|
|
| (142 intermediate revisions not shown) |
| Line 23: |
Line 23: |
| | </html> | | </html> |
| | == Sensing == | | == Sensing == |
| - | This part of the „BaKillus“-concept focuses on specific pathogen detection. Here we use quorum sensing to specifically recognize the presence of certain pathogens. The major aim of this subproject is transferring the quorum sensing two-component systems AgrC-II/AgrA of ''Staphylococcus aureus'' and ComDE of ''Streptococcus pneumoniae'' into ''Bacillus subtilis'', thus creating a pathogen detecting strain. With this strategy we will trigger our killing strategies only if a threshold concentration of autoinducers, produced by pathogens, is present in the environment. Thereby we aim to drastically increase the specificity of our antimicrobial strategies compared to commonly used broad-spectrum antibiotic therapy. | + | This part of the BaKillus-concept focuses on specific pathogen detection. Here we use quorum sensing to specifically recognize the presence of certain pathogens. The major aim of this subproject is transferring the quorum sensing two-component systems AgrC-II/AgrA of ''Staphylococcus aureus'' and ComDE of ''Streptococcus pneumoniae'' into ''Bacillus subtilis'', thus creating a pathogen detecting strain. With this strategy we will trigger our killing strategies only if a threshold concentration of autoinducers, produced by pathogens, is present in the environment Thereby we aim to drastically increase the specificity of our antimicrobial strategies compared to commonly used broad-spectrum antibiotic therapy. |
| | | | |
| | <html> | | <html> |
| Line 40: |
Line 40: |
| | | | |
| | ===Quorum sensing in ''S. aureus''=== | | ===Quorum sensing in ''S. aureus''=== |
| - | Quorum Sensing is a mechanism of bacterial cell-cell communication, granting bacteria the ability to measure the size of their surrounding cell population, and react to this by upregulation of specific genes. | + | Quorum Sensing is a mechanism of bacterial cell-cell communication, granting bacteria the ability to measure the density of their surrounding cell population, and react to this by upregulation of specific genes. |
| - | In ‘’S. aureus’’ this process is one of the major regulator of its virulence, making sure that energy-costly virulence factors are only produced when a large enough number of pathogens is present to overcome host-defence mechanisms. | + | In ''S. aureus'' this process is one of the major regulator of its virulence, making sure that energy-costly virulence factors are only produced when a large enough number of pathogens is present to overcome host-defence mechanisms. |
| | | | |
| - | In this mechanism a certain peptide termed autoinducing peptide (AIP) gets produced from a propeptide called AgrD and further processed and exported by a membrane protein AgrB to create a cyclic AIP. When a certain threshold concentration of AIP in the environment is reached, AIPs activate the receptor-histidine kinase AgrC. | + | In this mechanism a certain peptide termed autoinducing peptide (AIP) is produced from a pro-peptide called AgrD and further processed and exported by a membrane protein AgrB to create a cyclic AIP. When a certain threshold concentration of AIP in the environment is reached, AIPs activate the receptor-histidine kinase AgrC. |
| - | Upon induction AgrC phosphorylates the intracellular response regulator AgrA with its cytosolic histidine kinase domain [http://www.annualreviews.org/doi/abs/10.1146/annurev.genet.42.110807.091640 (3) ]. Phosphorylated AgrA then binds to distinct sites in the P2 and P3 promoter-region and activates these promoters. [http://jb.asm.org/content/193/21/6020 (2)] | + | Upon induction AgrC phosphorylates the intracellular response regulator AgrA with its cytosolic histidine kinase domain [http://www.annualreviews.org/doi/abs/10.1146/annurev.genet.42.110807.091640 (3)]. Phosphorylated AgrA then binds to distinct sites in the P2 and P3 promoter-region and activates these promoters. [http://jb.asm.org/content/193/21/6020 (2)] |
| - | The P2 promoter activates transcription of the AgrBDCA-Operon and thereby represents a positive feedback-loop. The P3 promoter activates transcription of the RNA-III transcript, involved of virulence gene expression [http://jb.asm.org/content/182/22/6517.full (4)]). | + | The P2 promoter activates transcription of the ''agrBDCA''-Operon and thereby represents a positive feedback-loop. The P3 promoter activates transcription of the RNA-III transcript, involved in virulence gene expression [http://jb.asm.org/content/182/22/6517.full (4)]). |
| | | | |
| | [[File:LMU14_Agr_QS.jpg|thumb|800px|center|'''Fig.1''': Quorum sensing in ''Staphylococcus aureus''. Taken from Novick and Geisinger, "Quorum Sensing in Staphylococci", Annual Review of Genetics, Vol. 42: 541-564 (2008)]] | | [[File:LMU14_Agr_QS.jpg|thumb|800px|center|'''Fig.1''': Quorum sensing in ''Staphylococcus aureus''. Taken from Novick and Geisinger, "Quorum Sensing in Staphylococci", Annual Review of Genetics, Vol. 42: 541-564 (2008)]] |
| | | | |
| - | Interestingly, accumulated mutations in the hypervariable region of the ''agrBDCA''-operon have led to four different pherotypes of ''S. aureus'', using different agrC-types, and producing different AIPs (AIPI, AIPII, AIPIII, AIPIV), which cross-inhibit each other and might correlate with certain diseases associated with ''S. aureus'' [http://www.sciencemag.org/content/287/5452/391.full.html (5)]. | + | Interestingly, accumulated mutations in the hypervariable region of the ''agrBDCA''-operon have led to four different pherotypes of ''S. aureus'', using different AgrC-types, and producing different AIPs (AIPI, AIPII, AIPIII, AIPIV), which cross-inhibit each other and might correlate with certain diseases associated with ''S. aureus'' [http://www.sciencemag.org/content/287/5452/391.full.html (5)]. |
| | | | |
| | For this project we used genomic DNA derived from the strain ''Staphylococcus aureus'' N315, which is a MRSA strain, belonging to the AIP-II group. | | For this project we used genomic DNA derived from the strain ''Staphylococcus aureus'' N315, which is a MRSA strain, belonging to the AIP-II group. |
| Line 55: |
Line 55: |
| | ==''Streptococcus pneumoniae''== | | ==''Streptococcus pneumoniae''== |
| | ===Virulence of ''S. pneumoniae'' === | | ===Virulence of ''S. pneumoniae'' === |
| - | All over the world, ''Streptococcus pneumoniae'' causes the life threating invasive diseases pneumonia, sepsis and meningitis. In early childhood the diplococcus colonizes the epithelium of the upper respiratory tract as a commensal bacterium. Under certain circumstances, it gets a threat by tissue invasion. Still up to date, not much is known regarding the mechanism behind the shifting from a colonizer organism to an invader or how ''S. pneumoniae'' is able to cross the blood-brain barrier to cause meningitis.[http://www.ncbi.nlm.nih.gov/pubmed/19880020] | + | All over the world, ''Streptococcus pneumoniae'' causes the life threating invasive diseases pneumonia, sepsis and meningitis. In early childhood the diplococcus colonizes the epithelium of the upper respiratory tract as a commensal bacterium. Under certain circumstances, it gets a threat by tissue invasion. Still up to date, not much is known regarding the mechanism behind the shifting from a colonizer organism to an invader or how ''S. pneumoniae'' is able to cross the blood-brain barrier to cause meningitis [http://www.ncbi.nlm.nih.gov/pubmed/19880020 (6)]. |
| - | To cross the line from an opportunist to a pathogen, ''S. pneumoniae'' needs to express specific virulence factors in a coordinated way. Apart from virulence factors like the polysaccharide capsule, the pore forming toxin pneumolysin or surface proteins like PspA and PspC, also biofilm production plays an important role for persistence in the human nasopharynx and contributes to pneumonia and meningitis [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1618759/]. | + | To cross the line from an opportunist to a pathogen, ''S. pneumoniae'' needs to express specific virulence factors in a coordinated way. Apart from virulence factors like the polysaccharide capsule, the pore forming toxin pneumolysin or surface proteins like PspA and PspC, also biofilm production plays an important role for persistence in the human nasopharynx and contributes to pneumonia and meningitis [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1618759 (7)]. |
| | | | |
| | One aspect of its infectiousness is the ability to perform natural transformation. Thus, it can take up extracellular DNA containing the information for resistance against diverse antibiotics and permanently incorporate the DNA into its genome via recombination. The main [https://2014.igem.org/Team:LMU-Munich/Project/Problem problem] of treating ''S. pneumoniae'' infections is the increasing spread of ß-lactam resistance. Luckily, the molecular mechanism behind natural transformation in ''S. pneumoniae'' is the focus of current research. | | One aspect of its infectiousness is the ability to perform natural transformation. Thus, it can take up extracellular DNA containing the information for resistance against diverse antibiotics and permanently incorporate the DNA into its genome via recombination. The main [https://2014.igem.org/Team:LMU-Munich/Project/Problem problem] of treating ''S. pneumoniae'' infections is the increasing spread of ß-lactam resistance. Luckily, the molecular mechanism behind natural transformation in ''S. pneumoniae'' is the focus of current research. |
| | | | |
| | ===Natural Transformation in ''S. pneumoniae''=== | | ===Natural Transformation in ''S. pneumoniae''=== |
| - | In ''S. pneumoniae'', biofilm production and competence for natural genetic transformation are triggered by a two component regulatory system, responsive to environmental stimuli. There, a peptide pheromone called competence-stimulating-peptide (CSP) binds to and consequently activates the membrane-embedded sensor kinase ComD by changing the conformation of the polytopic kinase. ComD autophosphorylates and subsequently transphosphorylates the cognate response regulator ComE. ComE acts as transcription activator via binding to the -10 promotor regions of early and late competence genes. These ComE binding sites (CEbs) consist of a 9 bp direct repeat seperated by a stretch of 12 nucleotides. | + | In ''S. pneumoniae'', biofilm production and competence for natural genetic transformation are triggered by a two component regulatory system, responsive to environmental stimuli. There, a peptide pheromone called competence-stimulating-peptide (CSP) binds to and consequently activates the membrane-embedded sensor kinase ComD by changing the conformation of the polytopic kinase. ComD autophosphorylates and subsequently transphosphorylates the cognate response regulator ComE. ComE acts as transcription activator via binding to the -10 promoter regions of early and late competence genes. These ComE binding sites (CEbs) consist of a 9 bp direct repeat seperated by a stretch of 12 nucleotides. |
| | | | |
| | [[File:LMU14 ComDE.png|thumb|left|700px|'''Fig. 2''': Competence regulation in ''Streptococcus pneumoniae''. See text for more details. (Jonsborg et al. 2009)]] | | [[File:LMU14 ComDE.png|thumb|left|700px|'''Fig. 2''': Competence regulation in ''Streptococcus pneumoniae''. See text for more details. (Jonsborg et al. 2009)]] |
| | | | |
| - | A positive feedback loop ensures the availability of the two component system by activating the expression of the ''comCDE'' operon, where ''comDE'' encodes for the two component system ComDE and ''comC'' for the precursor peptide pre-CSP. Pre-CSP is cleaved to the mature peptide pheromone and transported to the extraplasmic room by the ABC transporter ComAB, whose expression is also activated by ComE. [http://www.ncbi.nlm.nih.gov/pubmed/?term=regulation+of+natural+genetic+transformation+and+acquisition+of+transforming+DNA]. | + | A positive feedback loop ensures the availability of the two component system by activating the expression of the ''comCDE'' operon, where ''comDE'' encodes for the two component system ComDE and ''comC'' for the precursor peptide pre-CSP. Pre-CSP is cleaved to the mature peptide pheromone and transported to the extraplasmic room by the ABC transporter ComAB, whose expression is also activated by ComE. [http://www.ncbi.nlm.nih.gov/pubmed/?term=regulation+of+natural+genetic+transformation+and+acquisition+of+transforming+DNA (8)]. |
| - | In addition, ComE~P is able to trigger the expression of ''comX'' and ''comW '', encoding for the alternative σ factor X, and its stability factor ComW. This alternative σ factor is required for the transcription of the so called late competence genes involved in the process of DNA uptake and recombination. | + | In addition, ComE~P is able to trigger the expression of ''comX'' and ''comW '', encoding for the alternative σ-factor X, and its stability factor ComW. This alternative σ-factor is required for the transcription of the so called late competence genes involved in the process of DNA uptake and recombination. |
| | | | |
| | | | |
| - | We are working with the genomic DNA of the nonencapsulated apathogenic strain R6 of ''S. pneumoniae'', whose genome is completely sequenced [http://www.ncbi.nlm.nih.gov/pubmed/11544234]. CSP is a 17 amino acid long linear protein with the following amino acid sequence: Glu-Met-Arg-Leu-Ser-Lys-Phe-Arg-Asp-Phe-Ile-Leu-Gln-Arg-Lys-Lys-OH [http://www.microbesonline.org/cgi-bin/fetchLocus.cgi?locus=134925&disp=4]with a charged N-terminal residue and a positively charged C-terminal tail [http://www.ncbi.nlm.nih.gov/pubmed/16484185]. | + | We are working with the genomic DNA of the nonencapsulated apathogenic strain R6 of ''S. pneumoniae'', whose genome is completely sequenced [http://www.ncbi.nlm.nih.gov/pubmed/11544234 (9)]. CSP is a 17 amino acid long linear protein with the following amino acid sequence: Glu-Met-Arg-Leu-Ser-Lys-Phe-Phe-Arg-Asp-Phe-Ile-Leu-Gln-Arg-Lys-Lys-OH [http://www.microbesonline.org/cgi-bin/fetchLocus.cgi?locus=134925&disp=4 (10)] with a charged N-terminal residue and a positively charged C-terminal tail [http://www.ncbi.nlm.nih.gov/pubmed/16484185 (11)]. |
| | | | |
| | ===Sources=== | | ===Sources=== |
| | <small> | | <small> |
| - | [5] van der Poll, T., & Opal, S. M. (2009). Pathogenesis, treatment, and prevention of pneumococcal pneumonia. Lancet, 374(9700), 1543-1556. doi: 10.1016/S0140-6736(09)61114-4 | + | [1] den Heijer, C. D., van Bijnen, E. M., Paget, W. J., Pringle, M., Goossens, H., Bruggeman, C. A., . . . Team, A. S. (2013). Prevalence and resistance of commensal Staphylococcus aureus, including meticillin-resistant S aureus, in nine European countries: a cross-sectional study. Lancet Infect Dis, 13(5), 409-415. doi: 10.1016/S1473-3099(13)70036-7 |
| | | | |
| - | [6] Oggioni, M. R., Trappetti, C., Kadioglu, A., Cassone, M., Iannelli, F., Ricci, S., . . . Pozzi, G. (2006). Switch from planktonic to sessile life: a major event in pneumococcal pathogenesis. Mol Microbiol, 61(5), 1196-1210. doi: 10.1111/j.1365-2958.2006.05310.x | + | [2] Reyes, D., Andrey, D. O., Monod, A., Kelley, W. L., Zhang, G., & Cheung, A. L. (2011). Coordinated regulation by AgrA, SarA, and SarR to control agr expression in Staphylococcus aureus. J Bacteriol, 193(21), 6020-6031. doi: 10.1128/JB.05436-11 |
| | | | |
| - | [7] Johnsborg, O., & Havarstein, L. S. (2009). Regulation of natural genetic transformation and acquisition of transforming DNA in Streptococcus pneumoniae. FEMS Microbiol Rev, 33(3), 627-642. | + | [3] Novick, R. P., & Geisinger, E. (2008). Quorum sensing in staphylococci. Annu Rev Genet, 42, 541-564. doi: 10.1146/annurev.genet.42.110807.091640 |
| | | | |
| - | [8] Hoskins, J., Alborn, W. E., Jr., Arnold, J., Blaszczak, L. C., Burgett, S., DeHoff, B. S., . . . Glass, J. I. (2001). Genome of the bacterium Streptococcus pneumoniae strain R6. J Bacteriol, 183(19), 5709-5717. doi: 10.1128/JB.183.19.5709-5717.2001 | + | [4] Jarraud, S., Lyon, G. J., Figueiredo, A. M., Lina, G., Vandenesch, F., Etienne, J., . . . Novick, R. P. (2000). Exfoliatin-producing strains define a fourth agr specificity group in Staphylococcus aureus. J Bacteriol, 182(22), 6517-6522. |
| | | | |
| - | [9] Sequence obtained from http://www.microbesonline.org | + | [5] R. P. Novick, H.F. Ross, A. M. S. Figueiredo, G. Abramochkin (2000) Activation and Inhibition of the Staphylococcal AGR System. Science 21 Vol. 287 no. 5452 p. 391 |
| | | | |
| - | [10] Johnsborg, O., Kristiansen, P. E., Blomqvist, T., & Havarstein, L. S. (2006). A hydrophobic patch in the competence-stimulating Peptide, a pneumococcal competence pheromone, is essential for specificity and biological activity. J Bacteriol, 188(5), 1744-1749. doi: 10.1128/JB.188.5.1744-1749.2006 | + | [6] van der Poll, T., & Opal, S. M. (2009). Pathogenesis, treatment, and prevention of pneumococcal pneumonia. Lancet, 374(9700), 1543-1556. doi: 10.1016/S0140-6736(09)61114-4 |
| | + | |
| | + | [7] Oggioni, M. R., Trappetti, C., Kadioglu, A., Cassone, M., Iannelli, F., Ricci, S., . . . Pozzi, G. (2006). Switch from planktonic to sessile life: a major event in pneumococcal pathogenesis. Mol Microbiol, 61(5), 1196-1210. doi: 10.1111/j.1365-2958.2006.05310.x |
| | + | |
| | + | [8] Johnsborg, O., & Havarstein, L. S. (2009). Regulation of natural genetic transformation and acquisition of transforming DNA in Streptococcus pneumoniae. FEMS Microbiol Rev, 33(3), 627-642. |
| | + | |
| | + | [9] Hoskins, J., Alborn, W. E., Jr., Arnold, J., Blaszczak, L. C., Burgett, S., DeHoff, B. S., . . . Glass, J. I. (2001). Genome of the bacterium Streptococcus pneumoniae strain R6. J Bacteriol, 183(19), 5709-5717. doi: 10.1128/JB.183.19.5709-5717.2001 |
| | + | |
| | + | [10] Sequence obtained from http://www.microbesonline.org |
| | + | |
| | + | [11] Johnsborg, O., Kristiansen, P. E., Blomqvist, T., & Havarstein, L. S. (2006). A hydrophobic patch in the competence-stimulating Peptide, a pneumococcal competence pheromone, is essential for specificity and biological activity. J Bacteriol, 188(5), 1744-1749. doi: 10.1128/JB.188.5.1744-1749.2006 |
| | | | |
| | </small> | | </small> |
| Line 100: |
Line 110: |
| | | | |
| | | | |
| - | The two component system [http://parts.igem.org/Part:BBa_K1351036 ''agrCII-A''] for ''S. aureus'' has been designed as a composite part derived from Staphylococcus aureus N315 gDNA. | + | The two component system [http://parts.igem.org/Part:BBa_K1351036 ''agrCII-A''] for ''S. aureus'' has been designed as a composite part derived from ''Staphylococcus aureus'' N315 gDNA. |
| - | The two intermediated parts agrC-II and agrA were amplified by PCR using primers with modified RFC25 prefix and suffix. And then fused by using the restriction sites EcoRI and SpeI for agrC-II, as well as XbaI and PstI for agrA, thus creating the composite part [http://parts.igem.org/Part:BBa_K1351036 ''agrCII-A'']. | + | The two intermediated parts ''agrC-II'' and ''agrA'' were amplified by PCR using primers with modified RFC25 prefix and suffix. And then fused by using the restriction sites EcoRI and SpeI for ''agrC-II'', as well as XbaI and PstI for ''agrA'', thus creating the composite part [http://parts.igem.org/Part:BBa_K1351036 ''agrCII-A'']. |
| | | | |
| - | The operon encoding for the histidine sensor kinase ComD and the response regulator ComE ''comDE'' for ''S. pneumoniae'' sensing was designed in two different ways: On one hand, the native ''comDE'' operon with its native ribosome binding site (RBS) ([http://parts.igem.org/Part:BBa_K1351015 BBa_K1351015]), on the other hand the''comDE'' operon with the ''B. subtilis'' adapted RBS (TAAGGAGG) in the BioBrick standard rcf25 ([http://parts.igem.org/Part:BBa_K1351016 BBa_K1351016]). Unfortunately, the operon contains two EcoRI and one SpeI restriction site. The attempts to mutagenize these restriction sites were not complettely successfull as one recognition site remained. Thus, the ''comDE'' operon could not been sent to the registry - but this will be accomplished as soon as possible. | + | The operon encoding for the histidine sensor kinase ComD and the response regulator ComE ''comDE'' for ''S. pneumoniae'' sensing was designed in two different ways: On one hand, the native ''comDE'' operon with its native ribosome binding site (RBS) ([http://parts.igem.org/Part:BBa_K1351015 BBa_K1351015]), on the other hand the''comDE'' operon with the ''B. subtilis'' adapted RBS (TAAGGAGG) in the BioBrick standard RFC25 ([http://parts.igem.org/Part:BBa_K1351016 BBa_K1351016]). Unfortunately, the operon contains two EcoRI and one SpeI restriction site. The attempts to mutagenize these restriction sites were not completed successfully in time as one recognition site remained. Thus, the ''comDE'' operon could not been sent to the registry - but this will be accomplished as soon as possible. |
| | | | |
| | ===QS dependent Promoters=== | | ===QS dependent Promoters=== |
| | | | |
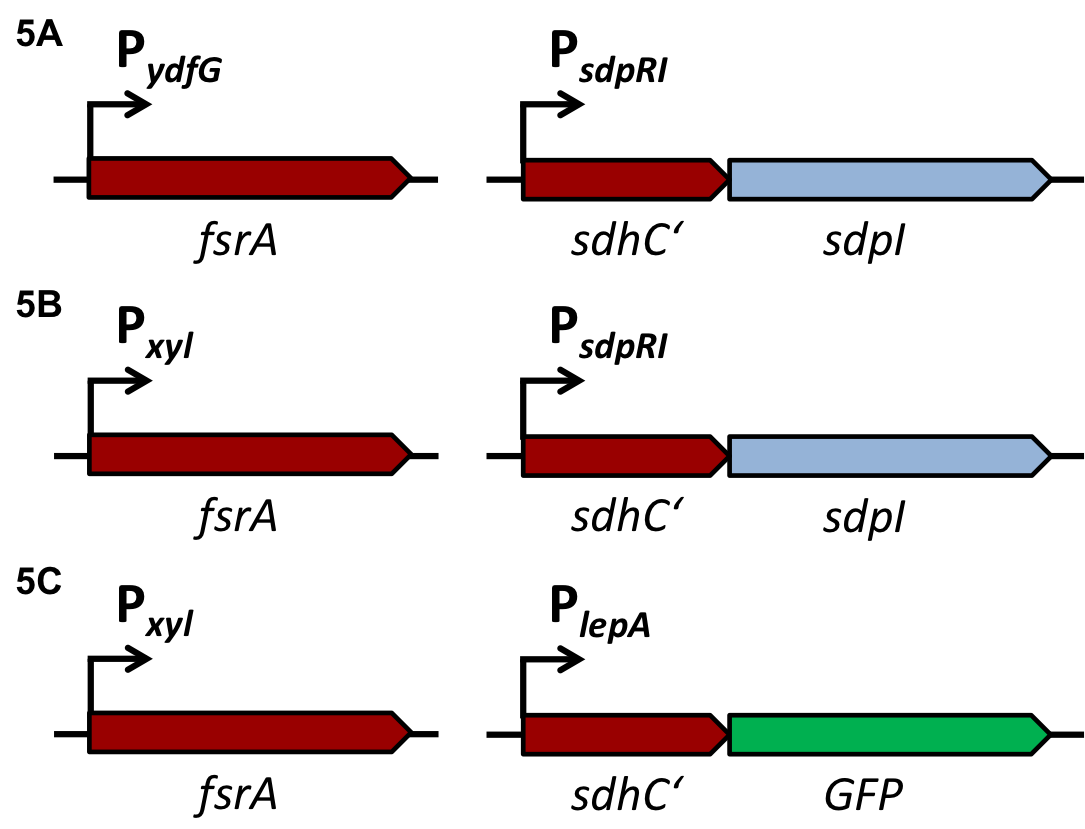
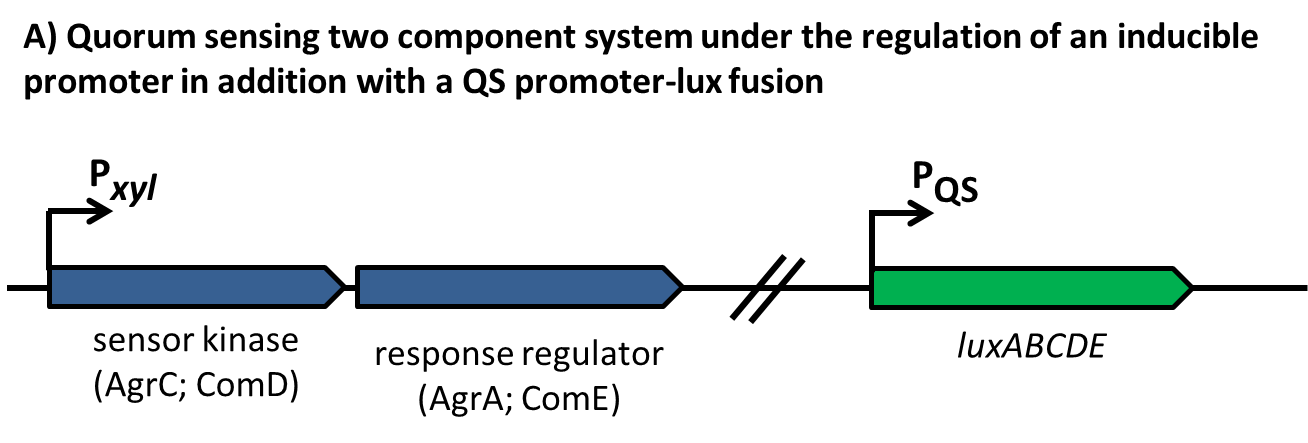
| - | In ''S. aureus'', the AgrA-sensitive promoters P2 [BBa_K1351037] and P3 [BBa_K1351038] with four AgrA binding sites downstream of the -35 signals were used for evaluation. The intergenic region with suspected regulator binding sites is shown in Fig. 3 | + | In ''S. aureus'', the AgrA-sensitive promoters P2 [http://parts.igem.org/Part:BBa_K1351037 BBa_K1351037] and P3 [http://parts.igem.org/Part:BBa_K1351038 BBa_K1351038] with four AgrA binding sites downstream of the -35 signals were used for evaluation. The intergenic region with suspected regulator binding sites is shown in Fig. 3 |
| | [[File:LMU14 P2P3.jpg|center|thumb|700px|'''Fig. 3''': AgrA dependent promoters P2 and P3 with the intergenic region containing several regulator binding sites for AgrA (arrows) and SarA/R (dashed lines) (figure taken from Reyes et al. 2000)]] | | [[File:LMU14 P2P3.jpg|center|thumb|700px|'''Fig. 3''': AgrA dependent promoters P2 and P3 with the intergenic region containing several regulator binding sites for AgrA (arrows) and SarA/R (dashed lines) (figure taken from Reyes et al. 2000)]] |
| | | | |
| Line 143: |
Line 153: |
| | As expected according to the strength of the -35 and -10 signals, P2 is the stronger promoter showing a higher background, compared to P3. | | As expected according to the strength of the -35 and -10 signals, P2 is the stronger promoter showing a higher background, compared to P3. |
| | | | |
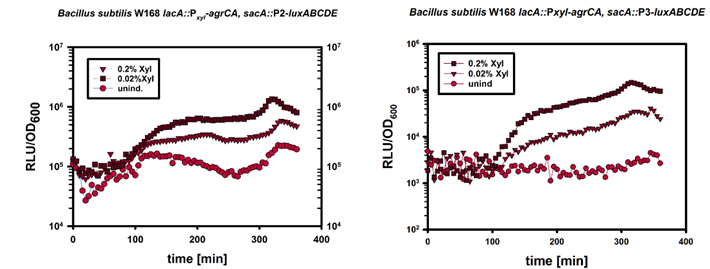
| - | In a next step we induced the expression of ''PxylA-agrCII-A'' by adding 0%, 0.02%, 0.2% Xylose at an OD600 of about 0.1. | + | In a next step we induced the expression of ''PxylA-agrCII-A'' by adding 0%, 0.02%, 0.2% Xylose at an OD<sub>600</sub> of about 0.1. |
| - | The result is depicted in Figure 4. The result is in accordance with our expactations since strong overexpression of the two-component system should lead to unspecific induction of the P2 and P3 promoters, and thereby can be used as a method to measure relative expression of ''agrC-IIA''. | + | The result is depicted in Figure 5. This result is in accordance with our expactations since strong overexpression of the two-component system should lead to unspecific induction of the P2 and P3 promoters, and thereby can be used as a method to measure relative expression of ''agrC-IIA''. |
| | | | |
| - | Using the strategy outlined in design, we tried to induce the AgrC-IIA two-component system, with synthetic AIP-II. The synthetic peptids have been dissolved in 100% DMSO to 1mM concentration and further diluted in dH20. | + | [[File:LMU14_P2_Xyl.png|thumb|800px|center|'''Fig.5''': Luminescence Reporterassay of the strain W168 ''lacA::PxylA-agrCA; sacA::''P2/P3-''luxABCDE'']] |
| - | The constructs have then been induced with a range of different concentrations of AIP-II at an OD600 of 0.1. | + | Using the strategy outlined in design, we tried to induce the AgrC-IIA two-component system, with synthetic AIP-II. The synthetic peptids have been dissolved in 100% DMSO to 1 mM concentration and further diluted in dH<sub>2</sub>0. |
| - | The result is shown in Figure 5. Unfortunatly we were not able to show induction of the system by addition of AIP-II. Several other approaches like the addition of a protease inhibitor cocktail to the growth media, several washing steps, using LB instead of MCSE inducing with 50µM AIP-II did not change this observation. | + | The constructs have then been induced with a range of different concentrations of AIP-II at an OD<sub>600</sub> of 0.1. |
| | + | The result is shown in Figure 6. Unfortunatly we were not able to show induction of the system by addition of AIP-II. Several other approaches like the addition of a protease inhibitor cocktail to the growth media, several washing steps, using LB instead of MCSE or inducing with 50 µM AIP-II did not change this observation. |
| | | | |
| - | There are several possible reasons why the induction didn't work as expected. One idea is that the extracellular proteases produced by ''Bacillus subtilis'' destroy the synthetic AIPs. So we tried adding protease inhibitor cocktails to the growth media and using the protease deficient strain ''Bacillus subtillis'' WB700. We successfully transformed the WB700 strain with the reporter construct pBS4S-P''xylA-agrCA''. However the reporterconstructs pBS3C-P2/P3-''luxABCDE'' refused transformation. | + | [[File:LMU14_Caind.png|thumb|800px|center|'''Fig.6''': Luminescence Reporterassay of the strain W168 ''lacA::PxylA-agrCA; sacA::''P2/P3-''luxABCDE'' induced with AIP-II]] |
| | + | There are several possible reasons why the induction didn't work as expected. One idea is that the extracellular proteases produced by ''Bacillus subtilis'' destroy the synthetic AIPs. So we tried adding protease inhibitor cocktails to the growth media and using the protease deficient strain ''Bacillus subtilis'' WB700. We successfully transformed the WB700 strain with the reporter construct pBS4S-P''xylA-agrCA''. However the reporter constructs pBS3C-P2/P3-''luxABCDE'' refused transformation. |
| | + | All in all this set of constructs needs several further protocol optimisations and testing. |
| | | | |
| | ==''S. pneumoniae'' Sensing== | | ==''S. pneumoniae'' Sensing== |
| Line 156: |
Line 169: |
| | First, we optimized the BioBrick <bbpart>BBa_K316018</bbpart> encoding for the ''comC'' promoter [http://parts.igem.org/wiki/index.php/Part:BBa_K316018] by sending our BioBrick <bbpart>BBa_K1351024</bbpart> to the registry. For the previous version, no DNA was available. | | First, we optimized the BioBrick <bbpart>BBa_K316018</bbpart> encoding for the ''comC'' promoter [http://parts.igem.org/wiki/index.php/Part:BBa_K316018] by sending our BioBrick <bbpart>BBa_K1351024</bbpart> to the registry. For the previous version, no DNA was available. |
| | | | |
| - | The BIG result we obtained in the night before wiki freeze performing a [https://2014.igem.org/Team:LMU-Munich/Notebook/Protocols luminescence assay] with different constructs in ''B. subtilis'' P<sub>''xyl''</sub> - ''comDE'' P<sub>''QS''</sub> - lux based on <bbpart>BBa_K1351016</bbpart>. Three different P<sub>''QS''</sub> were tested: P<sub>''comC''</sub>, P<sub>''comAB''</sub>, P<sub>''mreA''</sub>. The ''comDE'' operon expression was induced with 0 %, 0,02 % and 0,2 % xylose. Adding 0 mg/ml CSP, 100 mg/ml CSP and 1000 mg/ml CSP ''luxABCDE'' expression is only activated if ''B. subtilis'' was able to sense CSP due to the ''comDE'' operon. | + | The BIG result we obtained in the night of the wiki freeze performing a [https://2014.igem.org/Team:LMU-Munich/Notebook/Protocols luminescence assay] with different constructs in ''B. subtilis'' P<sub>''xyl''</sub> - ''comDE'' P<sub>''QS''</sub> - ''lux'' based on [http://parts.igem.org/Part:BBa_K1351016 BBa_K1351016]. Three different P<sub>''QS''</sub> were tested: P<sub>''comC''</sub>, P<sub>''comAB''</sub>, P<sub>''mreA''</sub>. The ''comDE'' operon expression was induced with 0 %, 0,02 % and 0,2 % xylose. Adding 0 mg/ml CSP, 100 mg/ml CSP and 1000 mg/ml CSP ''luxABCDE'' expression is only activated if ''B. subtilis'' was able to sense CSP due to the ''comDE'' operon. |
| - | For P<sub>''mreA''</sub>, no significant luminescence output could be measured concerning differing CSP concentrations. Nevertheless, at a xylose concentration of 0,02 % and induction of 1000 mg/ml CSP P<sub>''comC''</sub> and P<sub>''comAB''</sub> showed a 3 fold higher luminescence output compared with an induction of 0 mg/ml and 100 mg/ml CSP. | + | For P<sub>''mreA''</sub>, no significant luminescence output could be measured concerning differing CSP concentrations. Nevertheless, at a xylose concentration of 0,02 % and induction of 1000 mg/ml CSP P<sub>''comC''</sub> and P<sub>''comAB''</sub> showed a 3 fold increased luminescence output compared with an induction of 0 mg/ml and 100 mg/ml CSP. |
| | | | |
| | | | |
| Line 373: |
Line 386: |
| | }] | | }] |
| | }); | | }); |
| | + | |
| | + | $('#sensing-killing-graph-1').highcharts({ |
| | + | chart: { |
| | + | type: 'column', |
| | + | backgroundColor: null, |
| | + | plotBorderColor: '#000000', |
| | + | plotBorderWidth: 2 |
| | + | }, |
| | + | color:'#33CC33', |
| | + | title: { |
| | + | useHTML:true, |
| | + | text: 'P<sub style="font-style:italic">xyl</sub><span style="font-style:italic"> - comDE</span> | P<sub style="font-style:italic">comAB</sub><span style="font-style:italic"> - lux</span>', |
| | + | style: { |
| | + | fontSize: '28px'/*, |
| | + | fontFamily: 'Verdana, sans-serif',*/ |
| | + | |
| | + | }, |
| | + | }, |
| | + | subtitle: { |
| | + | text: null, |
| | + | style: { |
| | + | fontSize: '18px'/*, |
| | + | fontFamily: 'Verdana, sans-serif',*/ |
| | + | |
| | + | }, |
| | + | }, |
| | + | xAxis: { |
| | + | type: 'category', |
| | + | labels: { |
| | + | rotation: 0, |
| | + | style: { |
| | + | fontSize: '18px'/*, |
| | + | fontFamily: 'Verdana, sans-serif',*/ |
| | + | } |
| | + | } |
| | + | }, |
| | + | yAxis: { |
| | + | type: 'linear', |
| | + | gridLineWidth: 0, |
| | + | minorGridLineWidth: 0, |
| | + | min: 1, |
| | + | title: { |
| | + | text: 'RLU / OD<sub>600</sub>', |
| | + | useHTML:true, |
| | + | style: { |
| | + | fontSize: '18px', |
| | + | fontFamily: 'Arial, Helvetica, sans-serif', |
| | + | |
| | + | } |
| | + | } |
| | + | }, |
| | + | credits: false, |
| | + | legend: { |
| | + | enabled: false |
| | + | }, |
| | + | plotOptions: { |
| | + | column: { |
| | + | pointPadding: 0.3, |
| | + | color: '#33CC33', |
| | + | borderWidth:2, |
| | + | borderColor:'#000000' |
| | + | } |
| | + | }, |
| | + | |
| | + | tooltip: { |
| | + | shared: true, |
| | + | headerFormat: '<span style="font-size: 13px;font-weight:bold;"> {point.key}</span><br/>', |
| | + | useHTML: true |
| | + | }, |
| | + | |
| | + | |
| | + | series: [{ |
| | + | name: 'Data', |
| | + | data: [ |
| | + | ['0ng/ml CSP', 12869.7367 |
| | + | ], |
| | + | ['100 ng/ml CSP', 22602.16186 |
| | + | ], |
| | + | ['1000 ng/ml CSP', 103594.3583 |
| | + | ] |
| | + | ], |
| | + | marker: { |
| | + | enabled: false |
| | + | }, |
| | + | tooltip: { |
| | + | pointFormat: '<span style="font-size:13px">{series.name}</span>: {point.y}<br/>', |
| | + | valueDecimals: 0 |
| | + | } |
| | + | }, { |
| | + | color: '#FF0000', |
| | + | name: 'Error range', |
| | + | type: 'errorbar', |
| | + | data: [[12273,13466],[18394,26810],[91956,115231]], |
| | + | tooltip: { |
| | + | pointFormat: '<span style="font-size:13px">{series.name}</span>: {point.low}-{point.high}' |
| | + | }, |
| | + | stemWidth: 5, |
| | + | whiskerLength: 5, |
| | + | color: '#000000' |
| | + | }] |
| | + | }); |
| | + | |
| | }); | | }); |
| | </script> | | </script> |
| Line 387: |
Line 502: |
| | As part of the BaKillus strategy, we want to reduce the selective pressure on non-pathogenic bacteria to prevent resistance development. Towards this end, BaKillus is equipped with pathogen-binding peptides on its surface, thereby adhering specifically to the targeted bacteria. This approach ensures high local concentration of killing factors while minimizing harmful effects on untargeted species. | | As part of the BaKillus strategy, we want to reduce the selective pressure on non-pathogenic bacteria to prevent resistance development. Towards this end, BaKillus is equipped with pathogen-binding peptides on its surface, thereby adhering specifically to the targeted bacteria. This approach ensures high local concentration of killing factors while minimizing harmful effects on untargeted species. |
| | | | |
| - | Surface display of the pathogen-binding peptides is achieved by translational fusions to naturally occurring surface proteins of ''B. subtilis''. Cell-wall binding modules of autolysins are used for non-covalent interaction with the ''B. subtilis'' cell wall, whereas a sortase-based system functions via covalent attachment to the cell wall. In both cases, a linker between cell wall anchor and peptide allows flexibility of the fusion protein and increases its range. Peptides binding to different receptors on the surface of pathogens mediate specific binding. | + | Surface display of the pathogen-binding peptides is achieved by translational fusions to naturally occurring surface proteins of ''B. subtilis''. Cell wall-binding modules of autolysins are used for non-covalent interaction with the ''B. subtilis'' cell wall, whereas a sortase-based system functions via covalent attachment to the cell wall. In both cases, a linker between cell wall anchor and peptide allows flexibility of the fusion protein and increases its range. Peptides binding to different receptors on the surface of pathogens mediate specific binding. |
| | | | |
| | <html> | | <html> |
| Line 438: |
Line 553: |
| | Constructs for non-covalent and covalent surface display were cloned similar to those described by Chen ''et. al.'', 2008 [1], and Liew ''et. al.'', 2012 [6], respectively. | | Constructs for non-covalent and covalent surface display were cloned similar to those described by Chen ''et. al.'', 2008 [1], and Liew ''et. al.'', 2012 [6], respectively. |
| | | | |
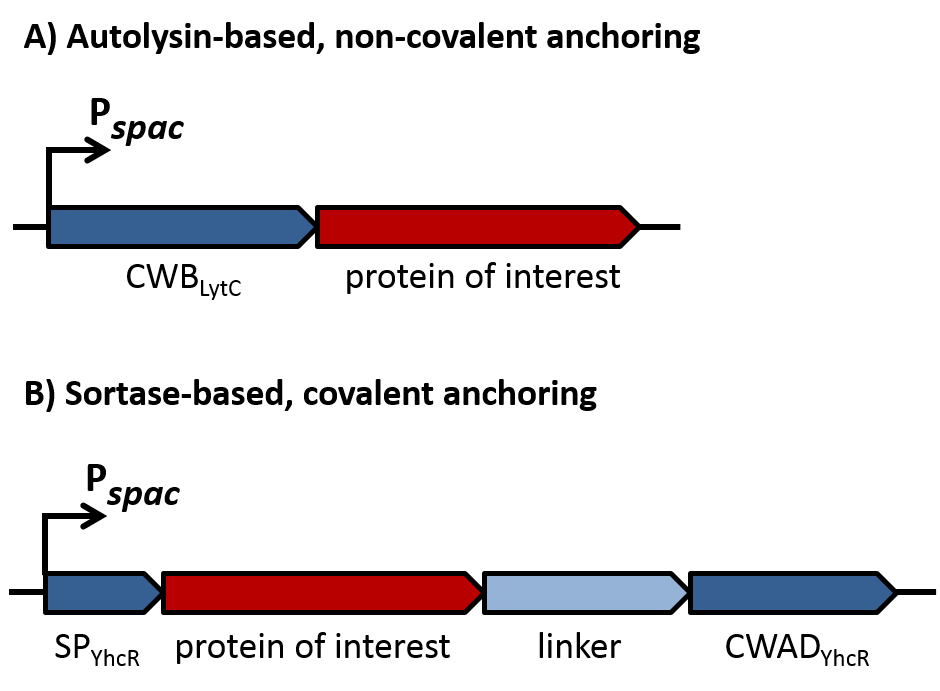
| - | As depicted in figure 1A, cell-wall binding domains (CWB) of LytC or LytE ([http://parts.igem.org/wiki/index.php?title=Part:BBa_K1351006 BBa_K1351006] or [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1351007 BBa_K1351007]) were translationally fused to a protein of interest for non-covalent anchoring of the latter to the cell wall. These CWB domains also included the native signal peptide (SP) of the respective proteins to direct export via the Sec pathway. For covalent anchoring, signal peptide ([http://parts.igem.org/Part:BBa_K1351008 BBa_K1351008]) and cell wall anchoring domain (CWAD, [http://parts.igem.org/Part:BBa_K1351010 BBa_K1351010]) of the sortase substrate YhcR and a 57 amino acid linker ([http://parts.igem.org/Part:BBa_K1351009 BBa_K1351009]) were assembled with the protein of interest as depicted in figure 1B to generate a YhcR-like fusion. | + | As depicted in figure 1A, cell-wall binding domains (CWB) of LytC or LytE ([http://parts.igem.org/wiki/index.php?title=Part:BBa_K1351006 BBa_K1351006] or [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1351007 BBa_K1351007]) should be translationally fused to a protein of interest for non-covalent anchoring of the latter to the cell wall. These CWB domains also included the native signal peptide (SP) of the respective proteins to direct export via the Sec pathway. For covalent anchoring, signal peptide ([http://parts.igem.org/Part:BBa_K1351008 BBa_K1351008]) and cell wall anchoring domain (CWAD, [http://parts.igem.org/Part:BBa_K1351010 BBa_K1351010]) of the sortase substrate YhcR and a 57 amino acid linker ([http://parts.igem.org/Part:BBa_K1351009 BBa_K1351009]) were assembled with the protein of interest as depicted in figure 1B to generate a YhcR-like fusion. |
| | | | |
| | [[File:LMU14_adhesion_fig1.png | 600px]] | | [[File:LMU14_adhesion_fig1.png | 600px]] |
| | | | |
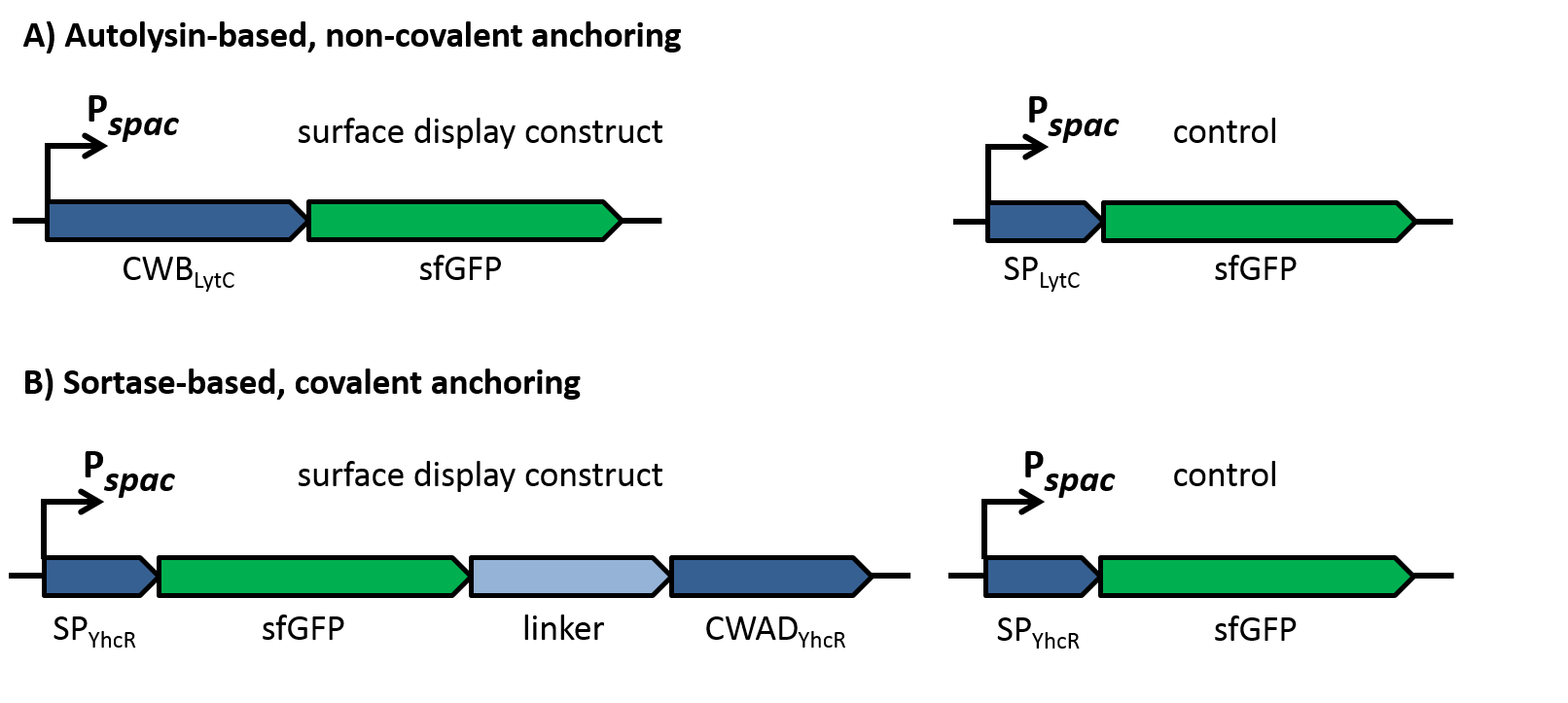
| - | To evaluate the functionality of the constructs, the latter were cloned into the replicative ''B. subtilis'' vector pBS0K under the control of the inducible promoter P<sub>''spac''</sub>. Two different proteins were used to test for successful display: Firstly, a superfolder GFP variant (sfGFP) was used for fluorescence microscopy to check if the constructs locate to the cell wall as shown in figure 2. This variant of GFP had been shown previously to fold in the periplasm of gram-negative bacteria [10] and could thus be expected to be fuctional also in the extracellular environment, in contrast to normal GFP that does not fold under reducing conditions. As controls for the sfGFP constructs, only the signal peptides without the respective cell wall binding or anchoring domains were fused to sfGFP, which should lead to secretion of the fusion into the growth medium. | + | To evaluate the functionality of the constructs, the latter should be cloned into the replicative ''B. subtilis'' vector pBS0K under the control of the inducible promoter P<sub>''spac''</sub>. Two different proteins should be for successful display: First, a superfolder GFP variant (sfGFP) was used for fluorescence microscopy to check if the constructs locate to the cell wall as shown in figure 2. This variant of GFP had been shown previously to fold in the periplasm of gram-negative bacteria [10] and could thus be expected to be fuctional also in the extracellular environment, in contrast to normal GFP that does not fold under reducing conditions. As controls for the sfGFP constructs, only the signal peptides without the respective cell wall binding or anchoring domains were fused to sfGFP, which should lead to secretion of the fusion into the growth medium. |
| | | | |
| | [[File:LMU14_adhesion_fig2.png | 960px]] | | [[File:LMU14_adhesion_fig2.png | 960px]] |
| | | | |
| - | Secondly, fusions to alkaline phosphatase (PhoA) were used to detect surface accessibility of the presented protein, as PhoA is only active when exported out of the cytoplasm [11]. Agar plates containing the PhoA substrate XP (5-bromo-4-chloro-3-indolyl-phosphate) indicate enzyme activity by turning blue where the substrate is hydrolyzed and further oxidized to an indigo dye [12]. PhoA was also fused to intracellular GFP as a negative control and to an extracellular domain of LiaI as a positive control. The PhoA-deficient ''B. subtilis'' strain MH3402 was used for expression of all PhoA constructs. | + | Secondly, fusions to alkaline phosphatase (PhoA) should be used to detect surface accessibility of the presented protein, as PhoA is only active when exported out of the cytoplasm [11]. Agar plates containing the PhoA substrate XP (5-bromo-4-chloro-3-indolyl-phosphate) indicate enzyme activity by turning blue where the substrate is hydrolyzed and further oxidized to an indigo dye [12]. PhoA was also fused to intracellular GFP as a negative control and to an extracellular domain of LiaI as a positive control. The PhoA-deficient ''B. subtilis'' strain MH3402 will be used for expression of all PhoA constructs. |
| | + | |
| | + | Unfortunately, the status of these constructs is still "cloning". |
| | | | |
| | ===Pathogen binding=== | | ===Pathogen binding=== |
| Line 453: |
Line 570: |
| | [[File:LMU14_adhesion_fig3.png | 600px]] | | [[File:LMU14_adhesion_fig3.png | 600px]] |
| | | | |
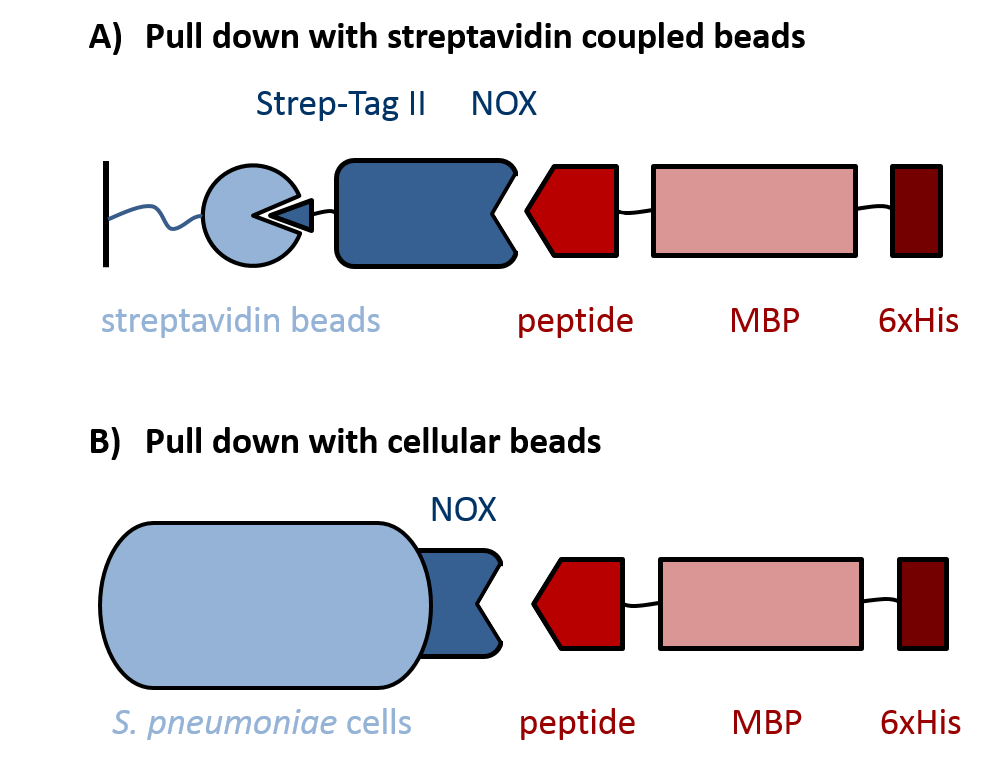
| - | Evaluation of peptides binding to ''S. pneumoniae'' was done by a biochemical pull-down assay with their binding partner NADH oxidase (NOX). As shown in figure 3, NOX was bound to streptavidin-coupled beads via an N-terminal fused Strep-Tag II, for which the pASK-IBA expression vector system was used. ''S. pneumoniae''-specific peptides C4P, CSP and L5P ([http://parts.igem.org/Part:BBa_K1351001 BBa_K1351001], [http://parts.igem.org/Part:BBa_K1351002 BBa_K1351002] and [http://parts.igem.org/Part:BBa_K1351003 BBa_K1351003], respectively) were fused C-terminally to a maltose-binding protein (MBP) and a 6xHis-tag. The MBP prevents degradation of the short peptides by proteases and was used for detection of the peptide fusions on an SDS gel. To increase specificity of the detection, the 6xHis-tag was used for Western Blot analysis of the pulled down proteins. Additionally, ''S. pneumoniae'' itself was used as cellular beads for a pull down of the same peptide-MBP-His constructs, again followed by analysis with SDS gel and Western Blot. | + | Evaluation of peptides binding to ''S. pneumoniae'' was done by a biochemical pull-down assay with their binding partner NADH oxidase (NOX). As shown in figure 3, NOX was bound to streptavidin-coupled beads via an N-terminal fused Strep-Tag II, for which the pASK-IBA expression vector system was used. ''S. pneumoniae''-specific peptides C4P, CSP and L5P ([http://parts.igem.org/Part:BBa_K1351001 BBa_K1351001], [http://parts.igem.org/Part:BBa_K1351002 BBa_K1351002] and [http://parts.igem.org/Part:BBa_K1351003 BBa_K1351003], respectively) were fused C-terminally to a maltose-binding protein (MBP) and a 6xHis-tag into the pKLD66 vector. This was achieved by digestion of the BioBricks and pKLD66 with NotI and HindIII and ligation of the products. The MBP prevents degradation of the short peptides by proteases and was used for detection of the peptide fusions on an SDS gel. To increase specificity of the detection, the 6xHis-tag was used for Western Blot analysis of the pulled down proteins. Additionally, ''S. pneumoniae'' itself was used as cellular beads for a pull down of the same peptide-MBP-His constructs, again followed by analysis with SDS gel and Western Blot. |
| | | | |
| | ===Sources=== | | ===Sources=== |
| Line 491: |
Line 608: |
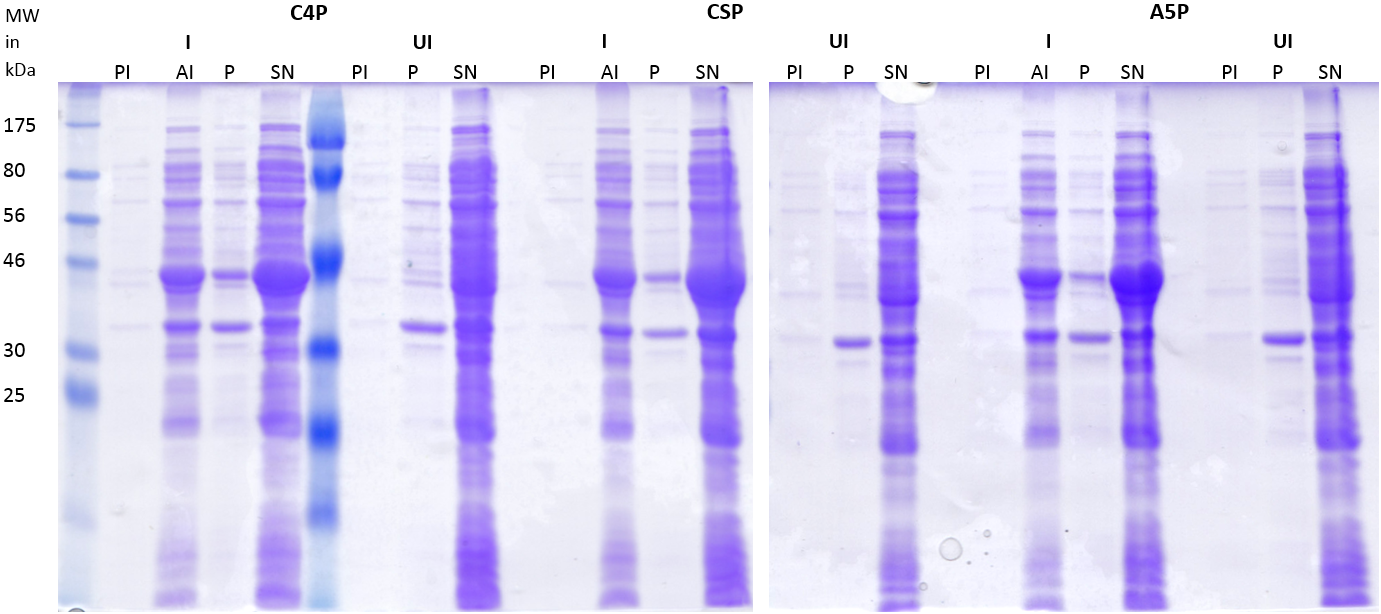
| | [[File:LMU14_Adhesion_Results_Figure3.PNG|thumb|center|700px|'''Figure 3:SDS-PAGE of the lysates and the fractions of the incubated S. pneumoniae cells.''' E. coli cells carrying the plasmids coding for C4P, CSP and A5P were lysed. ''S. pneumoniae'' cells were resuspended in the lysate and incubated for 30 minutes, followed by pelleting to separate the supernatant (SP) from the cell pellet (P). 3 μl of sample were applied per lane.]] | | [[File:LMU14_Adhesion_Results_Figure3.PNG|thumb|center|700px|'''Figure 3:SDS-PAGE of the lysates and the fractions of the incubated S. pneumoniae cells.''' E. coli cells carrying the plasmids coding for C4P, CSP and A5P were lysed. ''S. pneumoniae'' cells were resuspended in the lysate and incubated for 30 minutes, followed by pelleting to separate the supernatant (SP) from the cell pellet (P). 3 μl of sample were applied per lane.]] |
| | | | |
| - | The gel of the SDS-PAGE (Fig. 3) shows strong bands for the lysates at 40kDa, indicating a high amount of the His-MBP tagged peptides. The empty vector shows a relatively low expression of His-MBP. The pellet fractions of the ''S. pneumoniae'' cells incubated with the tagged C4P and CSP peptides show light bands at the same height indicating adhesion of the peptides to the cells (Fig. 4, white arrows). Although the cells incubated in buffer alone also show a band at the same height indicating a native protein of that size ((Fig. 3, black arrows) the amount of protein is clearly less. To determine whether the band represents the His-MBP tagged peptides, a western blot was performed (Figure 4). | + | The gel of the SDS-PAGE (Fig. 3) shows strong bands for the lysates at 40kDa, indicating a high amount of the His-MBP tagged peptides. The empty vector shows a relatively low expression of His-MBP. The pellet fractions of the ''S. pneumoniae'' cells incubated with the tagged C4P and CSP peptides show light bands at the same height indicating adhesion of the peptides to the cells (Fig. 4, white arrows). Although the cells incubated in buffer alone also show a band at the same height indicating a native protein of that size (Fig. 3, black arrows) the amount of protein is clearly less. To determine whether the band represents the His-MBP tagged peptides, a western blot was performed (Figure 4). |
| | | | |
| | [[File:LMU14_Adhesion_Results_Figure5.PNG|thumb|center|800px|'''Figure 4: Western-Blots of the lysates and the fractions of the incubated ''S. pneumoniae'' cells.''' Duplicates of the gel from Figure 4 (A) and another gel loaded with 10 μl/ lane (B) were processed by western blot with Anti-His antibodies.]] | | [[File:LMU14_Adhesion_Results_Figure5.PNG|thumb|center|800px|'''Figure 4: Western-Blots of the lysates and the fractions of the incubated ''S. pneumoniae'' cells.''' Duplicates of the gel from Figure 4 (A) and another gel loaded with 10 μl/ lane (B) were processed by western blot with Anti-His antibodies.]] |
| Line 531: |
Line 648: |
| | | | |
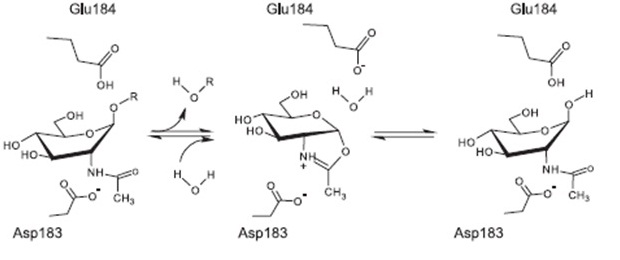
| | Subtilin is a small antimicrobial peptide (AMP) which belongs to the class of lanthionine-containing antibiotics (lantibiotics). These are heat-stable, ribosomally synthesized molecules with a molecular weight below 4 kDa. Their main characteristic is the high proportion of unusual amino acids, synthesized by post-translational side-chain modifications of precursor peptides. Most prominent examples are the polycyclic thioether amino acids, lanthionine and β–methyllanthionine and a number of dehydrated amino acids such as dehydroalanine (Dha) and dehydrobutyrine (Dhb). | | Subtilin is a small antimicrobial peptide (AMP) which belongs to the class of lanthionine-containing antibiotics (lantibiotics). These are heat-stable, ribosomally synthesized molecules with a molecular weight below 4 kDa. Their main characteristic is the high proportion of unusual amino acids, synthesized by post-translational side-chain modifications of precursor peptides. Most prominent examples are the polycyclic thioether amino acids, lanthionine and β–methyllanthionine and a number of dehydrated amino acids such as dehydroalanine (Dha) and dehydrobutyrine (Dhb). |
| - | Lantibiotics are promising candidates for future antimicrobials, as they inhibit the growth of many clinically relevant pathogens comprising even multidrug-resistant bacteria. Moreover, lantibiotics have only low tendency to generate resistance. Both features make them highly attractive for medical applications. | + | Lantibiotics are promising candidates for future antimicrobials, as they inhibit the growth of many clinically relevant pathogens comprising even multidrug-resistant bacteria. Moreover, lantibiotics have only low tendency to generate resistance. Both features make them highly attractive for medical applications. [http://www.unboundmedicine.com/medline/citation/24210177/Lantibiotics:_promising_candidates_for_future_applications_in_health_care_[1]] [http://www.ncbi.nlm.nih.gov/pubmed/16205711[2]] |
| | | | |
| | [[File:LMU14 killing subtilin structure.png|thumb|800px|center|Fig. 1. Schematic representation of the structure of the lantibiotic subtilin. Dha represents a dehydroalanine residue, Dhb represents a dehydrobutirine residue, and Abu represents a dehydrobutirine residue that has formed a thioether β-methyl-lanthionine bridge with a cysteine residue. | | [[File:LMU14 killing subtilin structure.png|thumb|800px|center|Fig. 1. Schematic representation of the structure of the lantibiotic subtilin. Dha represents a dehydroalanine residue, Dhb represents a dehydrobutirine residue, and Abu represents a dehydrobutirine residue that has formed a thioether β-methyl-lanthionine bridge with a cysteine residue. |
| - | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Quorum+sensing+control+of+lantibiotic+production%3B+nisin+and+subtilin+autoregulate+their+own+biosynthesis]]] | + | [http://www.ncbi.nlm.nih.gov/pubmed/?term=Quorum+sensing+control+of+lantibiotic+production%3B+nisin+and+subtilin+autoregulate+their+own+biosynthesis[3] ]]] |
| | | | |
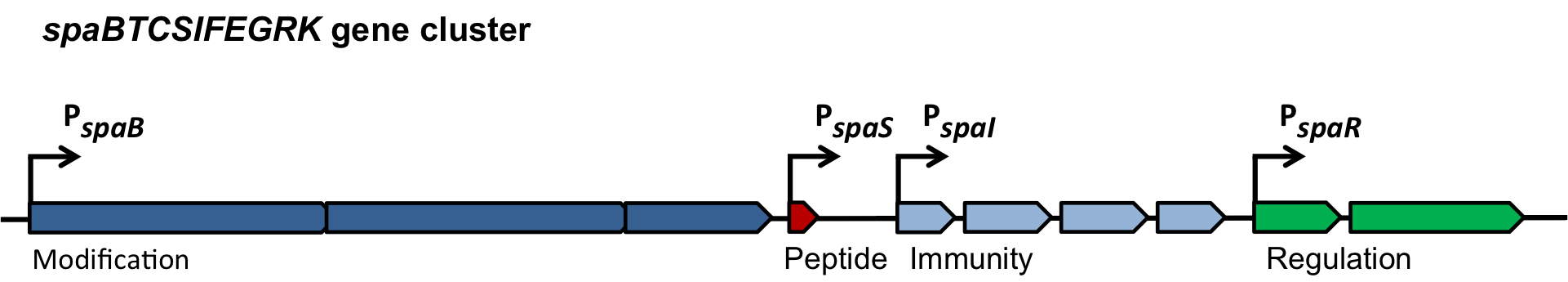
| | ===The ''spaBTCSIFEGRK'' gene cluster=== | | ===The ''spaBTCSIFEGRK'' gene cluster=== |
| | | | |
| - | [[File:LMU14 killing subtilin genecluster.png|thumb|800px|center|Fig. 2. Organisation of the subtilin biosynthetic gene cluster. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Quorum+sensing+control+of+lantibiotic+production%3B+nisin+and+subtilin+autoregulate+their+own+biosynthesis]]] | + | [[File:LMU14 killing subtilin genecluster.png|thumb|800px|center|Fig. 2. Organisation of the subtilin biosynthetic gene cluster. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Quorum+sensing+control+of+lantibiotic+production%3B+nisin+and+subtilin+autoregulate+their+own+biosynthesis [3] ]]] |
| | | | |
| | As true for many other lantibiotics, the genes for the subtilin biosynthesis are clustered. For subtilin, there is even a reasonable arrangement within the cluster. The genes ''spaBTC'' play a role for the posttranslational modifications and the transport of subtilin. They are regulated by their own promotor P<sub>''spaB''</sub> and polycistronically transcribed. ''spaS'' encodes the propeptide, which is derived from a short monocistronic mRNA. The next transcription entity, ''spaIFEG'', is coding for the immunity, whereas ''spaRK'' encodes a two-component-system that plays an important role for the regulation. What is missing in comparison to other lantibiotic gene clusters is a specific protease (encoded by ''lanP'') that cleaves off the leader sequence. In ''Bacillus subtilis'', this task is accomplished by unspecific extracellular proteases. | | As true for many other lantibiotics, the genes for the subtilin biosynthesis are clustered. For subtilin, there is even a reasonable arrangement within the cluster. The genes ''spaBTC'' play a role for the posttranslational modifications and the transport of subtilin. They are regulated by their own promotor P<sub>''spaB''</sub> and polycistronically transcribed. ''spaS'' encodes the propeptide, which is derived from a short monocistronic mRNA. The next transcription entity, ''spaIFEG'', is coding for the immunity, whereas ''spaRK'' encodes a two-component-system that plays an important role for the regulation. What is missing in comparison to other lantibiotic gene clusters is a specific protease (encoded by ''lanP'') that cleaves off the leader sequence. In ''Bacillus subtilis'', this task is accomplished by unspecific extracellular proteases. |
| Line 544: |
Line 661: |
| | | | |
| | | | |
| - | <i>Table 1: The role of the different proteins of the spa-locus</i> | + | <i>Table 1: The role of the different proteins of the ''spa''-locus</i> |
| | {| class="wikitable" | | {| class="wikitable" |
| | |- | | |- |
| Line 587: |
Line 704: |
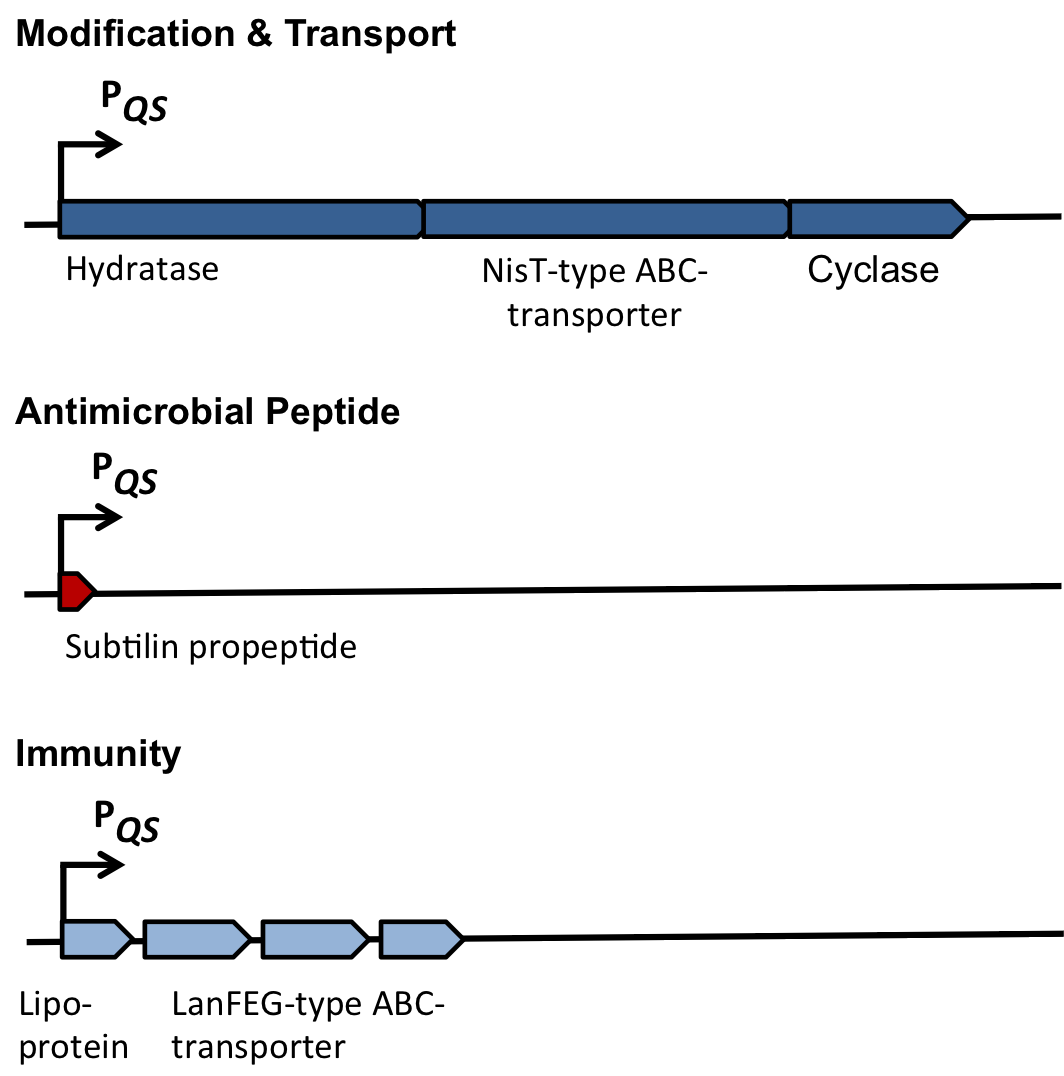
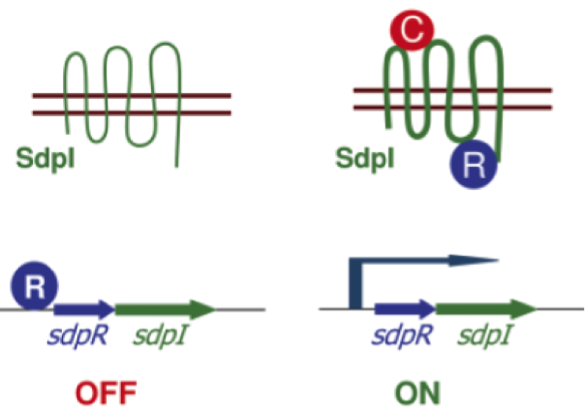
| | ===Self-protection from subtilin=== | | ===Self-protection from subtilin=== |
| | | | |
| - | Of course, the producer needs to be immune against the antimicrobial agent it produces. ''B. subtilis'' ATCC6633 protects itself against subtilin by two mechanisms that both act independently and confer some level of resistance, however full resistance is only achieved when both mechanisms are active. SpaI is a membrane-anchored lipoprotein. It is suggested that it binds the antimicrobial peptide and by this keeps it away from the membrane. SpaFEG is a LanFEG-like ABC-transporter that works by exporting subtilin from the cytoplasmic membrane to the culture supernatant. | + | Of course, the producer needs to be immune against the antimicrobial agent it produces. ''B. subtilis'' ATCC6633 protects itself against subtilin by two mechanisms that both act independently and confer some level of resistance, however full resistance is only achieved when both mechanisms are active. SpaI is a membrane-anchored lipoprotein. It is suggested that it binds the antimicrobial peptide and by this keeps it away from the membrane. SpaFEG is a LanFEG-like ABC-transporter that works by exporting subtilin from the cytoplasmic membrane to the culture supernatant. [http://www.ncbi.nlm.nih.gov/pubmed/15659659 [4]] [http://www.ncbi.nlm.nih.gov/pubmed/?term=Alkhatib%2C+Z.%3B+Abts%2C+A.%3B+Mavaro%2C+A.%3B+Schmitt%2C+L.%3B+Smits%2C+S.+H.%3A+Lantibiotics%3A+how+do+producers+become+self-protected%3F+Journal+of+biotechnology+2012%2C+159%2C+145-54. [5]] |
| | | | |
| | | | |
| | ===Dual regulation of subtilin biosynthesis and immunity=== | | ===Dual regulation of subtilin biosynthesis and immunity=== |
| | | | |
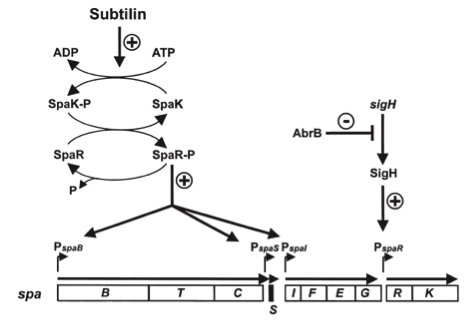
| - | [[File:LMU14 killing subtilin regulation.png|thumb|400px|right|Fig. 3. Dual control of subtilin biosynthesis and immunity. [http://www.ncbi.nlm.nih.gov/pubmed/11972779]]] | + | [[File:LMU14 killing subtilin regulation.png|thumb|400px|right|Fig. 3. Dual control of subtilin biosynthesis and immunity. [http://www.ncbi.nlm.nih.gov/pubmed/11972779 [6] ]]] |
| | | | |
| | Subtilin biosynthesis and immunity in ''B. subtilis'' ATCC6633 are subject to a dual control mechanism. | | Subtilin biosynthesis and immunity in ''B. subtilis'' ATCC6633 are subject to a dual control mechanism. |
| Line 599: |
Line 716: |
| | ===Mode of action=== | | ===Mode of action=== |
| | | | |
| - | Lantibiotics are active against a wide range of Gram-positive bacteria including multidrug-resistant pathogens. Some lantibiotics have a dual mode of action, subtilin is proposed to be among them. Subtilin forms a complex with the cell wall precursor lipid II, whereby the pyrophosphate moiety may play a key role for target recognition. Thereby, the cell wall biosynthesis is inhibited. Furthermore, the complexes aggregate and form a pore in the bacterial membrane by using lipid II as a docking molecule. This leads to a depolarisation of the membrane potential, the efflux of cytoplasma and in turn to rapid cell death. Just like nisin, which is highly similar, subtilin is still active in the nanomolar range and thus serves excellently as alternative killing strategy against multidrug-resistant pathogens. | + | Lantibiotics are active against a wide range of Gram-positive bacteria including multidrug-resistant pathogens. Some lantibiotics have a dual mode of action, subtilin is proposed to be among them. Subtilin forms a complex with the cell wall precursor lipid II, whereby the pyrophosphate moiety may play a key role for target recognition. Thereby, the cell wall biosynthesis is inhibited. Furthermore, the complexes aggregate and form a pore in the bacterial membrane by using lipid II as a docking molecule. This leads to a depolarisation of the membrane potential, the efflux of cytoplasma and in turn to rapid cell death. Just like nisin, which is highly similar, subtilin is still active in the nanomolar range and thus serves excellently as alternative killing strategy against multidrug-resistant pathogens. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Molecular+Mechanism+of+Target+parisot [7]] [http://www.researchgate.net/publication/7875075_Mode_of_action_of_lipid_II-targeting_lantibiotics [8]] |
| | | | |
| - | [[File:LMU14 killing subtilin modeofaction.png|frame|150px|center|Fig. 4. Proposed mechanism of pyrophosphate-mediated target engagement by the lanthionine antibiotic subtilin. Membrane breaches occur in a lipid II-mediated fashion and cell wall synthesis is impeded through engagement of pyrophosphate-containing intermediates into complexes. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Molecular+Mechanism+of+Target+parisot]]] | + | |
| | + | [[File:LMU14 killing subtilin modeofaction.png|frame|150px|center|Fig. 4. Proposed mechanism of pyrophosphate-mediated target engagement by the lanthionine antibiotic subtilin. Membrane breaches occur in a lipid II-mediated fashion and cell wall synthesis is impeded through engagement of pyrophosphate-containing intermediates into complexes. [http://www.ncbi.nlm.nih.gov/pubmed/?term=Molecular+Mechanism+of+Target+parisot [7] ]]] |
| | | | |
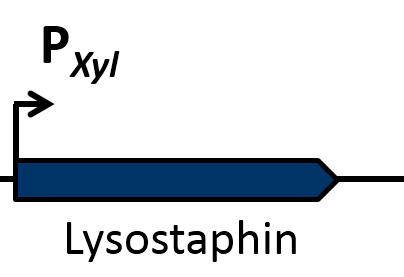
| | == Lysostaphin == | | == Lysostaphin == |
| Line 632: |
Line 750: |
| | === Sources: === | | === Sources: === |
| | <small> | | <small> |
| | + | |
| | + | [1] Dischinger, J.; Basi Chipalu, S.; Bierbaum, G.: Lantibiotics: promising candidates for future applications in health care. International journal of medical microbiology : IJMM 2014, 304, 51-62. |
| | + | |
| | + | [2] Cotter, P. D.; Hill, C.; Ross, R. P.: Bacteriocins: developing innate immunity for food. Nature reviews. Microbiology 2005, 3, 777-88. |
| | + | |
| | + | [3] Kleerebezem, M.: Quorum sensing control of lantibiotic production; nisin and subtilin autoregulate their own biosynthesis. Peptides 2004, 25, 1405-14. |
| | + | |
| | + | [4] Stein, T.; Heinzmann, S.; Dusterhus, S.; Borchert, S.; Entian, K. D.: Expression and functional analysis of the subtilin immunity genes spaIFEG in the subtilin-sensitive host Bacillus subtilis MO1099. Journal of bacteriology 2005, 187, 822-8. |
| | + | |
| | + | [5] Alkhatib, Z.; Abts, A.; Mavaro, A.; Schmitt, L.; Smits, S. H.: Lantibiotics: how do producers become self-protected? Journal of biotechnology 2012, 159, 145-54. |
| | + | |
| | + | [6] Stein, T.; Borchert, S.; Kiesau, P.; Heinzmann, S.; Kloss, S.; Klein, C.; Helfrich, M.; Entian, K. D.: Dual control of subtilin biosynthesis and immunity in Bacillus subtilis. Molecular microbiology 2002, 44, 403-16. |
| | + | |
| | + | [7] Parisot, J.; Carey, S.; Breukink, E.; Chan, W. C.; Narbad, A.; Bonev, B.: Molecular mechanism of target recognition by subtilin, a class I lanthionine antibiotic. Antimicrobial agents and chemotherapy 2008, 52, 612-8. |
| | + | |
| | + | [8] Bauer, R.; Dicks, L. M.: Mode of action of lipid II-targeting lantibiotics. International journal of food microbiology 2005, 101, 201-16. |
| | + | |
| | [B1] M.C.F. Bastos, H. Ceotto1, M.L.V. Coelho and J.S. Nascimento: Staphylococcal Antimicrobial Peptides: Relevant Properties and Potential Biotechnological Applications in Current Pharmaceutical Biotechnology (2009) S. 38-61 | | [B1] M.C.F. Bastos, H. Ceotto1, M.L.V. Coelho and J.S. Nascimento: Staphylococcal Antimicrobial Peptides: Relevant Properties and Potential Biotechnological Applications in Current Pharmaceutical Biotechnology (2009) S. 38-61 |
| | | | |
| Line 718: |
Line 853: |
| | | | |
| | First of all, a growth curve for ''S. pneumoniae'' was recorded to learn about the growth behavior of this organism. It was measured in the plate reader in 100 µl THB broth at 37 °C without shaking and measurement was taken every 10 min. The mean value of 10 measurements were determined and plotted. | | First of all, a growth curve for ''S. pneumoniae'' was recorded to learn about the growth behavior of this organism. It was measured in the plate reader in 100 µl THB broth at 37 °C without shaking and measurement was taken every 10 min. The mean value of 10 measurements were determined and plotted. |
| - | [[File:LMU14 killing subtilin growthcurve pneumo.png|thumb|500px|right|Fig. 1. Growth curve of ''S. pneumoniae'' measured in the plate reader in 100 µl THB broth.]] | + | [[File:LMU14 killing subtilin growthcurve pneumo.png|thumb|600px|center|Fig. 1. Growth curve of ''S. pneumoniae'' measured in the plate reader in 100 µl THB broth.]] |
| - | [[File:LMU14 killing subtilin Pneumo-halo.png|thumb|500px|right|Fig. 1. The spot-on-lawn assay shows a clear halo around the ''B. subtilis'' ATCC6633 spot. No clearing zone is visible around ''B. subtilis'' W168.]] | + | [[File:LMU14 killing subtilin Pneumo-halo.png|thumb|400px|center|Fig. 2. The spot-on-lawn assay shows a clear halo around the ''B. subtilis'' ATCC6633 spot. No clearing zone is visible around ''B. subtilis'' W168.]] |
| | As expected, ''S. pneumoniae'' shows rapid growth during its exponential growth phase. It reaches its maximal OD<sub>600</sub> just under 5 hours. In contrast to other bacteria, for ''S. pneumoniae'', there is no plateau during stationary phase, as autolysis of the cells starts and the OD<sub>600</sub> decreases again. | | As expected, ''S. pneumoniae'' shows rapid growth during its exponential growth phase. It reaches its maximal OD<sub>600</sub> just under 5 hours. In contrast to other bacteria, for ''S. pneumoniae'', there is no plateau during stationary phase, as autolysis of the cells starts and the OD<sub>600</sub> decreases again. |
| | | | |
| | | | |
| - | As ''S. pneumoniae'' is a target of the BaKillus killing strategies, a spot-on-lawn-assay should reveal the inhibitory impact of subtilin. Therefore, spots of the ''B. subtilis'' W168 wild type and the subtilin-producer ''B. subtilis'' ATCC6633, respectively, were spotted on a ''S. pneumoniae'' lawn. As depicted in figure XY, the subtilin producer significantly inhibits the growth of ''S. pneumoniae''. In contrast, ''B. subtilis'' W168 wild type does not show inhibitory impact. The experiment was performed three times and the mean value and the standard deviation are illustrated in the graph below. | + | As ''S. pneumoniae'' is a target of the BaKillus killing strategies, a spot-on-lawn-assay should reveal the inhibitory impact of subtilin. Therefore, spots of the ''B. subtilis'' W168 wild type and the subtilin-producer ''B. subtilis'' ATCC6633, respectively, were spotted on a ''S. pneumoniae'' lawn. As depicted in figure 2 and 3, the subtilin producer significantly inhibits the growth of ''S. pneumoniae''. In contrast, ''B. subtilis'' W168 wild type does not show inhibitory impact. The experiment was performed three times and the mean value and the standard deviation are illustrated in the graph below. |
| | | | |
| - | [[File:LMU14 killing subtilin Pneumo-inhibition diagramm.png|thumb|500px|center|Fig. 1. Bar graph .]] | + | [[File:LMU14 killing subtilin Pneumo-inhibition diagramm.png|thumb|700px|center|Fig. 3. Analysis of subtilin impact on ''S. pneumoniae''.]] |
| | | | |
| | ===Subtilin impact on ''Staphylococcus aureus''=== | | ===Subtilin impact on ''Staphylococcus aureus''=== |
| | | | |
| - | Moreover, the impact of subtilin on ''Staphylococcus aureus'' was tested in collaboration with iGEM Team Groningen, as they have access to S2 laboratories, in which ''S. aureus'' and even multidrug-resitant species are handled. A member of Team Groningen kindly performed spot-on-lawn assays with ''B. subtilis'' W168 wild type and the subtilin-producer ''B. subtilis'' ATCC6633 on different ''S. aureus'' species. The tested strains were CAL, NWZ (both MRSA, obtained at the UMCG hospital) and CECT240 (''Staphylococcus aureus'' subsp. ''aureus'', Rosenbach 1884). Strains were plated out in soft agar and spots from overnight cultures of ''Bacillus subtilis'' grown in either complex medium (LB) or defined medium (SMM) were added on top. [[File:LMU14 killing subtilin MRSA.png|thumb|700px|center|Fig. 1. Spot-on-lawn assay with the subtilin producer and on three different ''S. aureus'' lawns]] | + | Moreover, the impact of subtilin on ''Staphylococcus aureus'' was tested in collaboration with iGEM Team Groningen, as they have access to S2 laboratories, in which ''S. aureus'' and even multidrug-resitant species are handled. A member of Team Groningen kindly performed spot-on-lawn assays with ''B. subtilis'' W168 wild type and the subtilin-producer ''B. subtilis'' ATCC6633 on different ''S. aureus'' species. The tested strains were CAL, NWZ (both MRSA, obtained at the UMCG hospital) and CECT240 (''Staphylococcus aureus'' subsp. ''aureus'', Rosenbach 1884). Strains were plated out in soft agar and spots from overnight cultures of ''Bacillus subtilis'' grown in either complex medium (LB) or defined medium (SMM) were added on top. [[File:LMU14 killing subtilin MRSA.png|thumb|700px|center|Fig. 4. Spot-on-lawn assay with the subtilin producer and on three different ''S. aureus'' lawns]] |
| | After incubation for 24 h at 37 °C, halos could be detected for all spots of the subtilin producer. The exact results are shown in the chart below. However, the spots of the ''B. subtilis'' W168 culture did not inhibit the growth of any ''S. aureus'' strains (no pictures provided). | | After incubation for 24 h at 37 °C, halos could be detected for all spots of the subtilin producer. The exact results are shown in the chart below. However, the spots of the ''B. subtilis'' W168 culture did not inhibit the growth of any ''S. aureus'' strains (no pictures provided). |
| | | | |
| - | [[File:LMU14 killing MRSA diagramm.png|thumb|500px|center|Fig. 1..]] | + | [[File:LMU14 killing MRSA diagramm.png|thumb|500px|center|Fig. 5 Bar chart of the ''S. aureus'' spot-on-lawn assay.]] |
| | | | |
| | ===Heterologous expression=== | | ===Heterologous expression=== |
| Line 749: |
Line 884: |
| | <b>Antimicrobial peptide</b> | | <b>Antimicrobial peptide</b> |
| | | | |
| - | [http://parts.igem.org/Part:BBa_K1351012 BBa K1351012] and [http://parts.igem.org/Part:BBa_K1351013 BBa K1351013] are coding for SpaS, the subtilin pro-peptide. Their main difference are their overhangs, one being in RFC25, which allows the tagging or translational fusion of [http://parts.igem.org/Part:BBa_K1351012 BBa K1351012], whereas the other can be used according to the RFC10 BioBrick standard. Unfortunately none of them could be evaluated so far, as the development of the immunity BioBrick as well as of the modification and export BioBrick were required beforehand. | + | [http://parts.igem.org/Part:BBa_K1351012 BBa K1351012] and [http://parts.igem.org/Part:BBa_K1351013 BBa K1351013] are coding for SpaS, the subtilin pro-peptide. Their main difference are their overhangs, one being in RFC25, which allows the tagging or translational fusion of [http://parts.igem.org/Part:BBa_K1351012 BBa K1351012], whereas the other can be used according to the RFC10 BioBrick standard. Unfortunately non of them could be evaluated so far, as the development of the immunity BioBrick as well as of the modification and export BioBrick were required beforehand. |
| | | | |
| | <b>Immunity</b> | | <b>Immunity</b> |
| | | | |
| | The immunity against subtilin ([http://parts.igem.org/Part:BBa_K1351014 BBa K1351014]), was - just like the spaS BioBricks - submitted to the iGEM parts registry. Self-protection is mediated by SpaIFEG, a membrane-anchored lipoprotein and an ABC-transporter. | | The immunity against subtilin ([http://parts.igem.org/Part:BBa_K1351014 BBa K1351014]), was - just like the spaS BioBricks - submitted to the iGEM parts registry. Self-protection is mediated by SpaIFEG, a membrane-anchored lipoprotein and an ABC-transporter. |
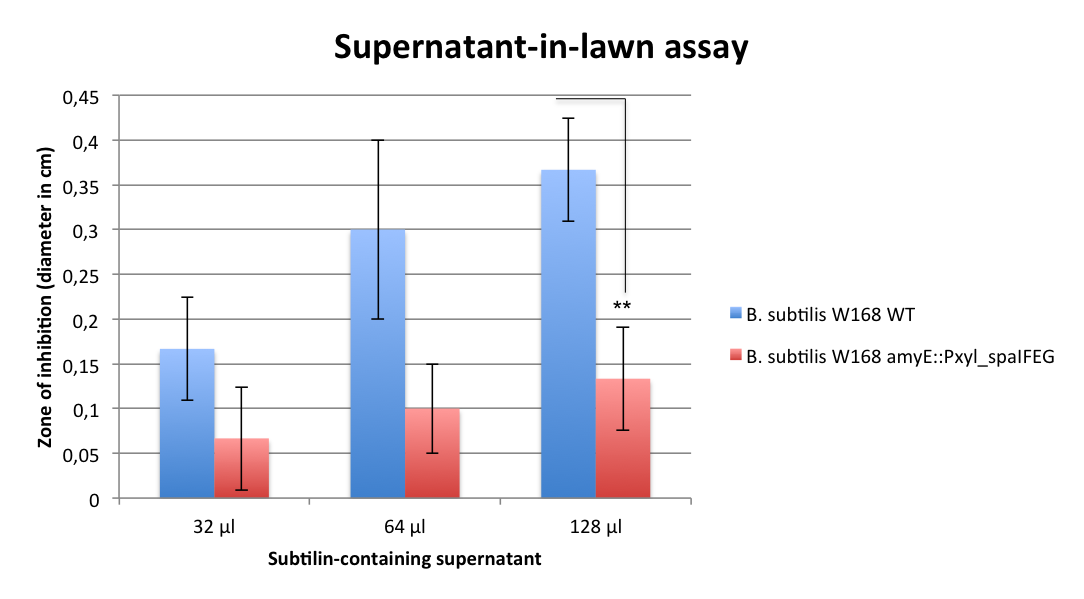
| - | [[File:LMU14 killing subtilin Immunity assay.png|thumb|600px|center|Fig. 1. Supernatant-in-lawn assay shows smaller halo diameters for ''B. subtilis'' with BioBRick-acquired immunity. ]] | + | [[File:LMU14 killing subtilin Immunity assay.png|thumb|700px|center|Fig. 6. Supernatant-in-lawn assay shows smaller halo diameters for ''B. subtilis'' with BioBRick-acquired immunity. ]] |
| - | The part was evaluated with a variation of an spot-on-lawn assay. In this case, holes were cut out of agar plates on which either ''B. subtilis'' W168 wild type or ''B. subtilis'' W168 ''amyE''::P<sub>''xyl''</sub>_''spaIFEG'' were plated. The holes were then filled with different amounts of sterile supernatant from overnight cultures of W168 or ATCC6633, respectively. The xylose-containing plates were incubated at 37 °C for 24 h and resulting halo diameters were measured. The supernatant derived from the negative control (W168) did not cause any growth inhibition to neither the wild type, nor to the strain who carries the immunity BioBrick (Figure XY). The subtilin-containing supernatant from ''B. subtilis'' ATCC6633 however, was able to inhibit the growth of both bacterial lawns. For the lowest amounts (32 µl), no significant difference between the halo diameters could be determined. The larger the amount of supernatant, and presumably, the higher the amount of subtilin in there, the bigger the zones of inhibition. For the largest volume (128 µl), a two-sided T-test could determine the p-value below 0.1 (p=0,007762603) and by this prove the functionality of this BioBrick. This is a major step for this project, as the immunity is the base for any strain which produces antimicrobial peptides. Now further steps can be taken and the remaining BioBricks can be evaluated. | + | The part was evaluated with a variation of an spot-on-lawn assay. In this case, holes were cut out of agar plates on which either ''B. subtilis'' W168 wild type or ''B. subtilis'' W168 ''amyE''::P<sub>''xyl''</sub>_''spaIFEG'' were plated. The holes were then filled with different amounts of sterile supernatant from overnight cultures of W168 or ATCC6633, respectively. The xylose-containing plates were incubated at 37 °C for 24 h and resulting halo diameters were measured. The supernatant derived from the negative control (W168) did not cause any growth inhibition to neither the wild type, nor to the strain who carries the immunity BioBrick (Figure 6). The subtilin-containing supernatant from ''B. subtilis'' ATCC6633 however, was able to inhibit the growth of both bacterial lawns. For the lowest amounts (32 µl), no significant difference between the halo diameters could be determined. The larger the amount of supernatant, and presumably, the higher the amount of subtilin in there, the bigger the zones of inhibition. For the largest volume (128 µl), a two-sided T-test could determine the p-value below 0.1 (p=0,007762603) and by this prove the functionality of this BioBrick. This is a major step for this project, as the immunity is the base for any strain which produces antimicrobial peptides. Now further steps can be taken and the remaining BioBricks can be evaluated. |
| | | | |
| | | | |
| | | | |
| - | [[File:LMU14 killing-subtilin supernatant-in-lawn.png|thumb|500px|center|Fig. 1..]] | + | [[File:Supernatant in-lawn.png|thumb|700px|center|Fig. 7 Evaluation of the ''spaIFEG'' BioBrick ]] |
| | | | |
| | ==Lysostaphin== | | ==Lysostaphin== |
| Line 789: |
Line 924: |
| | Thanks to the numerous killing strategies listed above, BaKillus has specifically dispatched the targeted pathogens, thus its job is almost done. However, to ensure the temporary mode of action and to avoid uncontrollable spread, we implemented the so called “suicide switch“ leading to time-delayed cell death of BaKillus. | | Thanks to the numerous killing strategies listed above, BaKillus has specifically dispatched the targeted pathogens, thus its job is almost done. However, to ensure the temporary mode of action and to avoid uncontrollable spread, we implemented the so called “suicide switch“ leading to time-delayed cell death of BaKillus. |
| | | | |
| - | During stationary phase of a Bacillus subtilis culture it has been observed that some cells start producing a toxin which causes cell lysis of sister cells whereas the toxin-producer cells stay unaffected. [1] This social and heterogenous behavior is called cannibalism and seems to be an even more promising basis for constructing the BaKillus “suicide switch“ since it has been shown that the toxin also kills a variety of Gram-positive bacteria, including MRSA-strains. [2] | + | During stationary phase of a ''Bacillus subtilis'' culture it has been observed that some cells start producing a toxin which causes cell lysis of sister cells whereas the toxin-producer cells stay unaffected. [http://www.sciencemag.org/content/301/5632/510.full [1]] This social and heterogenous behavior is called cannibalism and seems to be an even more promising basis for constructing the BaKillus “suicide switch“ since it has been shown that the toxin also kills a variety of Gram-positive bacteria, including MRSA-strains. [http://www.pnas.org/content/107/37/16286.full [2]] |
| | | | |
| | So how does the suicide switch work? | | So how does the suicide switch work? |
| Line 795: |
Line 930: |
| | | | |
| | | | |
| - | [1] Gonzalez-Pastor, J. E., et al. (2003). "Cannibalism by sporulating bacteria." Science 301(5632): 510-513. | + | [http://www.sciencemag.org/content/301/5632/510.full [1]] Gonzalez-Pastor, J. E., et al. (2003). "Cannibalism by sporulating bacteria." Science 301(5632): 510-513. |
| | | | |
| - | [2] Liu, W. T., et al. (2010). "Imaging mass spectrometry of intraspecies metabolic exchange revealed the cannibalistic factors of Bacillus subtilis." Proc Natl Acad Sci U S A | + | [http://www.pnas.org/content/107/37/16286.full [2]] Liu, W. T., et al. (2010). "Imaging mass spectrometry of intraspecies metabolic exchange revealed the cannibalistic factors of Bacillus subtilis." Proc Natl Acad Sci U S A |
| | | | |
| | <html> | | <html> |
| Line 807: |
Line 942: |
| | <article class="ac-small"> | | <article class="ac-small"> |
| | </html> | | </html> |
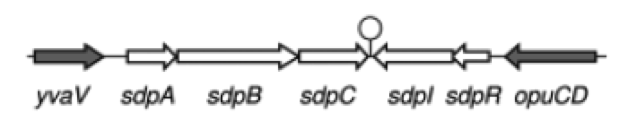
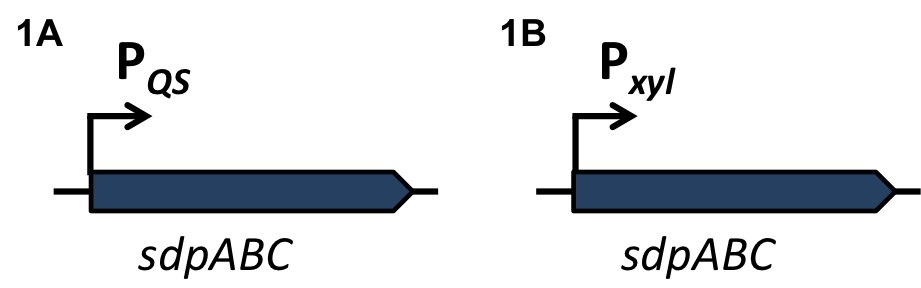
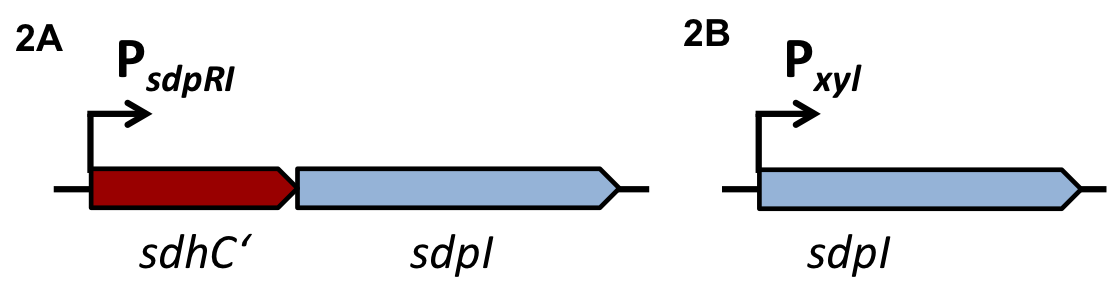
| - | '''The sdp-System of B. subtilis''' consists of two operons: The ''sdpABC'' operon, coding for the production and secretion of the cannibalism toxin SDP and the ''sdpRI'' operon responsible for the regulation and production of the immunity protein SdpI (Fig. 1). | + | '''The ''sdp''-System of ''B. subtilis''''' consists of two operons: The ''sdpABC'' operon, coding for the production and secretion of the cannibalism toxin SDP and the ''sdpRI'' operon responsible for the regulation and production of the immunity protein SdpI (Fig. 1). |
| | | | |
| - | [[File:LMU14 suicide background Fig.1.png|thumb|800px|center|Fig. 1. Gene organization for the sdpABC sdpRI operons. The hairpin symbolizes thetranscriptional terminators. [http://onlinelibrary.wiley.com/store/10.1111/j.1574-6976.2010.00253.x/asset/j.1574-6976.2010.00253.x.pdf;jsessionid=F0252E2CE3502A1AC14EF1CC930BE247.f04t01?v=1&t=i1e5c1ru&s=200f77df155555f934571349fb0543e7efe70ea4&systemMessage=Wiley+Online+Library+will+be+disrupted+on+the+18th+October+from+10%3A00+BST+%2805%3A00+EDT%29+for+essential+maintenance+for+approximately+two+hours+as+we+make+upgrades+to+improve+our+services+to+you[1]]] | + | [[File:LMU14 suicide background Fig.1.png|thumb|800px|center|Fig. 1. Gene organization for the ''sdpABC'' ''sdpRI'' operons. The hairpin symbolizes thetranscriptional terminators. [http://www.sciencemag.org/content/301/5632/510.full [1] ]]] |
| | | | |
| - | In vegetative cells, both operons are repressed by the unstable AbrB regulator. However, during early stages of sporulation AbrB itself is repressed by the master regulator of sporulation Spo0A, making ''sdpABC'' and ''sdpRI'' accessible for RNA polymerase. [http://onlinelibrary.wiley.com/store/10.1111/j.1574-6976.2010.00253.x/asset/j.1574-6976.2010.00253.x.pdf;jsessionid=F0252E2CE3502A1AC14EF1CC930BE247.f04t01?v=1&t=i1e5c1ru&s=200f77df155555f934571349fb0543e7efe70ea4&systemMessage=Wiley+Online+Library+will+be+disrupted+on+the+18th+October+from+10%3A00+BST+%2805%3A00+EDT%29+for+essential+maintenance+for+approximately+two+hours+as+we+make+upgrades+to+improve+our+services+to+you [1]] | + | In vegetative cells, both operons are repressed by the unstable AbrB regulator. However, during early stages of sporulation AbrB itself is repressed by the master regulator of sporulation Spo0A, making ''sdpABC'' and ''sdpRI'' accessible for RNA polymerase. [http://www.sciencemag.org/content/301/5632/510.full [1]] |
| | | | |
| | === The ''sdpABC'' Operon – Production and Secretion of the Cannibalism Toxin SDP === | | === The ''sdpABC'' Operon – Production and Secretion of the Cannibalism Toxin SDP === |
| | | | |
| - | The production of the Cannibalism Toxin SDP is a multi-step process. The ''sdpC'' sequence encodes the Pro-SdpC1-203,,which is translated by the ribosome.It is a precursor peptide which needs to be processed by a signal peptidases and the two membrane proteins SdpA and SdpB to become functional. This active form of SDP is a 42-amino-acid antimicrobial peptide (AMP) containing a disulfide bond between two cysteine residues located at the N-terminus.(Fig. 2). [http://jb.asm.org/content/195/14/3244.full[2]] | + | The production of the Cannibalism Toxin SDP is a multi-step process. The ''sdpC'' sequence encodes the Pro-SdpC1-203,,which is translated by the ribosome.It is a precursor peptide which needs to be processed by a signal peptidases and the two membrane proteins SdpA and SdpB to become functional. This active form of SDP is a 42-amino-acid antimicrobial peptide (AMP) containing a disulfide bond between two cysteine residues located at the N-terminus.(Fig. 2). [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3697648/ [2]] |
| | | | |
| - | [[File:Background Fig.2.png|thumb|800px|center|Fig. 2. SDP production requires multiple steps. In the cytosol, the full length SdpC (pro-SdpC1-203) is secreted via the Sec pathway. Following secretion, the signal peptidases SipS and SipT cleave the N-terminal signal peptide sequence of SdpC. Disulfide bond formation occurs independently of SdpAB. Finally, posttranslational cleavage of SdpC occurs via SdpAB to produce a 42-amino-acid SDP that will be secreted extracellulary as the active SDP peptide. http://jb.asm.org/content/195/14/3244.full [2]]] | + | [[File:Background Fig.2.png|thumb|800px|center|Fig. 2. SDP production requires multiple steps. In the cytosol, the full length SdpC (pro-SdpC1-203) is secreted via the Sec pathway. Following secretion, the signal peptidases SipS and SipT cleave the N-terminal signal peptide sequence of SdpC. Disulfide bond formation occurs independently of SdpAB. Finally, posttranslational cleavage of SdpC occurs via SdpAB to produce a 42-amino-acid SDP that will be secreted extracellulary as the active SDP peptide. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3697648/ [2] ]]] |
| | | | |
| - | SDP has been shown to be a very effective AMP against a variety of Gram-positive bacteria in the Phylum of the Firmicutes (Fig. 3). It rapidly collapses the proton motive force (PMF), thus inducing autolysis. [3] | + | SDP has been shown to be a very effective AMP against a variety of Gram-positive bacteria in the Phylum of the Firmicutes (Fig. 3). It rapidly collapses the proton motive force (PMF), thus inducing autolysis. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3839633/ [3]] |
| | | | |
| - | [[File:Background Fig.3.png|thumb|600px|center|Fig. 3. SDP inhibition curves for pathogenic microbes. Relative growth of the strains named abovewith the presence of increasing concentrations of SDP is shown in the curve. As a negative control the gram-negative bacteria ''K. pneumoniae'' and ''P. aeruginosa'' are depicted, which are unaffected by the toxin SDP, as it specifically targets gram-positive bacteria. ''B. subtilis'' is a gram-positive bacteria, but expresses the immunity protein SdpI and is therefore relatively resistent to the toxin SDP. the ''Stapylococcus'' species (also MRSA) though are quite drastically reduced in the presence of the SDP. [4]]] | + | [[File:Background Fig.3.png|thumb|600px|center|Fig. 3. SDP inhibition curves for pathogenic microbes. Relative growth of the strains named abovewith the presence of increasing concentrations of SDP is shown in the curve. As a negative control the gram-negative bacteria ''K. pneumoniae'' and ''P. aeruginosa'' are depicted, which are unaffected by the toxin SDP, as it specifically targets gram-positive bacteria. ''B. subtilis'' is a gram-positive bacteria, but expresses the immunity protein SdpI and is therefore relatively resistent to the toxin SDP. the ''Stapylococcus'' species (also MRSA) though are quite drastically reduced in the presence of the SDP. [http://www.pnas.org/content/107/37/16286.full [4] ]]] |
| | | | |
| | === The ''sdpIR'' Operon – Production and Regulation of the Immunity Protein SdpI === | | === The ''sdpIR'' Operon – Production and Regulation of the Immunity Protein SdpI === |
| | | | |
| - | In the absence of SDP the gene coding for the immunity protien DdpI is repressed by the autorepressor SdpR. If extracellular SDP binds the transmembrane protein SdpI it causes a conformational change. As a consequence SdpR is sequestered at the membrane, forming a complex with SDP and SdpI. Since the repression by SdpR is relieved the RNA polymerase can bind the ''sdpRI'' promotor region and start transcription (Fig. 4). [1] | + | In the absence of SDP the gene coding for the immunity protein SdpI is repressed by the autorepressor SdpR. If extracellular SDP binds the transmembrane protein SdpI it causes a conformational change. As a consequence SdpR is sequestered at the membrane, forming a complex with SDP and SdpI. Since the repression by SdpR is relieved the RNA polymerase can bind the ''sdpRI'' promotor region and start transcription (Fig. 4). [http://www.sciencemag.org/content/301/5632/510.full [1]] |
| | | | |
| - | [[File:Background Fig.4.png|thumb|800px|center|Fig. 4. Model of how SDP induces expression of the ''sdpRI'' immunity operon. The signal SDP interacts with the immunity protein SdpI to induce a conformational change that allows SdpI to bind the repressor SdpR, which is sequestered at the membrane. [1]]] | + | [[File:Background Fig.4.png|thumb|800px|center|Fig. 4. Model of how SDP induces expression of the ''sdpRI'' immunity operon. The signal SDP interacts with the immunity protein SdpI to induce a conformational change that allows SdpI to bind the repressor SdpR, which is sequestered at the membrane. [http://www.sciencemag.org/content/301/5632/510.full [1] ]]] |
| | | | |
| | === ECF41: An alternative sigma factor for building an orthogonal Delay Switch === | | === ECF41: An alternative sigma factor for building an orthogonal Delay Switch === |
| | | | |
| - | The alternative σ-factor ''ecf41''<sub>''bli aa 1-204''</sub>, which derives from ''B. licheniformis'' is a truncated version and binds to the target promoter P<sub>''ydfG''</sub>. It is an orthogonal σ-factor since there are no cross-reactions with other native promoters. [5] | + | The alternative σ-factor ''ecf41''<sub>''bli aa 1-204''</sub>, which derives from ''B. licheniformis'' is a truncated version and binds to the target promoter P<sub>''ydfG''</sub>. It is an orthogonal σ-factor since there are no cross-reactions with other native promoters. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3426412/ [5]] The introduction of this loop leads to a translational delay. |
| | | | |
| | === FsrA: a small RNA Represses Expression of Succinate Dehydrogenase === | | === FsrA: a small RNA Represses Expression of Succinate Dehydrogenase === |
| | | | |
| - | Regulation of bacterial iron homeostasis is often controlled by the iron-sensing ferric uptake repressor (Fur). The ''Bacillus subtilis'' Fur protein acts as an iron-dependent repressor for siderophore biosynthesis and iron transport proteins. Furthermore it also coordinates an iron-sparing response that acts to repress the expression of iron-rich proteins (e.g. succinate dehydrogenase) when iron is limited. It has been shown that the downregulation of succinate dehydrogenase (SDH) is most likely caused by complementary binding of a small RNA, named fsrA, to the leader region of the sdhCAB mRNA, namely SdhC' (Fig. 5). In this project this principle is exploited to downregulate the translation of the immunity protein SdpI in a delayed response to the detection of the QS-molecules of ''S. aureus'' or ''S. pneumoniae''. [6] | + | Regulation of bacterial iron homeostasis is often controlled by the iron-sensing ferric uptake repressor (Fur). The ''Bacillus subtilis'' Fur protein acts as an iron-dependent repressor for siderophore biosynthesis and iron transport proteins. Furthermore it also coordinates an iron-sparing response that acts to repress the expression of iron-rich proteins (e.g. succinate dehydrogenase) when iron is limited. It has been shown that the downregulation of succinate dehydrogenase (SDH) is most likely caused by complementary binding of a small RNA, named fsrA, to the leader region of the sdhCAB mRNA, namely SdhC' (Fig. 5). In this project this principle is exploited to downregulate the translation of the immunity protein SdpI in a delayed response to the detection of the QS-molecules of ''S. aureus'' or ''S. pneumoniae''. [http://www.pnas.org/content/105/33/11927.full.pdf [6]] |
| | | | |
| - | [[File:Background Fig.5.png|thumb|800px|center|Fig. 5. Predicted pairing between fsrA and the 5’-leader region of sdhC, the first gene of the sdhCAB operon. [6]]] | + | [[File:Background Fig.5.png|thumb|800px|center|Fig. 5. Predicted pairing between fsrA and the 5’-leader region of sdhC, the first gene of the ''sdhCAB'' operon. [http://www.pnas.org/content/105/33/11927.full.pdf [6] ]]] |
| | | | |
| | === Sources === | | === Sources === |
| | | | |
| - | [1] Gonzalez-Pastor, J. E. (2011). "Cannibalism: a social behavior in sporulating Bacillus subtilis." FEMS Microbiol Rev 35(3): 415-424. | + | [http://www.sciencemag.org/content/301/5632/510.full [1]] Gonzalez-Pastor, J. E. (2011). "Cannibalism: a social behavior in sporulating Bacillus subtilis." FEMS Microbiol Rev 35(3): 415-424. |
| | | | |
| - | [2] Perez Morales, T. G., et al. (2013). "Production of the cannibalism toxin SDP is a multistep process that requires SdpA and SdpB." J Bacteriol 195(14): 3244-3251. | + | [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3697648/ [2]] Perez Morales, T. G., et al. (2013). "Production of the cannibalism toxin SDP is a multistep process that requires SdpA and SdpB." J Bacteriol 195(14): 3244-3251. |
| | | | |
| - | [3] Lamsa, A., et al. (2012). "The Bacillus subtilis cannibalism toxin SDP collapses the proton motive force and induces autolysis." Mol Microbiol 84(3): 486-500. | + | [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3839633/ [3] Lamsa, A., et al. (2012). "The Bacillus subtilis cannibalism toxin SDP collapses the proton motive force and induces autolysis." Mol Microbiol 84(3): 486-500. |
| | | | |
| - | [4] Liu, W. T., et al. (2010). "Imaging mass spectrometry of intraspecies metabolic exchange revealed the cannibalistic factors of Bacillus subtilis." Proc Natl Acad Sci U S A | + | [http://www.pnas.org/content/107/37/16286.full [4]] Liu, W. T., et al. (2010). "Imaging mass spectrometry of intraspecies metabolic exchange revealed the cannibalistic factors of Bacillus subtilis." Proc Natl Acad Sci U S A |
| | | | |
| - | [5] Wecke, T., et al. (2012). "Extracytoplasmic function sigma factors of the widely distributed group ECF41 contain a fused regulatory domain." Microbiologyopen 1(2): 194-213. | + | [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3426412/ [5]] Wecke, T., et al. (2012). "Extracytoplasmic function sigma factors of the widely distributed group ECF41 contain a fused regulatory domain." Microbiologyopen 1(2): 194-213. |
| | | | |
| - | [6] Gaballa, A., et al. (2008). "The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins." Proc Natl Acad Sci U S A 105(33): 11927-11932. | + | [http://www.pnas.org/content/105/33/11927.full.pdf [6]] Gaballa, A., et al. (2008). "The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins." Proc Natl Acad Sci U S A 105(33): 11927-11932. |
| | | | |
| | | | |
| Line 863: |
Line 998: |
| | </html> | | </html> |
| | | | |
| - | The suicide switch module requires a high level of combinability regarding to its components. We therefore designed every new part (link to registry) in RFC10 or RFC25 standard. | + | The suicide switch module requires a high level of combinability regarding to its components. We therefore designed every [https://2014.igem.org/Team:LMU-Munich/Results/Registry new part] in RFC10 or RFC25 standard. |
| | | | |
| | === Production of the toxin SDP and the immunity protein SdpI === | | === Production of the toxin SDP and the immunity protein SdpI === |
| Line 879: |
Line 1,014: |
| | === Delayed loss of immunity === | | === Delayed loss of immunity === |
| | | | |
| - | A time delayed loss of the immunity protein SdpI in BaKillus can be achieved by interposing the alternative σ-factor ECF41<sub>''bli aa 1-204''</sub> which is also under the control of the quorum sensing promoter P<sub>''QS''</sub> (Fig.3A). When expressed, ECF41<sub>''bli aa 1-204''</sub> binds to the target promoter P<sub>''ydfG''</sub> leading either to the expression of fsrA, a small RNA, or transcription of sdpI-antisense, a RNA, a complementary RNA to the sdpI mRNA (Fig. 3B). | + | A time delayed loss of the immunity protein SdpI in BaKillus can be achieved by interposing the alternative σ-factor ECF41<sub>''bli aa 1-204''</sub> [http://parts.igem.org/Part:BBa_K823043 BBa_K823043] which is also under the control of the quorum sensing promoter P<sub>''QS''</sub> (Fig.3A). When expressed, ECF41<sub>''bli aa 1-204''</sub> binds to the target promoter P<sub>''ydfG''</sub> [http://parts.igem.org/Part:BBa_K823041 BBa_K823041] leading either to the expression of fsrA, a small RNA, or transcription of sdpI-antisense, a RNA, a complementary RNA to the sdpI mRNA (Fig. 3B). |
| | To veri- and quantify the time delay due to the interposition of Ecf41<sub>''bli aa 1-204''</sub> we replaced the fsrA coding sequence with the lux-cassette ''luxABCDE'' allowing lumi assay measurements (Fig. 3C). | | To veri- and quantify the time delay due to the interposition of Ecf41<sub>''bli aa 1-204''</sub> we replaced the fsrA coding sequence with the lux-cassette ''luxABCDE'' allowing lumi assay measurements (Fig. 3C). |
| | | | |
| | [[File:Design Fig.3.png|600px|center|Fig. 3.]] | | [[File:Design Fig.3.png|600px|center|Fig. 3.]] |
| | | | |
| - | For the loss of immunity we implemented two strategies, both causing translational inhibition of the sdpI mRNA. | + | For the loss of immunity we implemented two strategies, both causing translational inhibition of the ''sdpI'' mRNA. |
| | The first strategy is based on an artificial synthesized RNA called sdpI-antisense RNA, which is complementary to the sdpI mRNA sequence. The sdpI-antisense sequence is regulated by P<sub>''ydfG''</sub> (Fig. 4A) which is activated by the binding of Ecf41<sub>''bli aa 1-204''</sub>. As a consequence the sdpI-antisense RNA and the sdpI mRNA form a double-stranded duplex which is inaccessible for the ribosome making the cell SDP-sensitive. | | The first strategy is based on an artificial synthesized RNA called sdpI-antisense RNA, which is complementary to the sdpI mRNA sequence. The sdpI-antisense sequence is regulated by P<sub>''ydfG''</sub> (Fig. 4A) which is activated by the binding of Ecf41<sub>''bli aa 1-204''</sub>. As a consequence the sdpI-antisense RNA and the sdpI mRNA form a double-stranded duplex which is inaccessible for the ribosome making the cell SDP-sensitive. |
| | We tested the sdpI-antisense RNA in B. subtilis W168 with the xylose-inducable promoter P<sub>''xyl''</sub>. (Fig. 4B) | | We tested the sdpI-antisense RNA in B. subtilis W168 with the xylose-inducable promoter P<sub>''xyl''</sub>. (Fig. 4B) |
| Line 890: |
Line 1,025: |
| | [[File:Design Fig.4.png|600px|center|Fig. 4.]] | | [[File:Design Fig.4.png|600px|center|Fig. 4.]] |
| | | | |
| - | The second strategy involves fsrA, a small RNA regulated by P<sub>ydfG</sub> which binds to its binding site sdhC' (the 5'-UTR of sdhC mRNA) which is cloned between P<sub>''sdpRI''<sub> and ''sdpI'' (Fig. 5A). Binding of fsrA to the sdhC' site on the transcript inhibits the ribosomal translation process of the sdpI-mRNA. | + | The second strategy involves fsrA, a small RNA regulated by P<sub>ydfG</sub> which binds to its binding site sdhC' (the 5'-UTR of sdhC mRNA) which is cloned between P<sub>''sdpRI''</sub> and ''sdpI'' (Fig. 5A). Binding of fsrA to the sdhC' site on the transcript inhibits the ribosomal translation process of the sdpI-mRNA. |
| | To evaluate this small RNA and its binding site in ''B. subtilis'' we designed two strains. One ''∆abrB ∆sdpI'' mutant strain where we replaced the P<sub>''ydfG''</sub> promoter with the P<sub>''xyl''</sub> promoter (Fig. 5B). The second construct containing fsrA under control of P<sub>''xyl''</sub> and replaced P<sub>''sdpRI''</sub>-''sdhC'sdpI'' with the P<sub>''lepA''</sub>-''sdhC'gfp'', where GFP is consecutively expressed, (Fig. 5C) was transformed into ''B. subtilis'' W168. | | To evaluate this small RNA and its binding site in ''B. subtilis'' we designed two strains. One ''∆abrB ∆sdpI'' mutant strain where we replaced the P<sub>''ydfG''</sub> promoter with the P<sub>''xyl''</sub> promoter (Fig. 5B). The second construct containing fsrA under control of P<sub>''xyl''</sub> and replaced P<sub>''sdpRI''</sub>-''sdhC'sdpI'' with the P<sub>''lepA''</sub>-''sdhC'gfp'', where GFP is consecutively expressed, (Fig. 5C) was transformed into ''B. subtilis'' W168. |
| | | | |
| Line 921: |
Line 1,056: |
| | }, | | }, |
| | subtitle: { | | subtitle: { |
| - | text: 'technical replicas in three measurements', | + | text: 'three technical replicas of three independant measurements', |
| | x: -20 | | x: -20 |
| | }, | | }, |
| Line 2,529: |
Line 2,664: |
| | </script> | | </script> |
| | <br> | | <br> |
| - | The graph shows wildtype <i>Bacillus subtilis</i>, the ΔsdpI mutant | + | The graph shows wildtype <i>Bacillus subtilis</i>, the ΔsdpI mutant as well as a ΔsdpI mutant carrying our generated </html>([http://parts.igem.org/Part:BBa_K1351017 BBa_K1351017]) in addition the graph shows a wildype <i>Bacillus subtilis</i> carrying an antisense RNA [http://parts.igem.org/Part:BBa_K1351019 (BBa_K1351019)] sequence which should bind the mRNA of the sdpI immunity. The LUMI-assay shows a normal growth curve of <i>Bacillus subtilis</i> and a ΔsdpI mutant which degrades in early to late stationary phase. <html>The two from us constructed devices show in case of the inducible immunity in the ΔsdpI mutant an almost recovery in growth behavior when compared to wild type. The antisense carrying W168-strain showed normal wildtype growth and it seems like that an translational inhibition of the immunity was not functioning. <br><br><br> |
| | + | |
| | + | For the suicide switch we figured that a time delay was necessary before the suicide switch kicks in so that BaKillus has enough time to do his job. Therefore we decided to test if an insertion of an alternative sigma factor would lead to a time delay. For this purpose we designed two constructs: one having an inducible promoter in front of a lux-readout and the other device carrying an alternative sigma (ECF41 </html>[http://parts.igem.org/Part:BBa_K823043 BBa_K823043]<html>) factor after the same inducible promoter. The sigma factor has to find its own specific promoter again (P<sub>ydfG</sub> </html>[http://parts.igem.org/Part:BBa_K823041 BBa_K823041]<html>) which was cloned in front of the lux-operon.<br><br> |
| | + | The </html>[https://static.igem.org/mediawiki/2014/c/c7/LMU-Munich14_Luminescence_Assay.pdf LUMI-assay]<html> in the graph shows that we could achieve a time delay of about ten minutes using the alternative sigma factor route. For a reasonable delay we intend to implement a series of alternative sigma factors and their target promoters to achieve a cascade and therefore increase the time delay. |
| | + | <br> |
| | + | </html> |
| | + | [[File:14lmuecf.jpg|thumb|500px|center|Fig. 2. ECF41-mediated time delay]]<html> |
| | </article> | | </article> |
| | </div> | | </div> |
 "
"