Team:ETH Zurich/expresults/rr
From 2014.igem.org
(→Riboregulators) |
(→Riboregulators) |
||
| Line 13: | Line 13: | ||
| - | First, the experiments were conducted with cells | + | First, the experiments were conducted with cells transformed with two different pPaB plasmids. These plasmids contained either the pBR322 origin (pMB1) or the p15A origin, which yielded a stable two-plasmid system with about 15-20 copies per plasmid and cell (medium-copy). Also, exactly the published riboregulator sequences were used in the the first place to have a positive control for functionality. This is important, because the original riboregulator sequence includes two forbidden restriction sites (XbaI and SpeI). Second, the restriction sites were removed by two different approaches: a) multiple site-directed mutagenesis, b) blunting and ligation (Klenow and T4 DNA polymerase). Afterwards, the functionality of the construct was again tested to approve that no severe loss-of-function occurred due to the site-removal. In a third step, the construct was transferred into the [http://parts.igem.org/Part:pSB1C3 pSB1C3] backbone (pMB1 origin, high copy number of 100-300) to be in line with the BioBrick Standard (ref to standard). Below, you can find the corresponding graphs. |
Revision as of 18:07, 16 October 2014
Riboregulators
As many former iGEM teams have encountered unwanted basal expression (leakiness) (ref ETH2013, Groningen2012 amongst others) of their genes of interest (goi) , we decided to further investigate this severe obstacle to tightly controllable systems. We started looking into the available literature and found with riboregulators a promising approach to challenge the issue of leakiness (ref isaacs). In addition, we chose this approach, to study and characterize such regulators in combination with cell-to-cell communication (ref quorum sensing) and make the resulting parts available for the iGEM community in the Registry of Standard Biological Parts (ref registry).
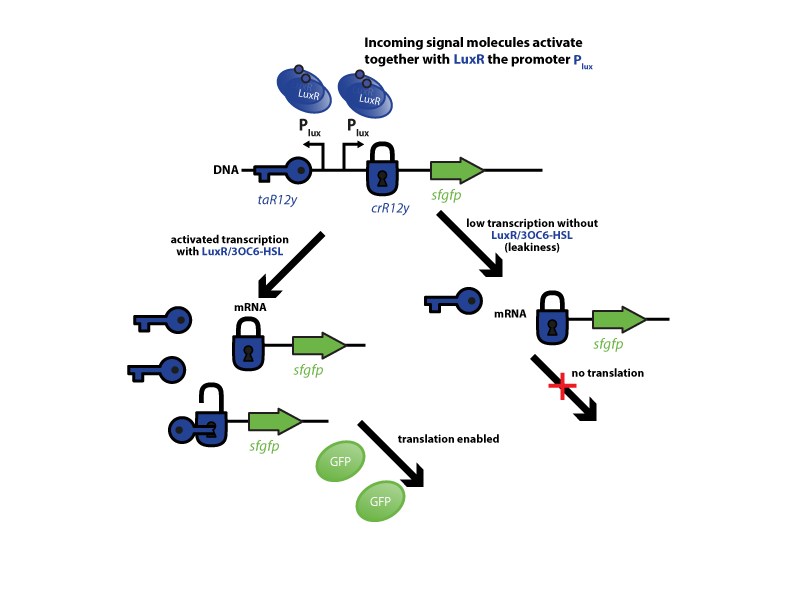
The riboregulator systems include two parts: 1) a cis-repressed RBS in front of the goi, and 2) a co-expressed trans-activating RNA.
To characterize this system for reliable cell-to-cell communication, we combined the riboregulator parts (ref to parts) with promoters of the quorum-sensing systems used by our team (LuxI/LuxR, LasI/LasR, and RhlI/RhlR (ref to qs parts)). In our system, the riboregulators were intended for the control of integrases (ref). However, to have an easily quantifiable output, we used GFP as our riboregulated goi and measured the output fluorescence with a Tecan plate reader (ref to materials). This approach is described in the figure below.
First, the experiments were conducted with cells transformed with two different pPaB plasmids. These plasmids contained either the pBR322 origin (pMB1) or the p15A origin, which yielded a stable two-plasmid system with about 15-20 copies per plasmid and cell (medium-copy). Also, exactly the published riboregulator sequences were used in the the first place to have a positive control for functionality. This is important, because the original riboregulator sequence includes two forbidden restriction sites (XbaI and SpeI). Second, the restriction sites were removed by two different approaches: a) multiple site-directed mutagenesis, b) blunting and ligation (Klenow and T4 DNA polymerase). Afterwards, the functionality of the construct was again tested to approve that no severe loss-of-function occurred due to the site-removal. In a third step, the construct was transferred into the [http://parts.igem.org/Part:pSB1C3 pSB1C3] backbone (pMB1 origin, high copy number of 100-300) to be in line with the BioBrick Standard (ref to standard). Below, you can find the corresponding graphs.
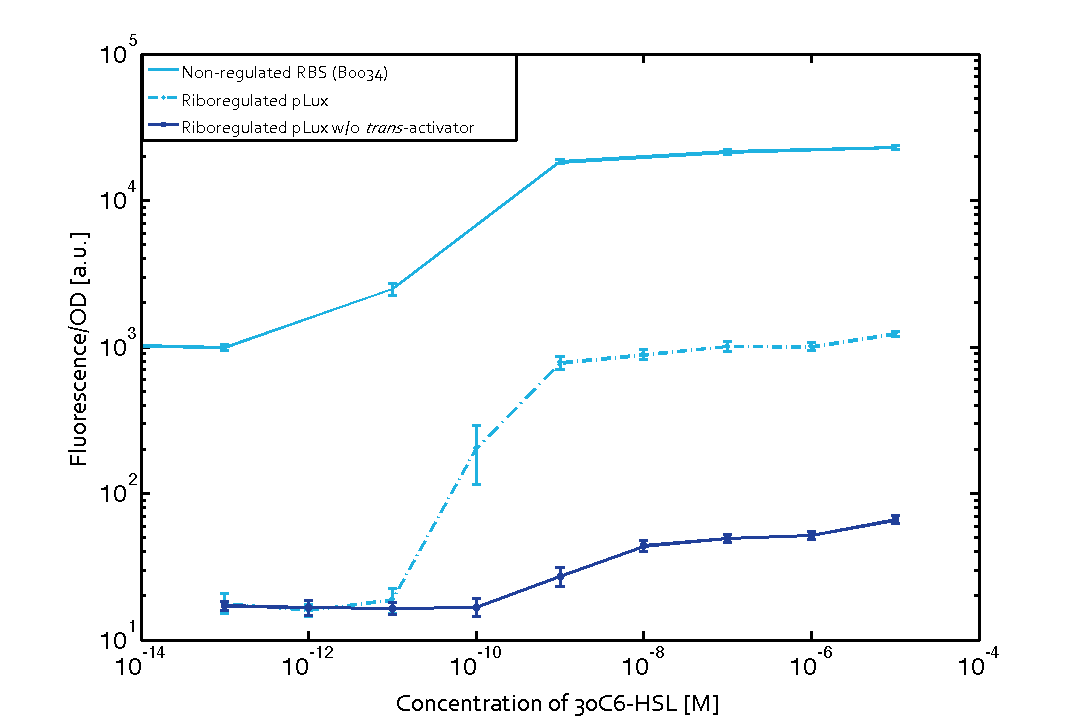
In more detail, we have characterized promoters of the three quorum-sensing systems. These are pLux, pLas, and pRhl. To have a reference, we first measured the transfer functions of the non-regulated GFP (ref constructs), i.e. the amount of fluorescence in dependence of the inducer concentration present. Next, we repeated the experiments with the corresponding riboregulated constructs (ref constructs) once with and without the forbidden restriction sites.
To conclude, we found a x-fold reduced basal GFP expression when using our riboregulator construct. Moreover, we achieved an increased signal-to-noise ratio of z-fold, as compared to the non-regulated system. Finally, the BioBrick conform part was confirmed to show no loss-of-funtion. However, the sensitivity towards the inducer was slightly reduced. This results are summarized in the graphs below.
 "
"