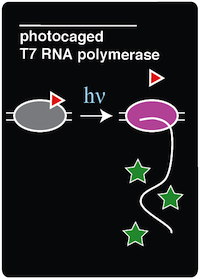
Team:Austin Texas/photocage
From 2014.igem.org
Nathanshin (Talk | contribs) |
Nathanshin (Talk | contribs) |
||
| Line 104: | Line 104: | ||
<h1>Experimental Methods</h1> | <h1>Experimental Methods</h1> | ||
| - | In order to create an in vivo light-activated GFP expression system, two plasmids were constructed and transformed into amberless E.coli cells. The first plasmid contains the tRNA and synthetase pair, which is necessary to incorporate ONBY into the amber stop codon of the T7 RNAP. The second plasmid contains the coding sequence for T7 RNAP, which has a mutation on the Tyrosine 639 residue, and a GFP coding sequence bound to an upstream T7 promoter. Once these components were assembled by Gibson Assembly, the two plasmids were transformed via electroporation into aliquots of Amberless E.coli. | + | In order to create an in vivo light-activated GFP expression system, two plasmids were constructed and transformed into amberless E.coli cells. The first plasmid contains the tRNA and synthetase pair, which is necessary to incorporate ONBY into the amber stop codon of the T7 RNAP. The second plasmid contains the coding sequence for T7 RNAP, which has a mutation on the Tyrosine 639 residue, and a GFP coding sequence bound to an upstream T7 promoter. The T7 RNAP coding sequence was taken from the BioBrick part BBa_K145001 and mutated to construct our desired plasmid. Once these components were assembled by Gibson Assembly, the two plasmids were transformed via electroporation into aliquots of Amberless E.coli. |
For this experiment, there were also other necessary control strains to test alongside the experimental strain. These controls included a T7-GFP construct with no amber stop codon in the O-helix (to serve as a positive control for expression with T7 polymerase), sfGFP amberless E.coli (to observe the expression of GFP by native polymerase), amberless E.coli (to serve as a cell background control), and LB supplemented with ncAA (to serve as a media background control). | For this experiment, there were also other necessary control strains to test alongside the experimental strain. These controls included a T7-GFP construct with no amber stop codon in the O-helix (to serve as a positive control for expression with T7 polymerase), sfGFP amberless E.coli (to observe the expression of GFP by native polymerase), amberless E.coli (to serve as a cell background control), and LB supplemented with ncAA (to serve as a media background control). | ||
Revision as of 03:41, 17 October 2014
| |||||||||||||||||||||||||||||
 "
"