Team:Oxford/biosensor characterisation
From 2014.igem.org
(Difference between revisions)
Olivervince (Talk | contribs) |
|||
| Line 56: | Line 56: | ||
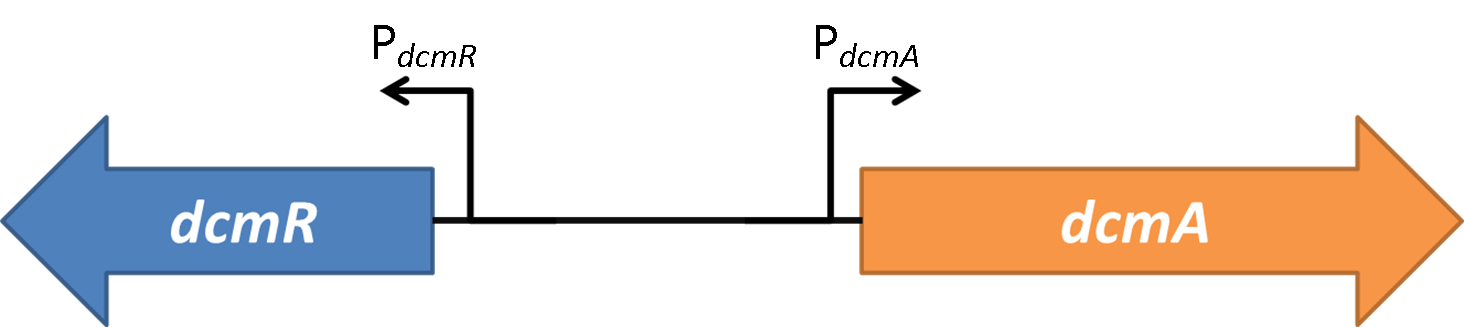
| - | In order to design and create a stable and sensitive system that responds to DCM we first need to characterise the regulatory nature of DcmR. Characterisation of this regulatory network has never been done before although it has been suggested to be a repressor [1]; we will be the first to fully characterise the mode of action of <font style="font-style: italic;">dcmR</font>. To do this we suppose the following hypotheses for DCM activating the transcription of <font style="font-style: italic;">dcmR</font>: either double repression or double activation. In other words, either DcmR represses <font style="font-style: italic;">dcmA</font> expression and DcmR is in | + | In order to design and create a stable and sensitive system that responds to DCM we first need to characterise the regulatory nature of DcmR. Characterisation of this regulatory network has never been done before although it has been suggested to be a repressor [1]; we will be the first to fully characterise the mode of action of <font style="font-style: italic;">dcmR</font>. To do this we suppose the following hypotheses for DCM activating the transcription of <font style="font-style: italic;">dcmR</font>: either double repression or double activation. In other words, either DcmR represses <font style="font-style: italic;">dcmA</font> expression and DcmR is in negatively modulated by the presence of DCM; or expression of <font style="font-style: italic;">dcmA</font> requires DcmR as an activator, with DcmR in turn only activated in the presence of DCM.<br><br> |
</div> | </div> | ||
| Line 88: | Line 88: | ||
<h1>DcmR and regulation of <font style="font-style: italic;">dcmA</font> expression</h1> | <h1>DcmR and regulation of <font style="font-style: italic;">dcmA</font> expression</h1> | ||
| - | Mutants with <font style="font-style: italic;">dcmA</font> and the intergenic region but without complete <font style="font-style: italic;">dcmR</font> express | + | Mutants with <font style="font-style: italic;">dcmA</font> and the intergenic region but without complete <font style="font-style: italic;">dcmR</font> express <font style="font-style: italic;">dcmA</font> constitutively. Re-integration of <font style="font-style: italic;">dcmR</font> restores regulation of <font style="font-style: italic;">dcmA</font> expression at the transcriptional level [1]. In addition, it has been shown that the region including <font style="font-style: italic;">dcmR</font>, the intergenic region and <font style="font-style: italic;">dcmA</font> is sufficient to confer a DCM dependent response in genetically engineered Methylobacterium extorquens DM4 [2]. <br><br> |
<h1>DcmR and DNA-binding</h1> | <h1>DcmR and DNA-binding</h1> | ||
| Line 101: | Line 101: | ||
<br><br> | <br><br> | ||
<h1>Characterising the DcmR - DCM - P_dcmA interaction</h1> | <h1>Characterising the DcmR - DCM - P_dcmA interaction</h1> | ||
| - | To find out whether the gene <font style="font-style: italic;">dcmR</font> acts as a repressor or an activator on the promoter of the <font style="font-style: italic;">dcmA</font> gene, we attempted to build the genetic circuit shown above on the right. Having <font style="font-style: italic;">dcmR</font> under inducible TetR expression should allow us to have very good control of the amount of DcmR present. | + | To find out whether the gene <font style="font-style: italic;">dcmR</font> acts as a repressor or an activator on the promoter of the <font style="font-style: italic;">dcmA</font> gene, we attempted to build the genetic circuit shown above on the right. Having <font style="font-style: italic;">dcmR</font> under inducible TetR expression should allow us to have very good control of the amount of DcmR present. Additionally a translational fusion with DcmR and a mCherry fluorescence tag will act as another confirmation to the amount of DcmR present. |
<br><br> | <br><br> | ||
We then extensively modelled the circuit to discover how the response of the system would differ if it was either of the two circuit systems. Click the modelling bubbles (pink) to find out exactly how we achieved this. | We then extensively modelled the circuit to discover how the response of the system would differ if it was either of the two circuit systems. Click the modelling bubbles (pink) to find out exactly how we achieved this. | ||
| Line 136: | Line 136: | ||
<h1>Predicting the mCherry fluorescence</h1> | <h1>Predicting the mCherry fluorescence</h1> | ||
| - | We simplified the first double repression by modelling it as an activation of <font style="font-style: italic;">dcmR</font> by ATC, albeit parameterised by different constants. This assumption is justified by the fact that we are able to precisely control the addition of ATC and | + | We simplified the first double repression by modelling it as an activation of <font style="font-style: italic;">dcmR</font> by ATC, albeit parameterised by different constants. This assumption is justified by the fact that we are able to precisely control the addition of ATC and measure the fluorescence of the mCherry. |
<br> | <br> | ||
<br> | <br> | ||
| Line 165: | Line 165: | ||
<br> | <br> | ||
| - | The differential equation that | + | The differential equation that describes this first step of the system is: |
| Line 179: | Line 179: | ||
<br><br><br><br><br><br> | <br><br><br><br><br><br> | ||
| - | Solving this ODE in Matlab (with zero basal transcription rate) predicts the following the response of the system: | + | Solving this ODE in Matlab (with a zero basal transcription rate) predicts the following the response of the system: |
<br><br> | <br><br> | ||
This model works assuming that sufficient TetR is always present. | This model works assuming that sufficient TetR is always present. | ||
Revision as of 12:14, 17 October 2014
 "
"








 Deterministic models are very powerful tools for synthetic biology. They describe the behaviour of the bacteria at the population level and use Ordinary Differential Equations (ODEs) to relate each activation and repression. By constructing a cascade of differential equations one can build a realistic model of the average behaviour of the system.
Deterministic models are very powerful tools for synthetic biology. They describe the behaviour of the bacteria at the population level and use Ordinary Differential Equations (ODEs) to relate each activation and repression. By constructing a cascade of differential equations one can build a realistic model of the average behaviour of the system.



 Oxford iGEM 2014
Oxford iGEM 2014











 This simplifies the equation to:
This simplifies the equation to:
 As we want our model to accurately predict the fluorescence, we will substitute the fluorescence value in place of the [DcmR] and rearrange:
As we want our model to accurately predict the fluorescence, we will substitute the fluorescence value in place of the [DcmR] and rearrange:
 Substituting in the value for δ1 that we found above and the basal steady state fluorescence level from the data (471 to 3 s.f.) gives the basal transcription rate as:
Substituting in the value for δ1 that we found above and the basal steady state fluorescence level from the data (471 to 3 s.f.) gives the basal transcription rate as:

 alongside the correct inputs:
alongside the correct inputs:
 The graph below shows the model's predictions plotted in the same figure as the data points that the wet-lab team obtained for the system:
The graph below shows the model's predictions plotted in the same figure as the data points that the wet-lab team obtained for the system:
 Plotting the model's output as a by interpolating between the calculated values makes the graph clearer:
Plotting the model's output as a by interpolating between the calculated values makes the graph clearer:


 The first system that we tested is shown here. This is the bottom level of our synthetic circuit. Testing just this plasmid allowed us to obtain some important information to allow us to characterise the genetic system.
The first system that we tested is shown here. This is the bottom level of our synthetic circuit. Testing just this plasmid allowed us to obtain some important information to allow us to characterise the genetic system.
 The second system that we tested was the whole synthetic system without any DCM added. This allowed us to analyse just the effect of DcmR on the PdcmA promoter when it was compared to the result from system 1.
The second system that we tested was the whole synthetic system without any DCM added. This allowed us to analyse just the effect of DcmR on the PdcmA promoter when it was compared to the result from system 1.
 Therefore we know that DcmR represses PdcmA with DCM repressing this repression. This means that the system is a double repressor!
Therefore we know that DcmR represses PdcmA with DCM repressing this repression. This means that the system is a double repressor!





























































