Team:Paris Saclay/Notebook/August/20
From 2014.igem.org
(→Digestion of CAD and PMCS5 at 37°C) |
m (→Photo of the Day) |
||
| (5 intermediate revisions not shown) | |||
| Line 21: | Line 21: | ||
|} | |} | ||
| - | === | + | ===Construction of the fusion protein (color)=== |
====PCR==== | ====PCR==== | ||
| Line 91: | Line 91: | ||
[https://2014.igem.org/Team:Paris_Saclay/Protocols/ protocol] | [https://2014.igem.org/Team:Paris_Saclay/Protocols/ protocol] | ||
| - | === | + | ===Lemon scent=== |
====Bacterial culture==== | ====Bacterial culture==== | ||
''by Laetitia'' | ''by Laetitia'' | ||
| + | [[File:20 08 bacterial culture.jpg| 500px|right]] | ||
Here are the results of the bacterial culture of yesterday | Here are the results of the bacterial culture of yesterday | ||
| - | |||
Bacteria with the pGEMTeasy without the DNA insert are blue and the bacteria with the pGEMTeasy and the DNA insert (PS, GS or LS gene) are white | Bacteria with the pGEMTeasy without the DNA insert are blue and the bacteria with the pGEMTeasy and the DNA insert (PS, GS or LS gene) are white | ||
| Line 114: | Line 114: | ||
Preculture 20ml of DH5a pPSI (apra 1/2000) | Preculture 20ml of DH5a pPSI (apra 1/2000) | ||
| - | |||
====CAD PCR==== | ====CAD PCR==== | ||
| Line 122: | Line 121: | ||
{| class="wikitable centre" width="50%" | {| class="wikitable centre" width="50%" | ||
| - | |+ PCR of | + | |+ PCR of CAD |
|- | |- | ||
! scope=col | components | ! scope=col | components | ||
! scope=col | volumes | ! scope=col | volumes | ||
|- | |- | ||
| - | |DNA of | + | |DNA of CAD |
|"very small amount" | |"very small amount" | ||
|- | |- | ||
| Line 197: | Line 196: | ||
#ladder 10µl | #ladder 10µl | ||
| - | [[File:Paris_Saclay_140820.jpg]] | + | [[File:Paris_Saclay_140820.jpg|center]] |
====Migration of pPS1 digested by PacI for the purification==== | ====Migration of pPS1 digested by PacI for the purification==== | ||
| Line 255: | Line 254: | ||
|2.5 µl | |2.5 µl | ||
|} | |} | ||
| + | |||
| + | ==Photo of the Day== | ||
| + | [[File:Paris Saclay 20_august.jpg|300px|center]] | ||
| + | |||
| + | '''Members present:''' | ||
| + | * Instructors and advisors: Alice, Solenne and Sylvie. | ||
| + | * Students: Eugène, Hoang Vu, Laëtitia, Mélanie, Romain, Sean and Terry. | ||
{{Team:Paris_Saclay/notebook_footer}} | {{Team:Paris_Saclay/notebook_footer}} | ||
Latest revision as of 15:08, 14 October 2014
Contents |
Wednesday 20th August
Lab work
Petri dishes
by Laetitia
| Antibiotic | Concentration |
|---|---|
| XGal | 80μg/ml |
| IPTG | 2*10-3 |
| Ampicillin | 10-3 |
Construction of the fusion protein (color)
PCR
| components | volumes |
|---|---|
| DNA | .5µl |
| GoTaq buffer 5X | 10μl |
| DMSO | 1.5µl |
| H2O | 22.75µl |
| MgCl2 | 4µl |
| dNTP | 1µl |
| iPS83 | 5µl |
| iPS84 | 5µl |
| GoTaq polymerase | .25µl |
| step | temperature (°C) | time (min) |
|---|---|---|
| 1 | 95 | 2 |
| 2 | 95 | .5 |
| 3 | 58 | .5 |
| 4 | 72 | 1.75 |
| 5 | 72 | 5 |
Cloning in pGEMTesay
by Sean
Lemon scent
Bacterial culture
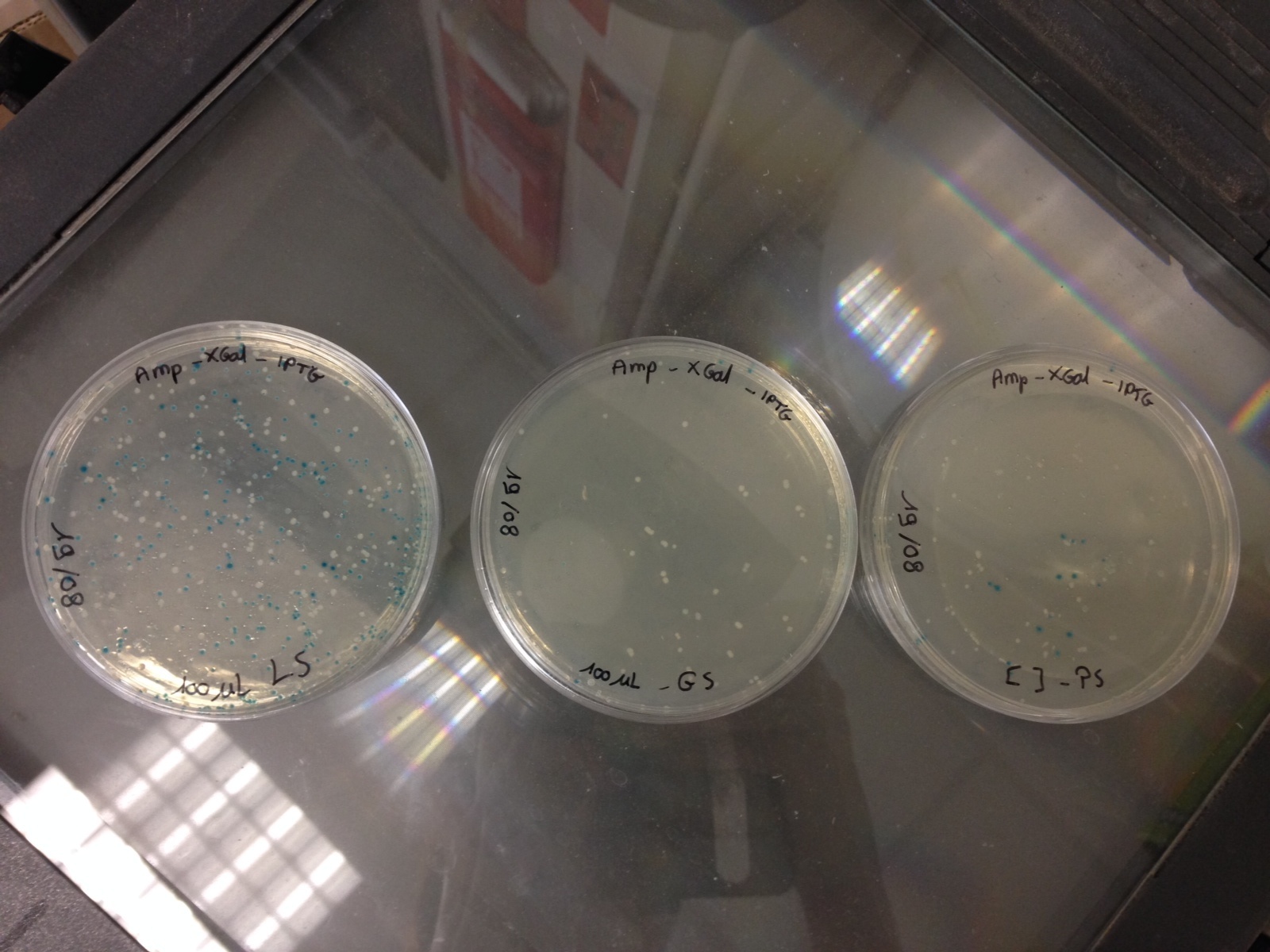
by Laetitia
Here are the results of the bacterial culture of yesterday
Bacteria with the pGEMTeasy without the DNA insert are blue and the bacteria with the pGEMTeasy and the DNA insert (PS, GS or LS gene) are white
Then, we have chosen 3 white colonies for each DNA insert and we have launched 3 corresponding liquid cultures :
- 5mL LB + ampiciline (1/1000e)
- 1 white colony
The 9 cultures are incubated at 37°C
by Melanie
Preculture 20ml of DH5a pPSI (apra 1/2000)
CAD PCR
Today we received our synthetic gene CAD (cinnamyl alcohol deshydrogenase) :)
so we do a PCR (with GoTaq):
| components | volumes |
|---|---|
| DNA of CAD | "very small amount" |
| GoTaq buffer 5X | 10μl |
| DMSO | 1.5µl |
| H2O | 23.25µl |
| MgCl2 | 4µl |
| dNTP | 1µl |
| iPS79 | 5µl |
| iPS80 | 5µl |
| GoTaq polymerase | .25µl |
| step | temperature (°C) | time (min) |
|---|---|---|
| 1 | 95 | 2 |
| 2 | 95 | .5 |
| 3 | 58 | .5 |
| 4 | 72 | 1.75 |
| 5 | 72 | 5 |
CAD cloning in pGEMTeasy
by Sean
Electrophoresis of pPSI digested by PacI
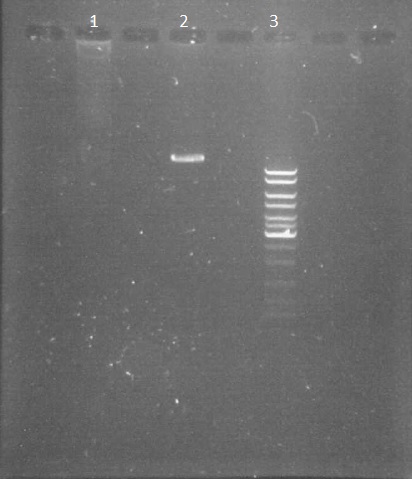
by Eugene
We made an electrophoresis of pPSI to verify if the digestion made 19th August has worked. It is useful before starting the purification of the plasmid.
- non-digested pPSI 3µl
- digested pPSI 3µl
- ladder 10µl
Migration of pPS1 digested by PacI for the purification
by Huang vu and Laetitia
pPS1 previously digested by PacI is filled inside an agarose gel :
-1g agarose
-1mL TAE
-2 µL BET
The totality of the sample is filled (around 38µL) and we filled 20µl of ladder. Migration 100V
After the migration, we want to purify the band corresponding of pPS1 digested by Pac1. While exposing the gel to UV, we will cut the band and save it inside an Eppendorf (weight = 1.080g) , before the purification
Digestion of CAD and PMCS5 at 37°C
by Eugene
| components | volumes |
|---|---|
| CAD | 5 μl |
| 10x buffer fastDigest | 1 μl |
| PacI | 0.5 µl |
| H2O | 3.5 µl |
| components | volumes |
|---|---|
| PMCS5 | 15 μl |
| 10x buffer fastDigest | 2 μl |
| PacI | 0.5 µl |
| H2O | 2.5 µl |
Photo of the Day
Members present:
- Instructors and advisors: Alice, Solenne and Sylvie.
- Students: Eugène, Hoang Vu, Laëtitia, Mélanie, Romain, Sean and Terry.
 "
"