Subtilin
Subtilin is a small antimicrobial peptide (AMP) which belongs to the class of lanthionine-containing antibiotics (lantibiotics). These are heat-stable, ribosomally synthesized molecules with a molecular weight below 4 kDa. Their main characteristic is the high proportion of unusual amino acids, synthesized by post-translational side-chain modifications of precursor peptides. Most prominent examples are the polycyclic thioether amino acids, lanthionine and β–methyllanthionine and a number of dehydrated amino acids such as dehydroalanine (Dha) and dehydrobutyrine (Dhb).
Lantibiotics are encouraging candidates for future antimicrobials, as they inhibit the growth of many clinically relevant pathogens comprising even multidrug-resistant bacteria. Moreover, lantibiotics have only low tendency to generate resistance. Both features make them highly attractive for medical applications.

Fig. 1. Schematic representation of the structure of the lantibiotic subtilin. Dha represents a dehydroalanine residue, Dhb represents a dehydrobutirine residue, and Abu represents a dehydrobutirine residue that has formed a thioether β-methyl-lanthionine bridge with a cysteine residue. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Quorum+sensing+control+of+lantibiotic+production%3B+nisin+and+subtilin+autoregulate+their+own+biosynthesis]
The spaBTCSIFEGRK gene cluster

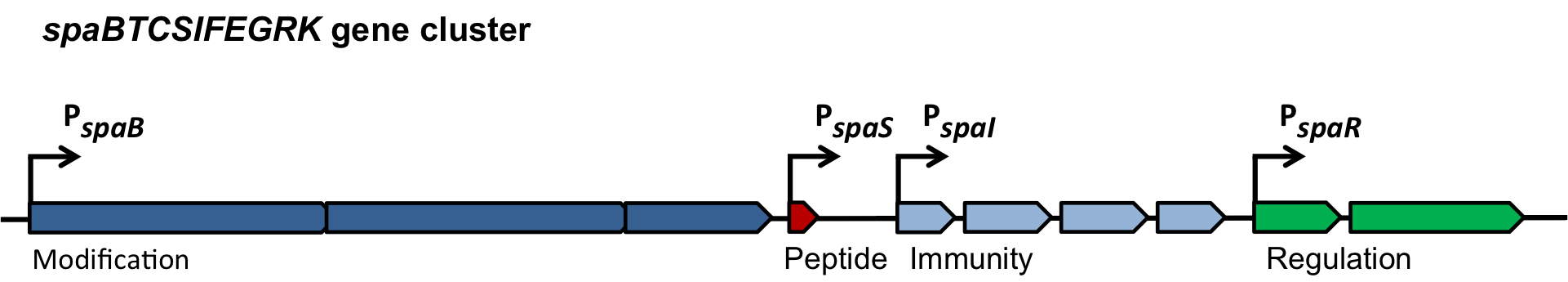
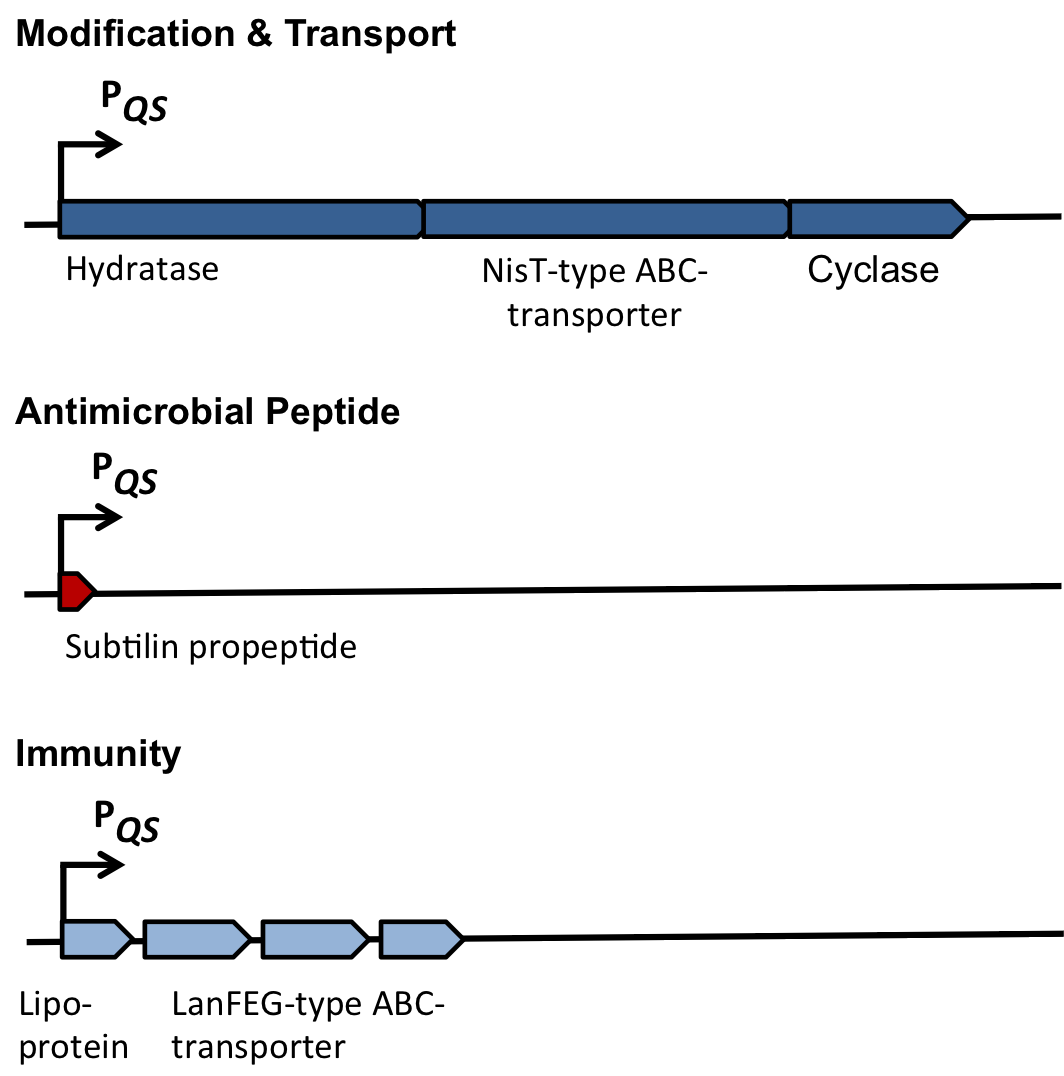
Fig. 2. Organisation of the subtilin biosynthetic gene cluster. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Quorum+sensing+control+of+lantibiotic+production%3B+nisin+and+subtilin+autoregulate+their+own+biosynthesis]
As true for many other lantibiotics, the genes for the subtilin biosynthesis are clustered. For subtilin, there is even a reasonable arrangement within the cluster. The genes spaBTC play a role for the posttranslational modifications and the transport of subtilin. They are regulated by their own promotor PspaB and polycistronically transcribed. spaS encodes the propeptide, which is derived from a short monocistronic mRNA. The next transcription entity, spaIFEG, is coding for the immunity, whereas spaRK encode an two-component-system that plays an important role for the regulation. What is missing in comparison to other lantibiotic gene clusters is a specific protease (encoded by lanP) that cleaves off the leader sequence. In Bacillus subtilis, this task is accomplished by unspecific extracellular proteases.
TABLE MISSING
Self-protection from subtilin
Of course, the producer needs to be immune against the antimicrobial agent it produces. B. subtilis ATCC6633 protects itself against subtilin by two mechanisms that both act independently and confer some level of resistance, however full resistance is only achieved when both mechanisms are active. SpaI is a membrane-anchored lipoprotein. It is suggested that it binds the antimicrobial peptide and by this keeps it away from the membrane. SpaFEG is a LanFEG-like ABC-transporter that works by exporting subtilin from the cytoplasmic membrane to the culture supernatant.
Dual regulation of subtilin biosynthesis and immunity
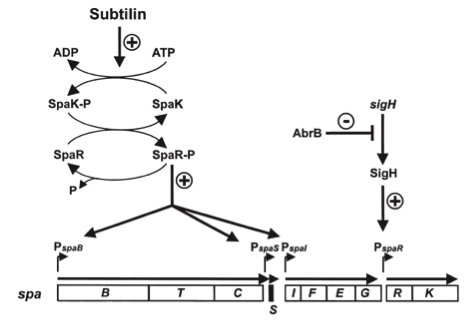
Fig. 3. Dual control of subtilin biosynthesis and immunity. [http://www.ncbi.nlm.nih.gov/pubmed/11972779]
Subtilin biosynthesis and immunity in B. subtilis ATCC6633 are subject to a dual control mechanism.
To begin with, subtilin production is positively regulated by sigma factor H (SigH). sigH itself is repressed by the transition state regulator AbrB during exponential growth phase. Thus, subtilin production is highest at the transition to stationary phase. Moreover, when a threshold level of subtilin concentration is reached in the extracellular space, the peptide acts as a pheromone. Subtilin activates the two-component system SpaRK, consisting of a histidine kinase and a response regulator, which in turn induces expression of spaBTC, spaS and spaIFEG. This dual control mechanism allows the coordination of subtilin biosynthesis with the physiological state of the cell.
Mode of action
Lantibiotics are active against a wide range of Gram-positive bacteria including multidrug-resistant pathogens. Some lantibiotics have a dual mode of action, subtilin is proposed to be among them. Subtilin forms a complex with the cell wall precursor lipid II, whereby the pyrophosphate moiety may play a key role for target recognition. Thereby, the cell wall biosynthesis is inhibited. Furthermore, the complexes aggregate and form a pore in the bacterial membrane by using lipid II as a docking molecule. This leads to a depolarisation of the membrane potential, the efflux of cytoplasma and in turn to rapid cell death. Just like nisin, which is highly similar, subtilin is still active in the nanomolar range and thus serves excellently as alternative killing strategy against multidrug-resistant pathogens.

Fig. 4. Proposed mechanism of pyrophosphate-mediated target engagement by the lanthionine antibiotic subtilin. Membrane breaches occur in a lipid II-mediated fashion and cell wall synthesis is impeded through engagement of pyrophosphate-containing intermediates into complexes. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Molecular+Mechanism+of+Target+parisot]
Lysostaphin
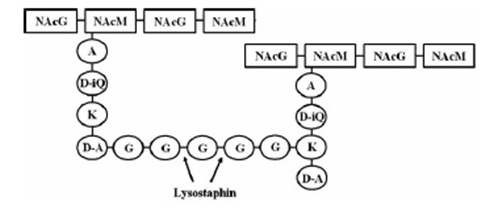
targeted pentaglycinbridge in the cellwall of some Staphylococcus species [1]
Lysostaphin is a murein-hydrolase, which is naturally produced by Staphylococcus simulans biovar staphylolyticus in order to defeat other, competing Staphylococcus species. It cleaves specifically the pentaglycinbridge between the tetrapeptides of the peptidoglycan in the cell wall. In addition to killing the cells, lysostaphin also dissolves the biofilms of lysostaphin-sensitive bacteria. [http://www.ncbi.nlm.nih.gov/pubmed/19149589 [A1]]
This targeted pentaglycinbridge is a feature of the cell wall of some Staphylococcus species, leading to the high specificity of lysostaphin against S. carnosus, S. epidermidis and Staphylococcus aureus, which is particularly sensitive towards lysostaphin. http://www.ncbi.nlm.nih.gov/pubmed/18607587 A2
The immunity of S. simulans is accomplished by substitution of the glycine-residues in the cell wall by serine which is mediated by the immunity factor Lif [http://www.ncbi.nlm.nih.gov/pubmed/1914958 [A1]] .

maturation of Lysostaphin [A1]
The preprolysostaphin consists of a signal peptide to mediate the export, 15 tandem repeats of 13 aminoacids length and two protein domains: the peptidase-domain and the C-terminal targeting domain. The signal peptide is cleaved during the export, the tandem repeats are cleaved by an additionally secreted cysteine protease. This repeats are not necessary for correct protein folding or maturation of the protein, they just inhibit protein activity. Once cleaved off, the maturated protein is 4.5 fold more active. [http://www.ncbi.nlm.nih.gov/pubmed/1914958 [A1]]
Because of its promising attributes, the usage of lysostaphin as coating for implants [http://www.ncbi.nlm.nih.gov/pubmed/21709102 [A3] ] and treatment of S. aureus in the nasal area via cream [http://www.ncbi.nlm.nih.gov/pubmed/12709327[A4]] is currently in the focus of medical research.
Dispersin
DispersinB (DspB) is a biofilm-dissolving enzyme, specifically a 1,6-beta-N-acetyl-glucosaminidase. It is produced by Aggregatibacter actinomycetemcomitans, a pathogen which causes aggressive forms of Parodontitis. [http://www.ncbi.nlm.nih.gov/pubmed/23103508 [A5]] In its natural habitat, A. actinomycetecombitans uses this enzyme to release single cells from the biofilm in order to colonize new habitats. [http://www.ncbi.nlm.nih.gov/pubmed/15878175[A6]] DspB is specific towards its substrate, PNAG, (PGA in E. coli), which is an important component of the extracellular matrix and which can make up to 90% of the dry weight of a biofilm. [http://www.ncbi.nlm.nih.gov/pubmed/15576769 [A7]] The process of dissolving biofilms is accomplished by cleaving monosaccharide-residues from the polysaccharide, beginning at the non-reducing end of the PNAG. [http://www.ncbi.nlm.nih.gov/pubmed/15878175[A6]] This glycosamin-hydrolase has a typical TIM-barrel-structure, consisting of 8α- and 8β-barrels, with a large cavity in the middle, which is considered to be the substrate-binding pocket. [http://www.ncbi.nlm.nih.gov/pubmed/23103508 [A5]]
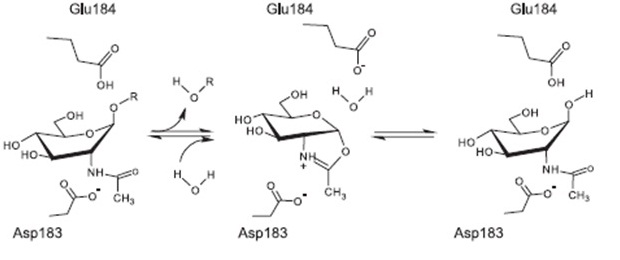
Proposed mechanism of reaction [http://www.ncbi.nlm.nih.gov/pubmed/17949435 [A8]
The currently proposed mechanism of reaction suggests a substrate assisted nucleophilic attack (see chart below) [http://www.ncbi.nlm.nih.gov/pubmed/17949435[A8]]
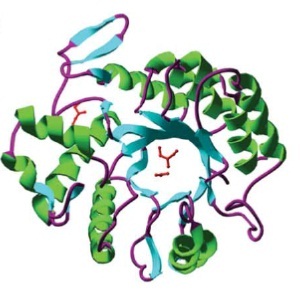
Barrel structure of DspB [http://www.ncbi.nlm.nih.gov/pubmed/15878175[A6]
DspB is active against many bacteria, both gram-positive and gram-negative, and shows some very promising features for its use in medical devices. It can prevent the growth of biofilms in coated catheters if used in combination with triclosan, and can be used to clear medical devices of biofilms. [http://www.ncbi.nlm.nih.gov/pubmed/19447791[A9]]
Sources:
[A1] M.C.F. Bastos, H. Ceotto1, M.L.V. Coelho and J.S. Nascimento: Staphylococcal Antimicrobial Peptides: Relevant Properties and Potential Biotechnological Applications in Current Pharmaceutical Biotechnology (2009) S. 38-61
[A2] Jaspal K. Kumar: Lysostaphin: an antistaphylococcal agent in Appl Microbiol Biotechnol (2008) S.555-561 )
[A3] Rohan Satishkumar, Sriram Sankar, Yuliya Yurko, Amy Lincourt, John Shipp, B. Todd Heniford and Alexey Vertegel: Evaluation of the Antimicrobial Activity of Lysostaphin-Coated Hernia Repair Meshes in Antimicrobial Agents and Chemotherapy (2011)
[A4] John F. Kokai-Kun, Scott M. Walsh, Tanya Chanturiya, and James J. Mond: Lysostaphin Cream Eradicates Staphylococcus aureus Nasal Colonization in a Cotton Rat Model in Antimicrobial Agents and Chemotherapy, 47 (2003)
[A5] Erika Fazekas, Lili Kandra, Gyongyi Gyemant: Model for b-1,6-N-acetylglucosamine oligomer hydrolysis catalysed by DispersinB, a biofilm degrading enzyme, in Carbohydrate Research 363 (2012), S. 7-13
[A6] N. RamasubbuL. M. Thomas, C. Ragunath and J. B. Kaplan: Structural Analysis of Dispersin B, a Biofilm-releasing Glycoside Hydrolase from the Periodontopathogen Actinobacillus actinomycetemcomitans, in Journal of Molecular Biology 340 (2005) S. 475-486
[7] Jeffrey B. Kaplan, Kabilan Velliyagounder, Chandran Ragunath, Holger Rohde,
Dietrich Mack, Johannes K.-M. Knobloch, and Narayanan Ramasubbu: Genes Involved in the Synthesis and Degradation of Matrix Polysaccharide in Actinobacillus actinomycetemcomitans and Actinobacillus pleuropneumoniae Biofilms, in JOURNAL OF BACTERIOLOGY, 186 (2004) S.8213-8220
[A8] Suba G. A. Manuel, Chandran Ragunath, Hameetha B. R. Sait, Era A. Izano, Jeffrey B. Kaplan and Narayanan Ramasubbu: Role of active-site residues of dispersin B, a biofilm-releasing b-hexosaminidase from a periodontal pathogen, in substrate hydrolysis, in The FEBS journal (2007)
[A9] Darouiche RO, Mansouri MD, Gawande PV, et al. Antimicrobial and antibiofilm efficacy of triclosan and DispersinB combination. J Antimicrob Chemother 2009; 64: 88-93
 "
"