Team:Oxford/biosensor characterisation
From 2014.igem.org
(Difference between revisions)
| Line 215: | Line 215: | ||
<br><br><br><br><br><br> | <br><br><br><br><br><br> | ||
| - | We modelled the probability of a molecule of GFP being created using the Michaelis-Menten model (α_1), incorporating a basal transcription rate (beta1). For the degradation, we assumed a simple proportional relationship | + | We modelled the probability of a molecule of GFP being created using the Michaelis-Menten model (α_1), incorporating a basal transcription rate (beta1). For the degradation, we assumed a simple proportional relationship: the more GFP you have, the more likely it is that a molecule degrades (δ_1). The constant of proportionality will be a function of the intrinsic life time of the protein in the cell. We considered there to be no DCM originally, then a large step in DCM at time=0. This is similar to placing the detector in a DCM polluted source, to make the model more realistic the level of DCM would go down as it is degraded but we had no time to obtain data for this rate. |
<br><br> | <br><br> | ||
| - | To decide if GFP was produced, we looked at the percentage of “reactions” which were | + | To decide if GFP was produced, we looked at the percentage of “reactions” which were productive, and then we compared this to a second random number (again taken from a uniform distribution from 0 to 1). If the random number was lower, then a GFP was created. If it was higher, then a GFP was degraded. In this way we make a weighted random choice about whether GFP was created or degraded. |
<br><br> | <br><br> | ||
| Line 226: | Line 226: | ||
<br><br> | <br><br> | ||
| - | Stochastic modelling is useful because it can show us the stochastic effects which are often | + | Stochastic modelling is useful because it can show us the stochastic effects which are often observed in individual bacteria. By calculating the variation of the mean of multiple GFP producing bacteria, we can also work out the standard deviation. Then, if we assume that the system varies with respect to the normal distribution, we can produce error bounds for the production of GFP, such that we can say that 90% of the time we can expect the production of GFP from a single bacterium to be within these two curves. This could be useful for seeing if results are unexpected, or, if there are multiple outliers, that our model is incorrect. If we average an increasing number bacteria, then the mean curve tends towards the deterministic response. This is to be expected, as we are now looking at the system as a whole and fluctuations in the production from individual bacteria are averaged out. In terms of their use, when looking at small amounts of bacterium the stochastic model would be better, because real random fluctuations can be seen. For larger bacterial populations, the deterministic response models the growth very well. The stochastic model can also model large groups but requires large number of realisations which causes simulations to take a lot longer to run. |
<br><br> | <br><br> | ||
When running the models, we picked arbitrary constants to view the general response. If we had more time we would have attempted to work out the basal rate, transcription rate and degradation rate of the GFP from DCM. | When running the models, we picked arbitrary constants to view the general response. If we had more time we would have attempted to work out the basal rate, transcription rate and degradation rate of the GFP from DCM. | ||
| Line 398: | Line 398: | ||
<h1>Wetlab data showing response in level of mCherry expressed with different concs of ATC</h1> | <h1>Wetlab data showing response in level of mCherry expressed with different concs of ATC</h1> | ||
| - | By making a translational fusion of mCherry at the | + | By making a translational fusion of mCherry at the C terminus of the dcmR gene under the tet promoter and tet operator system (see our <a href="https://2014.igem.org/Team:Oxford/biosensor_construction">Construction page</a> for details) we could measure mCherry fluorescence to gain information about dcmR induction by ATC. Expression was induced with various amounts of ATC and the following fluorescence data acquired. Exposure time was 0.2 seconds. As no calibration data was obtained using purified mCherry, the results have been left in fluorescence arbitrary units. Images were analysed using imageJ software.<br> |
mCherry fluorescence increases with amount of ATC used confirming that the dcmR gene was expressed under the control of the tet promoter and operator system.<br><br> | mCherry fluorescence increases with amount of ATC used confirming that the dcmR gene was expressed under the control of the tet promoter and operator system.<br><br> | ||
<img src="https://static.igem.org/mediawiki/parts/5/51/Oxford_DcmR-mCherry_expression_induced_by_0ng_ATC.png" | <img src="https://static.igem.org/mediawiki/parts/5/51/Oxford_DcmR-mCherry_expression_induced_by_0ng_ATC.png" | ||
| Line 447: | Line 447: | ||
<h1>Introduction</h1> | <h1>Introduction</h1> | ||
| - | As you can see from the biochemistry bubble above, our team was only able to obtain fluorescence data for the first half of the genetic circuit (ATC induced mCherry response). On top of this, the wet lab team were unable to obtain data that measured how the fluorescence of a single culture changed with time, again because of time constraints. This is slightly limiting because it means that we don’t have any dynamic data for any part of our system, and therefore can’t test the modelling predictions of the speed of the biosensor’s response. | + | As you can see from the biochemistry bubble above, our team was only able to obtain fluorescence data for the first half of the genetic circuit (ATC-induced mCherry response). On top of this, the wet lab team were unable to obtain data that measured how the fluorescence of a single culture changed with time, again because of time constraints. This is slightly limiting because it means that we don’t have any dynamic data for any part of our system, and therefore can’t test the modelling predictions of the speed of the biosensor’s response. |
<br><br> | <br><br> | ||
The original data is shown on the right with error bars showing the standard error of the measurements. | The original data is shown on the right with error bars showing the standard error of the measurements. | ||
| Line 463: | Line 463: | ||
<img src="https://static.igem.org/mediawiki/2014/4/46/Oxford_data2.png" style="float:right;position:relative; width:80%;margin-left:10%;margin-right:10%;" /> | <img src="https://static.igem.org/mediawiki/2014/4/46/Oxford_data2.png" style="float:right;position:relative; width:80%;margin-left:10%;margin-right:10%;" /> | ||
<br><br> | <br><br> | ||
| - | We then ran the model for the correct amount of time (2 hours 20mins incubation with ATC) and ran it for lots of different concentrations of ATC over the range that the wet-lab team | + | We then ran the model for the correct amount of time (2 hours 20mins incubation with ATC) and ran it for lots of different concentrations of ATC over the range that the wet-lab team tested. The parameters are still arbitrary at this point (the same as above) and the results of the graphs are therefore arbitrary are as well, but the input values are now correct. The graphic below shows how we used the existing model to obtain the same graphs as the wet-lab team had obtained. |
<br><br> | <br><br> | ||
The numerical inputs that were used to model this data set were therefore: | The numerical inputs that were used to model this data set were therefore: | ||
Revision as of 18:56, 17 October 2014
 "
"








 Deterministic models are very powerful tools for synthetic biology. They describe the behaviour of the bacteria at the population level and use Ordinary Differential Equations (ODEs) to relate each activation and repression. By constructing a cascade of differential equations one can build a realistic model of the average behaviour of the system.
Deterministic models are very powerful tools for synthetic biology. They describe the behaviour of the bacteria at the population level and use Ordinary Differential Equations (ODEs) to relate each activation and repression. By constructing a cascade of differential equations one can build a realistic model of the average behaviour of the system.



 Oxford iGEM 2014
Oxford iGEM 2014














 This simplifies the equation to:
This simplifies the equation to:
 As we want our model to accurately predict the fluorescence, we will substitute the fluorescence value in place of the [DcmR] and rearrange:
As we want our model to accurately predict the fluorescence, we will substitute the fluorescence value in place of the [DcmR] and rearrange:
 Substituting in the value for δ1 that we found above and the basal steady state fluorescence level from the data (471 to 3 s.f.) gives the basal transcription rate as:
Substituting in the value for δ1 that we found above and the basal steady state fluorescence level from the data (471 to 3 s.f.) gives the basal transcription rate as:

 alongside the correct inputs:
alongside the correct inputs:
 The graph below shows the model's predictions plotted in the same figure as the data points that the wet-lab team obtained for the system:
The graph below shows the model's predictions plotted in the same figure as the data points that the wet-lab team obtained for the system:
 Plotting the model's output as a by interpolating between the calculated values makes the graph clearer:
Plotting the model's output as a by interpolating between the calculated values makes the graph clearer:


 The first system that we tested is shown here. This is the bottom level of our synthetic circuit. Testing just this plasmid allowed us to obtain some important information to allow us to characterise the genetic system.
The first system that we tested is shown here. This is the bottom level of our synthetic circuit. Testing just this plasmid allowed us to obtain some important information to allow us to characterise the genetic system.
 The second system that we tested was the whole synthetic system without any DCM added. This allowed us to analyse just the effect of DcmR on the PdcmA promoter when it was compared to the result from system 1.
The second system that we tested was the whole synthetic system without any DCM added. This allowed us to analyse just the effect of DcmR on the PdcmA promoter when it was compared to the result from system 1.
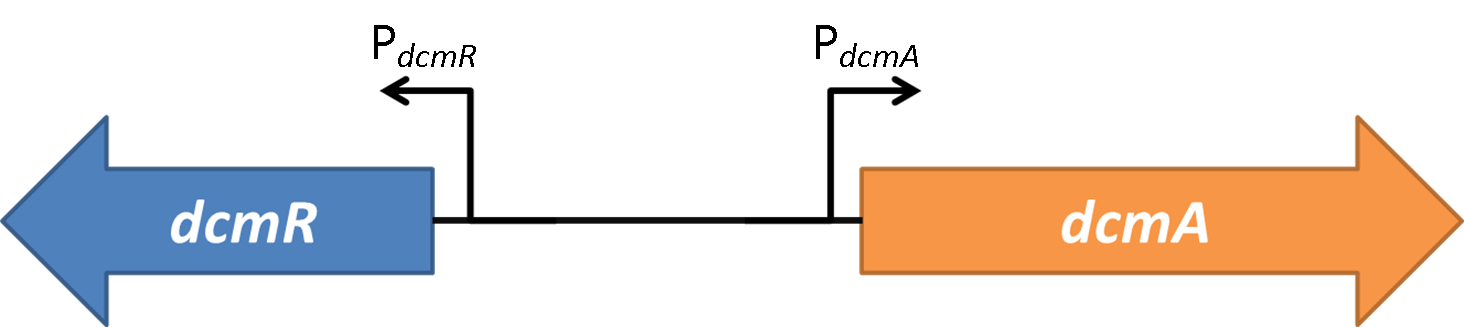
 Therefore we know that DcmR represses PdcmA with DCM repressing this repression. This means that the system is a double repressor!
Therefore we know that DcmR represses PdcmA with DCM repressing this repression. This means that the system is a double repressor!





























































