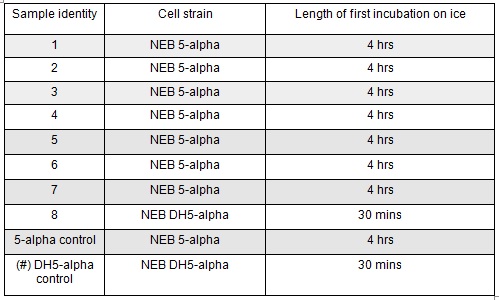
Next we transformed the Gibson assembly products into chemically competent E.coli cells. We decided to run a positive control for each kind of cell used using the pUC19 plasmid provided with the cells. Unfortunately we were limited with the amount of cells available and had to use two different cell strains; this resulted in some variation in the length of the first incubation with the DNA on ice as we began the transformation in the morning but then decided to pause the process so that the cells would be plated out later in the day in order to not have too much growth overnight. The NEB 5-alpha cells were also on ice slightly longer before the DNA was added.
The cells were plated out after 1.5 hour incubation. It was noted that one of the sample appeared to have leaked from the Eppendorf into the petri dish containing all the samples in the incubator. We think it was the 5-alpha control as this was the only sample that appeared to have lost volume. Samples 1-8 were plated on Kanamycin plates (i) that we made up during the final incubation period. The remaining sample was re-suspended in ~100μl of SOC to create a concentrated solution of cells that were also plated out on kanamycin plates (ii). The two controls were spread on Ampicillin plates made up by Nick. All plates were placed in the 37⁰c incubator overnight. The results in the morning were as follows:
Apart from the concentrated sample 1ii, none of the samples prepared using the gel extraction protocol grew. Sample 1ii also had limited growth compared to the other plates with colonies:
Samples 1i and 5i differed only in their clean-up method; 5i has lots of growth and 1i has none. This comparison holds true for Samples 2+6, 3+7 and 4+8 with the same result:
This suggests that we should use the Promega PCR clean-up kit and not the QIAquick Gel extraction. We will still run our samples on a gel to check the quality of our PCR reactions. We may investigate the Promega Gel extraction kit as this will enable us to pick our desired band out of a PCR that has also generated non-specific fragments which we cannot do with the Promega PCR clean-up protocol.
The samples that were treated with the master-mix and ligase supplied by Ciaran had more colonies than those that were treated with the NEB supplied master-mix as shown by the comparison between samples 5i and 7i:
This suggests that in future we should use the master-mix supplied by Ciaran.
All the controls grew showing that the lack of growth of samples 1- 4 was not due to inefficient transformation:
There was no observable difference between sample 8 and sample 7 which were treated the same but which used different chemically competent cells. Thus we can continue to use wither the NEB 5-alpha or DH5-alpha chemically competent cells.
There was no observable difference between the samples that correspond to A Gibson assembly reactions (template for PCR primers and therefore fragment = pCM66) and those that correspond to B Gibson assembly reactions (PCR primers and therefore fragment include Pamp).
We will confirm that the plasmid in each plate of transformed cells is the desired product by growing up the cultures overnight and extracting the plasmid for sequencing. We are fairly confident that it is the correct product as the fragment we were inserting was the kanamycin resistance itself which is necessary for growth on the kanamycin plates.
Part C
 "
"