Team:Austin Texas/photocage
From 2014.igem.org
| Line 86: | Line 86: | ||
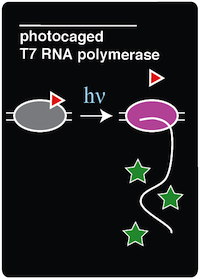
We recreated a light-activatable T7 RNA polymerase (RNAP) for the spatio-temporal control of protein expression. The light-activatable T7 RNAP was created by mutating a tyrosine codon at position 639 of a domain crucial for the polymerization of RNA during transcription. Y639 was mutated to an amber codon, allowing us to incorporate a ncAA at this position. We used ortho-nitrobenzyl tyrosine (ONBY), which is a "photocaged" ncAA ('''Figure 1'''). Thus, if our synthetase/tRNA pair works, position 639 should contain ONBY in place of tyrosine. This work is essentially a recapitulation of earlier work done by [Chou et al. 2010]. | We recreated a light-activatable T7 RNA polymerase (RNAP) for the spatio-temporal control of protein expression. The light-activatable T7 RNAP was created by mutating a tyrosine codon at position 639 of a domain crucial for the polymerization of RNA during transcription. Y639 was mutated to an amber codon, allowing us to incorporate a ncAA at this position. We used ortho-nitrobenzyl tyrosine (ONBY), which is a "photocaged" ncAA ('''Figure 1'''). Thus, if our synthetase/tRNA pair works, position 639 should contain ONBY in place of tyrosine. This work is essentially a recapitulation of earlier work done by [Chou et al. 2010]. | ||
| + | |||
When the photocaged ONBY is incorporated at position 639, RNAP activity is inhibited. The polymerase can become activated only upon de-caging of the ONB group from the ONBY. This is accomplished by exposing the cells to 365 nm light. When exposed to 365 nm light, the ONB group is released, resulting in a normal tyrosine amino acid at position 639. T7 RNAP was selected because of its orthogonal nature, which allows us to selectively induce the expression of specific genes that are preceded by the T7 RNAP promoter. Because T7 promoters are not natively found in ''E. coli'', a gene downstream of a T7 promoter may be exclusively expressed through the introduction of 365 nm light. | When the photocaged ONBY is incorporated at position 639, RNAP activity is inhibited. The polymerase can become activated only upon de-caging of the ONB group from the ONBY. This is accomplished by exposing the cells to 365 nm light. When exposed to 365 nm light, the ONB group is released, resulting in a normal tyrosine amino acid at position 639. T7 RNAP was selected because of its orthogonal nature, which allows us to selectively induce the expression of specific genes that are preceded by the T7 RNAP promoter. Because T7 promoters are not natively found in ''E. coli'', a gene downstream of a T7 promoter may be exclusively expressed through the introduction of 365 nm light. | ||
| Line 94: | Line 95: | ||
[[Image:Uncaging_of_ONBY.jpg | 300px|left|thumb| '''Figure 2.''' The caged T7 RNAP is decaged via exposure to 365 nm light. Figure reproduced from '''Chou et al. 2010'''.]] | [[Image:Uncaging_of_ONBY.jpg | 300px|left|thumb| '''Figure 2.''' The caged T7 RNAP is decaged via exposure to 365 nm light. Figure reproduced from '''Chou et al. 2010'''.]] | ||
| + | |||
Incorporation of ONBY at position 639 of T7 RNAP halts activity because of the bulky nature of ONBY (Chou et al. 2010). This ONB side group effectively renders T7 RNAP inactive. However, the bulky ONB group is able to be removed through irradiation with 365 nm light. The wavelength of light used to "decage" the amino acid proved to be another advantage of this system because 365 nm light is not toxic to the cell (Chou et al 2010). Once the ONB group is removed, a normal tyrosine residue is left in its place, restoring T7 RNA polymerase activity (''Figure 2'''). | Incorporation of ONBY at position 639 of T7 RNAP halts activity because of the bulky nature of ONBY (Chou et al. 2010). This ONB side group effectively renders T7 RNAP inactive. However, the bulky ONB group is able to be removed through irradiation with 365 nm light. The wavelength of light used to "decage" the amino acid proved to be another advantage of this system because 365 nm light is not toxic to the cell (Chou et al 2010). Once the ONB group is removed, a normal tyrosine residue is left in its place, restoring T7 RNA polymerase activity (''Figure 2'''). | ||
| + | |||
In order to incorporate the ncAA into amberless E.coli (which is described '''[here]'''), a ''Methanocaldoccus jannaschii'' tyrosyl-tRNA synthetase/tRNA pair was previously mutated to selectively charge and incorporate ONBY. Six residues (Tyr 32, Leu 65, Phe 108, Gln 109, Asp 158, and Leu 162) on the original synthetase were randomized and the library was selected for its ability to charge ONBY while discriminating against other canonical amino acids. The resulting mutant ONBY synthetase contained five mutations (Deiters et al. 2006). The following are the residues that were mutated on the synthetase: Tyr32→Gly32, Leu65→Gly65, Phe108→Glu108, Asp158→Ser158, and Leu162→Glu162. The Asp158→Ser158 and Tyr32→Gly32 mutations are believed to result in the loss of hydrogen bonds with the natural substrate, which would disfavor binding to tyrosine. Additionally, the Tyr 32→Gly 32 and Leu 65→Gly 65 mutations are believed to increase the size of the substrate-binding pocket to accommodate for the size of the bulky o-nitrobenzyl group (Deiters et al. 2006). | In order to incorporate the ncAA into amberless E.coli (which is described '''[here]'''), a ''Methanocaldoccus jannaschii'' tyrosyl-tRNA synthetase/tRNA pair was previously mutated to selectively charge and incorporate ONBY. Six residues (Tyr 32, Leu 65, Phe 108, Gln 109, Asp 158, and Leu 162) on the original synthetase were randomized and the library was selected for its ability to charge ONBY while discriminating against other canonical amino acids. The resulting mutant ONBY synthetase contained five mutations (Deiters et al. 2006). The following are the residues that were mutated on the synthetase: Tyr32→Gly32, Leu65→Gly65, Phe108→Glu108, Asp158→Ser158, and Leu162→Glu162. The Asp158→Ser158 and Tyr32→Gly32 mutations are believed to result in the loss of hydrogen bonds with the natural substrate, which would disfavor binding to tyrosine. Additionally, the Tyr 32→Gly 32 and Leu 65→Gly 65 mutations are believed to increase the size of the substrate-binding pocket to accommodate for the size of the bulky o-nitrobenzyl group (Deiters et al. 2006). | ||
| Line 102: | Line 105: | ||
In order to create an in vivo light-activated GFP expression system, two plasmids were constructed and transformed into amberless E.coli cells. The first plasmid contains the tRNA and synthetase pair, which is necessary to incorporate ONBY into the amber stop codon of the T7 RNAP. The second plasmid contains the coding sequence for T7 RNAP, which has a mutation on the Tyrosine 639 residue, and a GFP coding sequence bound to an upstream T7 promoter. Once these components were assembled by Gibson Assembly, the two plasmids were transformed via electroporation into aliquots of Amberless E.coli. | In order to create an in vivo light-activated GFP expression system, two plasmids were constructed and transformed into amberless E.coli cells. The first plasmid contains the tRNA and synthetase pair, which is necessary to incorporate ONBY into the amber stop codon of the T7 RNAP. The second plasmid contains the coding sequence for T7 RNAP, which has a mutation on the Tyrosine 639 residue, and a GFP coding sequence bound to an upstream T7 promoter. Once these components were assembled by Gibson Assembly, the two plasmids were transformed via electroporation into aliquots of Amberless E.coli. | ||
| + | |||
After transformation, the amberless E. coli cells containing both the synthetase/tRNA pair plasmid and the amber T7 RNAP/GFP plasmid were assayed for their ability to express GFP upon irradiation with 365 nm light in different intervals of time. The cells were irradiated for 0 minutes, 1 minute, 5 minutes, 15 minutes, and 30 minutes in order to observe the temporal control of GFP expression. | After transformation, the amberless E. coli cells containing both the synthetase/tRNA pair plasmid and the amber T7 RNAP/GFP plasmid were assayed for their ability to express GFP upon irradiation with 365 nm light in different intervals of time. The cells were irradiated for 0 minutes, 1 minute, 5 minutes, 15 minutes, and 30 minutes in order to observe the temporal control of GFP expression. | ||
| + | |||
For this experiment, there were also other necessary control strains to test alongside the experimental strain. These controls included a T7-GFP construct with no amber stop codon in the O-helix (to serve as a positive control for expression with T7 polymerase), sfGFP amberless E.coli (to observe the expression of GFP by native polymerase), amberless E.coli (to serve as a cell background control), and LB supplemented with ncAA (to serve as a media background control). | For this experiment, there were also other necessary control strains to test alongside the experimental strain. These controls included a T7-GFP construct with no amber stop codon in the O-helix (to serve as a positive control for expression with T7 polymerase), sfGFP amberless E.coli (to observe the expression of GFP by native polymerase), amberless E.coli (to serve as a cell background control), and LB supplemented with ncAA (to serve as a media background control). | ||
| + | |||
After irradiation with light, these cells were allowed to grow overnight before taking fluorescent measurements. This additional growth was necessary to allow the "decaged" T7 RNAP to polymerize mRNA transcripts of the GFP coding sequence. | After irradiation with light, these cells were allowed to grow overnight before taking fluorescent measurements. This additional growth was necessary to allow the "decaged" T7 RNAP to polymerize mRNA transcripts of the GFP coding sequence. | ||
| Line 115: | Line 121: | ||
To demonstrate the functionality of the light-activatable T7 RNAP, a GFP reporter was placed under the control of a T7 reporter. Cells with this plasmid were grown in culture with ONBY for 4-6 hours. These cells were then exposed to varying amounts of 365 nm light, ranging from 0 to 30 minutes. During this exposure, the ONB functional group at position ONBY639 should be released, resulting in de-caging of Y639. | To demonstrate the functionality of the light-activatable T7 RNAP, a GFP reporter was placed under the control of a T7 reporter. Cells with this plasmid were grown in culture with ONBY for 4-6 hours. These cells were then exposed to varying amounts of 365 nm light, ranging from 0 to 30 minutes. During this exposure, the ONB functional group at position ONBY639 should be released, resulting in de-caging of Y639. | ||
| + | |||
After this exposure, the cells were grown for an additional 16 hours, allowing the newly decaged T7 RNAP to transcribe the GFP reporter gene, which ultimately results in fluorescence. As can be seen in '''Figure 3''', while the fluorescence background at time zero is not as low as we would like, there is a clear and dramatic increase in fluorescence that is directly dependent on the amount of time the culture was exposed to 365 nm light. This result is consistent with previous results using this system (Chou et al. 2010). | After this exposure, the cells were grown for an additional 16 hours, allowing the newly decaged T7 RNAP to transcribe the GFP reporter gene, which ultimately results in fluorescence. As can be seen in '''Figure 3''', while the fluorescence background at time zero is not as low as we would like, there is a clear and dramatic increase in fluorescence that is directly dependent on the amount of time the culture was exposed to 365 nm light. This result is consistent with previous results using this system (Chou et al. 2010). | ||
| Line 122: | Line 129: | ||
The data from our photocage project provides strong evidence that we have, in fact, replicated the light-activated protein expression system that Chou et al. had constructed. | The data from our photocage project provides strong evidence that we have, in fact, replicated the light-activated protein expression system that Chou et al. had constructed. | ||
From the data, we are able to observe that GFP expression has a positive correlation with the amount of time that it is irradiated by light. Although the difference in fluorescence between 0 minutes and 1 minute appears to be negligible, there was a clear change in fluorescence at the 5 minute, 15 minute, and 30 minute time intervals. | From the data, we are able to observe that GFP expression has a positive correlation with the amount of time that it is irradiated by light. Although the difference in fluorescence between 0 minutes and 1 minute appears to be negligible, there was a clear change in fluorescence at the 5 minute, 15 minute, and 30 minute time intervals. | ||
| - | + | ||
| - | Although the increases in fluorescence are clearly present, there are still a few problems regarding the system. For one, the level of background expression of the | + | |
| + | Although the increases in fluorescence are clearly present, there are still a few problems regarding the system. For one, the level of background expression of the un-induced T7 RNAP is relatively high. The uninduced caged T7 RNAP has roughly 25 times more expression compared to the negative control. The relatively high level of fluorescence may be explained by the efficient nature of T7 RNAP. Even if a few polymerases were "decaged", they would have 16 hours to bind to the promoter and transcribe the RNA transcript for GFP. In order to decrease the level of background, a second amber stop codon was mutated into _______. | ||
The replication of such a system provides the foundation for potential projects for our future iGEM teams. By replacing the GFP reporter with coding sequences for other proteins, we will be able to explore novel applications of light-activated protein expression. | The replication of such a system provides the foundation for potential projects for our future iGEM teams. By replacing the GFP reporter with coding sequences for other proteins, we will be able to explore novel applications of light-activated protein expression. | ||
Revision as of 02:58, 17 October 2014
| |||||||||||||||||||||||||||||
 "
"