Team:Tianjin/Project
From 2014.igem.org
(Difference between revisions)
(Prototype team page) |
|||
| (4 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | |||
| - | |||
| - | |||
{{CSS/Main}} | {{CSS/Main}} | ||
| + | <div id="content" | ||
| + | border:0px;border-color: transparent;> | ||
| + | </div> | ||
| + | <html xmlns="http://www.w3.org/1999/xhtml"> | ||
| + | <head> | ||
| + | <meta http-equiv="Content-Type" content="text/html; charset=utf-8" /> | ||
| + | <title>Team:Tianjin2014/Safety</title> | ||
| + | <style type="text/css"> | ||
| + | <!-- | ||
| + | body { | ||
| + | margin-left: -330px; | ||
| + | margin-right: 20px; | ||
| + | margin-top:0px; | ||
| + | margin-bottom: 1%; | ||
| + | background-color: #663300; | ||
| + | } | ||
| + | #search-controls{display:none;} | ||
| + | .firstHeading{display:none;border:0px;} | ||
| + | #siteSub{display:none;} | ||
| + | #content{ | ||
| + | border-color: transparent;} | ||
| + | .firstHeading{display:none;border:0px;} | ||
| + | .fixed{ | ||
| + | height:300px; | ||
| + | width:200px; | ||
| + | position:fixed; | ||
| + | top:290px; | ||
| + | right:10px; | ||
| + | border: medium dashed #C67B14; | ||
| + | background-color: #993300; | ||
| + | } | ||
| + | #jq_fix{height:100px;width:100px;background:#313;} | ||
| + | a:link { | ||
| + | color: #FFFFCC; | ||
| + | text-decoration: none; | ||
| + | } | ||
| + | a:visited { | ||
| + | text-decoration: none; | ||
| + | color: #CCCC99; | ||
| + | } | ||
| + | a:hover { | ||
| + | text-decoration: none; | ||
| + | color: #FF9933; | ||
| + | } | ||
| + | a:active { | ||
| + | text-decoration: none; | ||
| + | } | ||
| + | #project { | ||
| + | position:absolute; | ||
| + | width:80px; | ||
| + | height:24px; | ||
| + | z-index:2; | ||
| + | } | ||
| + | #Layer8 { | ||
| + | position:absolute; | ||
| + | width:180px; | ||
| + | height:150px; | ||
| + | z-index:1; | ||
| + | top: 30px; | ||
| + | left: 0; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #home { | ||
| + | position:absolute; | ||
| + | width:80px; | ||
| + | height:24px; | ||
| + | z-index:3; | ||
| + | } | ||
| + | #judging { | ||
| + | position:absolute; | ||
| + | width:80px; | ||
| + | height:24px; | ||
| + | z-index:4; | ||
| + | } | ||
| + | #Layer11 { | ||
| + | position:absolute; | ||
| + | width:115px; | ||
| + | height:50px; | ||
| + | z-index:1; | ||
| + | left: 0; | ||
| + | top: 30px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #notebook { | ||
| + | position:absolute; | ||
| + | width:80px; | ||
| + | height:24px; | ||
| + | z-index:5; | ||
| + | } | ||
| + | #Layer13 { | ||
| + | position:absolute; | ||
| + | width:130px; | ||
| + | height:50px; | ||
| + | z-index:1; | ||
| + | top: 30px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #team { | ||
| + | position:absolute; | ||
| + | width:101px; | ||
| + | height:24px; | ||
| + | z-index:10; | ||
| + | } | ||
| + | #safety { | ||
| + | position:absolute; | ||
| + | width:80px; | ||
| + | height:24px; | ||
| + | z-index:11; | ||
| + | } | ||
| + | #human-practice { | ||
| + | position:absolute; | ||
| + | width:130px; | ||
| + | height:24px; | ||
| + | z-index:8; | ||
| + | } | ||
| + | #Layer17 { | ||
| + | position:absolute; | ||
| + | width:130px; | ||
| + | height:75px; | ||
| + | z-index:9; | ||
| + | left: 0px; | ||
| + | top: 24px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | .STYLE18 { | ||
| + | font-family: Arial, Helvetica, sans-serif; | ||
| + | font-weight: bold; | ||
| + | font-size: x-large; | ||
| + | color: #006600; | ||
| + | } | ||
| + | .STYLE29 { | ||
| + | font-size: 36px; | ||
| + | font-family: Jokerman; | ||
| + | } | ||
| + | .STYLE33 { | ||
| + | font-size: 18px; | ||
| + | font-family: "Comic Sans MS"; | ||
| + | } | ||
| + | .STYLE40 { | ||
| + | font-family: "Kristen ITC"; | ||
| + | color: #C67B18; | ||
| + | } | ||
| + | .STYLE43 {font-family: Jokerman; | ||
| + | font-size: 72px; | ||
| + | color: #C67B18; | ||
| + | } | ||
| + | .STYLE24 {font-size: 96px} | ||
| + | .STYLE45 {font-family: "Kristen ITC"; font-size: 24px; color: #FFFFFF; font-weight: bold; } | ||
| + | .STYLE46 {font-family: "Kristen ITC"} | ||
| + | .STYLE47 {font-size: 24px; font-family: "Kristen ITC"; font-weight: bold; } | ||
| + | .STYLE48 {font-weight: bold; font-size: 24px;} | ||
| + | #Layer18 { | ||
| + | position:absolute; | ||
| + | width:260px; | ||
| + | height:115px; | ||
| + | z-index:9; | ||
| + | left: 28px; | ||
| + | top: 458px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #Layer19 { | ||
| + | position:absolute; | ||
| + | width:137px; | ||
| + | height:78px; | ||
| + | z-index:1; | ||
| + | left: 0; | ||
| + | top: 30; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #Layer7 { | ||
| + | position:absolute; | ||
| + | width:222px; | ||
| + | height:149px; | ||
| + | z-index:1; | ||
| + | left: 0; | ||
| + | top: 30; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #Layer9 { | ||
| + | position:absolute; | ||
| + | width:200px; | ||
| + | height:115px; | ||
| + | z-index:12; | ||
| + | } | ||
| + | #Layer10 { | ||
| + | position:absolute; | ||
| + | width:452px; | ||
| + | height:115px; | ||
| + | z-index:12; | ||
| + | left: 38px; | ||
| + | top: 400px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #Layer12 { | ||
| + | position:absolute; | ||
| + | width:551px; | ||
| + | height:508px; | ||
| + | z-index:1; | ||
| + | left: 457px; | ||
| + | top: 0px; | ||
| + | } | ||
| + | #Layer14 { | ||
| + | position:absolute; | ||
| + | width:200px; | ||
| + | height:115px; | ||
| + | z-index:13; | ||
| + | left: 0; | ||
| + | top: 0; | ||
| + | } | ||
| + | #Layer15 { | ||
| + | position:absolute; | ||
| + | width:200px; | ||
| + | height:115px; | ||
| + | z-index:12; | ||
| + | left: 528px; | ||
| + | top: 410px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #Layer16 { | ||
| + | position:absolute; | ||
| + | width:200px; | ||
| + | height:115px; | ||
| + | z-index:12; | ||
| + | left: 528px; | ||
| + | top: 410px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #Layer20 { | ||
| + | position:absolute; | ||
| + | width:200px; | ||
| + | height:115px; | ||
| + | z-index:13; | ||
| + | left: 528px; | ||
| + | top: 410px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | #Layer21 { | ||
| + | position:absolute; | ||
| + | width:200px; | ||
| + | height:115px; | ||
| + | z-index:14; | ||
| + | left: 528px; | ||
| + | top: 410px; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | .STYLE61 {font-family: "Comic Sans MS"; font-size: 18px; color: #C67B14; } | ||
| + | .STYLE62 {font-size: 36px; font-family: "Kristen ITC"; } | ||
| + | .STYLE69 {font-family: Jokerman; font-size: 20px; } | ||
| + | .STYLE76 {font-size: 90px} | ||
| + | .STYLE77 {font-size: 120px} | ||
| + | #Layer22 { | ||
| + | position:absolute; | ||
| + | width:200px; | ||
| + | height:115px; | ||
| + | z-index:12; | ||
| + | left: 0; | ||
| + | top: 30; | ||
| + | visibility: hidden; | ||
| + | } | ||
| + | --> | ||
| + | ul,li{list-style:none;} | ||
| + | #content{width:600px;margin:0 auto;border:1px solid #f00;} | ||
| + | ul li.item{width:400px;text-align:center;margin:20px 100px;background:#00f;color:#fff;font-size:14px;font-weight:bold;height:100px;line-height:100px;} | ||
| + | #float_banner{position:absolute;top:0;left:50%;width:900px;margin-left:-450px;height:30px;line-height:30px;text-align:center;background:#000;color:#fff;font-size:14px;font-weight:bold;z-index:2;} | ||
| + | .STYLE78 {font-family: Jokerman; font-size: 20px; font-weight: bold; } | ||
| + | .STYLE79 {font-size: 18px; font-family: "Comic Sans MS"; font-weight: bold; } | ||
| + | </style> | ||
| + | <script type="text/JavaScript" language="javascript"> | ||
| + | <!-- | ||
| - | |||
| - | + | $(document).ready(function(){ | |
| - | + | //滚动事件 | |
| + | $(window).scroll(function(){ | ||
| + | //显示滚动条位置 | ||
| + | $(".fixed").html(document.body.scrollTop||document.documentElement.scrollTop); | ||
| + | var t = document.body.scrollTop||document.documentElement.scrollTop; | ||
| + | //当导航条不可见时,重新定义导航条位置 | ||
| + | if(t>100){ | ||
| + | $("#jq_fix").css({ | ||
| + | "position":"fixed", | ||
| + | "top" :"0px", | ||
| + | "left" :"0px" | ||
| + | }); | ||
| + | }else{ | ||
| + | //回到导航条原来位置时,还原导航条位置 | ||
| + | $("#jq_fix").css({ | ||
| + | "position":"", | ||
| + | "top" :"", | ||
| + | "left" :"" | ||
| + | }); | ||
| + | } | ||
| + | }); | ||
| + | }); | ||
| + | function MM_findObj(n, d) { //v4.01 | ||
| + | var p,i,x; if(!d) d=document; if((p=n.indexOf("?"))>0&&parent.frames.length) { | ||
| + | d=parent.frames[n.substring(p+1)].document; n=n.substring(0,p);} | ||
| + | if(!(x=d[n])&&d.all) x=d.all[n]; for (i=0;!x&&i<d.forms.length;i++) x=d.forms[i][n]; | ||
| + | for(i=0;!x&&d.layers&&i<d.layers.length;i++) x=MM_findObj(n,d.layers[i].document); | ||
| + | if(!x && d.getElementById) x=d.getElementById(n); return x; | ||
| + | } | ||
| + | function MM_showHideLayers() { //v6.0 | ||
| + | var i,p,v,obj,args=MM_showHideLayers.arguments; | ||
| + | for (i=0; i<(args.length-2); i+=3) if ((obj=MM_findObj(args[i]))!=null) { v=args[i+2]; | ||
| + | if (obj.style) { obj=obj.style; v=(v=='show')?'visible':(v=='hide')?'hidden':v; } | ||
| + | obj.visibility=v; } | ||
| + | } | ||
| - | + | function MM_preloadImages() { //v3.0 | |
| - | + | var d=document; if(d.images){ if(!d.MM_p) d.MM_p=new Array(); | |
| - | + | var i,j=d.MM_p.length,a=MM_preloadImages.arguments; for(i=0; i<a.length; i++) | |
| - | < | + | if (a[i].indexOf("#")!=0){ d.MM_p[j]=new Image; d.MM_p[j++].src=a[i];}} |
| - | + | } | |
| - | + | //--> | |
| - | + | </script> | |
| - | + | </head> | |
| - | </ | + | |
| - | </ | + | |
| - | <tr> <td colspan="3" height=" | + | <body> |
| - | < | + | <div class="fixed"> |
| - | <tr> | + | <table width="100%" border="5" cellpadding="0" cellspacing="0" bordercolor="#993300" bgcolor="#993300"> |
| + | <tr bgcolor="#3A2108"> | ||
| + | <td height="40"><span class="STYLE78"><a href="#part1">Overview</a></span></td> | ||
| + | </tr> | ||
| + | <tr bgcolor="#3A2108"> | ||
| + | <td height="40"><span class="STYLE78"><a href="#part2">Background</a></span></td> | ||
| + | </tr> | ||
| + | <tr bgcolor="#3A2108"> | ||
| + | <td height="40"><span class="STYLE78"><a href="#part3">Design</a></span></td> | ||
| + | </tr> | ||
| + | <tr bgcolor="#3A2108"> | ||
| + | <td height="40" class="STYLE78"><a href="#part4"><strong>Results</strong></a></td> | ||
| + | </tr> | ||
| + | <tr bgcolor="#3A2108"> | ||
| + | <td height="40" class="STYLE69"><a href="#part5"><strong>Future Work</strong></a> </td> | ||
| + | </tr> | ||
| + | <tr bgcolor="#3A2108"> | ||
| + | <td height="40" class="STYLE78"><a href="https://2014.igem.org/Team:Tianjin/Project2#part6"><strong>Modeling</strong></a></td> | ||
| + | </tr> | ||
| + | <tr bgcolor="#3A2108"> | ||
| + | <td height="35" bgcolor="#663300"><span class="STYLE78"><a href="#top">Go to the top</a></span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div> | ||
| + | <table width="1230" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td width="19%" bgcolor="#3A2108"><p><a href="https://2014.igem.org/Main_Page"><img src="https://static.igem.org/mediawiki/2014/5/5d/Team:Tianjin_image_IGEM_official_logo.png" alt="image igem" width="205" height="114" border="0" align="top" /></a><br /> | ||
| + | </p> | ||
| + | <p> </p> | ||
| + | <p> </p> | ||
| + | <p> </p></td> | ||
| + | <td width="43%" bgcolor="#3A2108"><p> </p> | ||
| + | <p> </p> | ||
| + | <p> </p> | ||
| + | <p><span class="STYLE43"><span class="STYLE24"><a name="top" id="top"></a><span class="STYLE77">T</span></span><span class="STYLE76">rans<em>f</em> ibre</span></span><br /> | ||
| + | </p> | ||
| + | <p> </p> | ||
| + | <p><span class="STYLE40">Welcome to Team Tianjin! </span><br /> | ||
| + | </p></td> | ||
| + | <td width="38%" bgcolor="#3A2108"><img src="https://static.igem.org/mediawiki/2014/e/eb/Tianjin-head3.gif" width="456" height="226" align="right" /></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <table width="1230" border="0" cellpadding="0" cellspacing="0" bgcolor="#663300"> | ||
| + | <tr> | ||
| + | <td width="157" height="21" bgcolor="#663300"><div id="home"> | ||
| + | <table width="100" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#663300"><span class="STYLE45"><a href="https://2014.igem.org/Team:Tianjin">Home|</a></span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </div></td> | ||
| + | <td width="169" bgcolor="#663300"><div class="STYLE46" id="project" onmouseover="MM_showHideLayers('Layer8','','show')" onmouseout="MM_showHideLayers('Layer8','','hide')"><a href="https://2014.igem.org/Team:Tianjin/Project" class="STYLE48">Project|</a></div></td> | ||
| + | <td width="174" bgcolor="#663300"><div id="judging" onmouseover="MM_showHideLayers('Layer11','','show')" onmouseout="MM_showHideLayers('Layer11','','hide')"> | ||
| + | <table width="110" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#663300"><a href="https://2014.igem.org/Team:Tianjin/Judging" class="STYLE47">Judging|</a></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </div></td> | ||
| + | <td width="200" bgcolor="#663300"><div id="notebook" onmouseover="MM_showHideLayers('Layer13','','show')" onmouseout="MM_showHideLayers('Layer13','','hide')"> | ||
| + | <table width="110" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#663300"><a href="https://2014.igem.org/Team:Tianjin/Notebook" class="STYLE47">Notebook|</a></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </div></td> | ||
| + | <td width="146" bgcolor="#663300"><div id="team" onmouseover="MM_showHideLayers('Layer7','','show')" onmouseout="MM_showHideLayers('Layer7','','hide')"> | ||
| + | <div id="layer"> | ||
| + | <table width="100" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#663300"><span class="STYLE45"><a href="https://2014.igem.org/Team:Tianjin/Team">Team|</a></span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div></td> | ||
| + | <td width="159" bgcolor="#663300"><div id="safety" onmouseover="MM_showHideLayers('Layer19','','show')" onmouseout="MM_showHideLayers('Layer19','','hide')"> | ||
| + | <div id="layer2"> | ||
| + | <table width="100" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#663300"><span class="STYLE45"><a href="https://2014.igem.org/Team:Tianjin/Safety">Safety|</a></span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div></td> | ||
| + | <td width="225" bgcolor="#663300"><div id="human-practice" onmouseover="MM_showHideLayers('Layer22','','show')" onmouseout="MM_showHideLayers('Layer22','','hide')"> | ||
| + | <div id="layer3"> | ||
| + | <table width="100" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#663300"><span class="STYLE45"><a href="https://2014.igem.org/Team:Tianjin/Human_Practice">Human Practice|</a></span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </div> | ||
| + | </div></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <br /> | ||
| + | <table width="1230" border="0" align="center" cellpadding="0" cellspacing="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#993300"><table width="809" border="10" align="center" cellpadding="0" cellspacing="0" bordercolor="#C67B14" bgcolor="#993300"> | ||
| + | <tr> | ||
| + | <td width="100%" bgcolor="#993300"><table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#C67B14"><span class="STYLE62"><a name="part1" id="part1"></a><strong>①Overview</strong></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97" class="STYLE33"><p>As to lower organisms, there seems an indestructible barrier between biological signals and electrical signals. There have been bits of research results on the transformation from electrical signals to biological signals, for example, the phosphoinositide phosphatase activity coupled to an intrinsic voltage sensor in Ciona intestinalis. However, the opposite direction of transformation is almost a blank space, especially in microbes. Thus, we want to build a bridge between them. With this, microbes can accomplish detection with electronic output signals, and even ‘operate’ machines!</p> | ||
| + | <p>The bridge is the transformed curli fiber linked to nanogold. When inducer is added in the system, ‘transfibers’ take shape and nanogolds are linked together as wires, so the circuit is ‘on’. In terms of detection, it can be characterized by electric current change. Meanwhile, other machines can be switched on by means of this.</p> | ||
| + | <p>To learn more, follow us and enjoy the travel of ‘TRANS’!</p></td> | ||
| + | </tr> | ||
| + | </table></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td height="56" bgcolor="#C67B14"><span class="STYLE29"><a name="part2" id="part2"></a><span class="STYLE62"><strong>②Background</strong></span></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><p class="STYLE33"><strong>Curli fiber</strong></p> | ||
| + | <p class="STYLE33"><strong>1.What is curli fiber?</strong></p> | ||
| + | <p class="STYLE33">Bacteria are able to integrate and survive in a remarkably diverse collection of environments.</p> | ||
| + | <p class="STYLE33">In recent years, bacterial communities have been better appreciated as an integral part of most microbial lifestyles. These communities, or biofilms, are prominent during infections and are generally characterized by an extracellular matrix that can help sculpt three-dimensional structures, which promote the survival of its inhabitants in the face of environmental stresses. Enteric bacteria such as Escherichia coli and Salmonella spp. express proteinaceous extracellular fibers called “<strong>curli</strong>” that are involved in surface and cell-cell contacts that promote community behavior and host colonization.</p> | ||
| + | <p class="STYLE33">Curli are the major proteinaceous component of a complex extra-cellular matrix. Curli fibers are involved in adhesion to surfaces, cell aggregation and biofilm formation. Curli also mediate host cell adhesion and invasion, and they are potent inducers of the host inflammatory response.</p> | ||
| + | <p class="STYLE33">Structurally and biochemically, curli belong to a growing class of fibers known as amyloids. Amyloid fiber formation is responsible for several human diseases including Alzheimer's, Huntington's, and prion diseases. Curli share all of the biophysical properties of amyloids: ordered β-sheet-rich fibers resistant to proteases and other harsh denaturants with a capacity to bind the dyes CR and ThT. In contrast to disease-associated amyloids, curli assembly is not the result of protein misfolding. Instead, curli are the product of a dedicated biogenesis pathway providing a paradigm for understanding controlled amyloidogenesis. </p> | ||
| + | <p class="STYLE33"><strong>2.Why do we choose curli?</strong> </p> | ||
| + | <p class="STYLE33">Curli assembly is guided by the products of seven curlispecific genes (csg) encoded on two divergently transcribed operons, <em>csgDEFG</em> and <em>csgBAC</em>. CsgD is the master regulator of curli biogenesis and is required for transcription of the csgBAC operon.</p> | ||
| + | <p class="STYLE33">Our model system is based on the self-assembly of the secreted major curli subunit CsgA. Secreted CsgA monomers are templated on CsgB, which is anchored to the cell surface, to form curli fibre; moreover, CsgA secreted from one cell can interact with CsgB on other cells. Situated at the cell surface, CsgB nucleates soluble, unstructured CsgA into a highly ordered amyloid fiber. </p> | ||
| + | <p class="STYLE33">To transport the major submit CsgA and the minor submit CsgB into the extracellular milieu, the lipoprotein CsgG oligomerizes into a pore-like structure in the outer membrane.<br /> | ||
| + | In addition, the secretion and localization of curli fiber subunits through CsgG are mediated by three additional proteins: CsgC, CsgE and CsgF. CsgC was predicted to have oxidoreductase activity, regulating CsgG outer membrane assembly and pore activity. CsgE could be considered a CsgA-specific chaperone. Besides, CsgF associates with the outer membrane and is required for cell association of the minor curli fiber subunit CsgB.</p> | ||
| + | <p class="STYLE33">As explained above, curli have two remarkable advantages. One is the self-assembly of CsgA monomers into mature amyloids----curli. Apart from it, using synthetic riboregulators, we can implement inducible transcriptional and translational control over the expression of CsgA subunits engineered to display various peptide tags, which can interface with inorganic materials, such as gold nanoparticles, to form a conductive structure. Utilizing these advantages, we can design a transducer which can convert the change of gene expression level that stimulated by inducer directly into the electric signal via inductive synthesis(or destroy)of nanowire between the electrodes.</p> | ||
| + | <p class="STYLE33"><strong>Gold Nanoparticles (AuNPs)</strong> </p> | ||
| + | <p class="STYLE33">AuNPs are the most stable metal nanoparticles, and they present fascinating aspects such as their assembly of multiple types involving materials science, the behavior of the individual particles, size-related electronic, magnetic and optical properties (quantum size effect), and their applications to catalysis and biology.</p> | ||
| + | <p class="STYLE33">Since we can interface curli fiber with inorganic materials to produce amyloid-based wires that are either externally controllable or undergo autonomous patterning, a material with completely bio-compatibility, conductivity and stability is required. AuNPs can be linked to some peptides, providing fascinating propoties. For instance, the assembly of relatively short polypeptides on curli can provide protein-like complex structures and conductivity in conjugates with AuNPs.</p> | ||
| + | <p class="STYLE33"><strong>Nanowire-based devices</strong></p> | ||
| + | <p class="STYLE33">Devices based on nanowires have emerged as one of the most powerful and general platforms for ultrasensitive, direct electrical detection of biological and chemical species and for building functional interfaces to biological systems. With collective efforts and creation of researchers all around the world, there comes many kinds of nanosensors for various uses such as ultrasensitive detection of proteins and individual virus particles as well as recording, stimulation, and inhibition of neuronal signals in nanowire–neuron hybrid structures.</p> | ||
| + | <p class="STYLE33">The advantages of nanowires lie in many ways. Firstly, nanowires are enabled reproducible synthesis of nanowires of homogeneous composition and diameter with controllable electronic and optical properties. Moreover, branched nanowire structures with unique functions have been built in at the stage of synthesis. Significantly, In general, the similarity in size of nanowires and natural nanostructures in biological systems makes nanowires an obvious choice for creating highly sensitive tools that can probe biological systems. The great characteristic of direct and label-free readout (i.e., without the use of bound dyes and fluorescent probes) is particularly attractive for many applications in medicine and life sciences.</p> | ||
| + | <p class="STYLE33">An important point about this detection process, which is quite distinct from common optically based assays, is that it occurs in real time, and the binding process can be viewed as it happens on a computer logging the conductance of one or more devices.</p> | ||
| + | <p class="STYLE33"><strong>What is MMP-7?</strong></p> | ||
| + | <p class="STYLE33">Matrilysin, also known as matrix metalloproteinase-7 (MMP-7), pump-1 protease (PUMP-1), or uterine metalloproteinase is an enzyme in humans that is encoded by the MMP-7 gene. <br /> | ||
| + | The primary role of cleaved/activated MMP-7 is to break down extracellular matrix by degrading macromolecules including casein, type I, II, IV, and V gelatins, fibronectin, and proteoglycan. Most MMP's are secreted as inactive proproteins which are activated when cleaved by extracellular proteinases. The enzyme encoded by this gene degrades proteoglycans, fibronectin, elastin and casein and differs from most MMP family members in that it lacks a conserved C-terminal protein domain.</p> | ||
| + | <p class="STYLE33">MMP-7 cleaves collagen III/IV/V/IX/X/XI and proteoglycan indicating that MMP inhibitors can potentially be used in therapies that involved in inhibition tissue degradation, remodeling, anti-angiogenesis and inhibition of tumor invasion. The upregulation of MMP-7 is associated with many malignant tumors including esophagus, stomach, colon, liver, pancreas, and renal cell carcinomas. High MMP-7 expression facilitates cancer invasion and angiogenesis by degrading extracellular matrix macromolecules and connective tissues. Theses degradations are associated with many mechanisms including embryogenesis, postpartum uterine involution, tissue repair, angiogenesis, bone remodeling, arthritis, decubitus ulcer, and tumor metastasis/invasion. Benifiting from the characteristic of MMP-7, it may be used to predict the level of cancer.</p> | ||
| + | <p class="STYLE33">Most MMP's are secreted as inactive proproteins which are activated when cleaved by extracellular proteinases. The enzyme encoded by this gene degrades proteoglycans, fibronectin, elastin and casein and differs from most MMP family members in that it lacks a conserved C-terminal protein domain.</p> | ||
| + | <p class="STYLE33"><strong>What is CheZ?</strong></p> | ||
| + | <p class="STYLE33">Bacterial chemotaxis results from the ability of flagellated bacteria to control the frequency of switching between smooth-swimming and tumbling episodes in response to changes in concentration of extracellular substances. High levels of phosphorylated CheY protein are the intracellular signal for inducing the tumbling mode of swimming. The CheZ protein has been shown to control the level of phosphorylated CheY by regulating its rate of dephosphorylation. In short, protein CheZ plays an important role in bacterial chemotaxis signal transduction pathway by accelerating the dephosphorylation of phosphorylated CheY (CheY-P).</p> | ||
| + | <p class="STYLE33"><strong class="STYLE48">References</strong></p> | ||
| + | <p class="STYLE33">[1] Blanco L P, Evans M L, Smith D R, et al. Diversity, biogenesis and function of microbial amyloids[J]. Trends in microbiology, 2012, 20(2): 66-73.</p> | ||
| + | <p class="STYLE33">[2] Barnhart M M, Chapman M R. Curli biogenesis and function[J]. Annual review of microbiology, 2006, 60: 131.</p> | ||
| + | <p class="STYLE33">[3] Chen A Y, Deng Z, Billings A N, et al. Synthesis and patterning of tunable multiscale materials with engineered cells[J]. Nature materials, 2014.</p> | ||
| + | <p class="STYLE33">[4] Daniel M C, Astruc D. Gold nanoparticles: assembly, supramolecular chemistry, quantum-size-related properties, and applications toward biology, catalysis, and nanotechnology[J]. Chemical reviews, 2004, 104(1): 293-346.</p> | ||
| + | <p class="STYLE33">[5] Patolsky F, Timko B P, Zheng G, et al. Nanowire-based nanoelectronic devices in the life sciences[J]. MRS bulletin, 2007, 32(02): 142-149.</p> | ||
| + | <p class="STYLE33">[6] Wilipedia</p> | ||
| + | <p class="STYLE33">[7] Yokoyama Y, Grünebach F, Schmidt SM, Heine A, H?ntschel M, Stevanovic S, Rammensee HG, Brossart P (2008). "Matrilysin (MMP-7) is a novel broadly expressed tumor antigen recognized by antigen-specific T cells". Clin. Cancer Res. 14 (17): 5503–11. </p> | ||
| + | <p class="STYLE33">[8] Edman K, Furber M, Hemsley P, Johansson C, Pairaudeau G, Petersen J, Stocks M, Tervo A, Ward A, Wells E, Wissler L (2011). "The discovery of MMP7 inhibitors exploiting a novel selectivity trigger". ChemMedChem 6 (5): 769–73.</p> | ||
| + | <p class="STYLE33">[9] Sanna M G, Simon M I. In vivo and in vitro characterization of Escherichia coli protein CheZ gain-and loss-of-function mutants[J]. Journal of bacteriology, 1996, 178(21): 6275-6280.</p></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#C67B14"><span class="STYLE29"><a name="part3" id="part3"></a><strong class="STYLE62">③Design</strong><br /> | ||
| + | </span> | ||
| + | <table width="100%" border="3" cellpadding="0" cellspacing="0" bordercolor="#C67B14" bgcolor="#993300"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><p class="STYLE33">To achieve the convert from bio-respond to electrical output, we designed the transmitter "transfibre", and the mechanics of which can be summarized as follow: On appearance of the substance of interest, our bacteria will inductively grow a large quantity of modified curli fibers which can adsorb nanogold. These nanowires are highly exclusive in conductivity from the culture, thus, they will conductively bridges two separated electrodes that pre-arranged in the culture and the change of current can be monitored, which present the approximate concentration of the target substance. </p> | ||
| + | <p class="STYLE33">Our design work of this bio-electrical bridge can be summarized into following parts. </p> | ||
| + | <p class="STYLE33"><strong>Part 1: the controllable expression of the curli fiber</strong> </p> | ||
| + | <p class="STYLE33">1.Background </p> | ||
| + | <p class="STYLE33">The delicate curli-assembly system in E. coli K-12 is revealed lately and chose for our "transfibre" prototype. Curli fibers, that play key role in establishment and persistence of biofilm communities, are coded by the Csg gene cluster where seven genes(CsgABCDEFG) located, and their function are listed here: <br /> | ||
| + | CsgA: At the cell surface, CsgA, The main subunit, alone soluble in vivo, that can be nucleated into an amyloid fiber by CsgB. <br /> | ||
| + | CsgB: Cell-surface-associated nucleation agent. <br /> | ||
| + | CsgC: CsgC is less well understood and it was predicted to have oxidoreductase activity. Within the periplasm, CsgC may regulate CsgG outer membrane assembly and pore activity through modification of C230 in CsgG. <br /> | ||
| + | CsgE: a small periplasmic protein, is proposed to direct CsgA to the CsgG secretion complex and helps mediate secretion specificity via an N-terminal sequence in the mature CsgA. CsgE also inhibits CsgA polymerization in vitro, and hence CsgE could be considered a CsgA-specific chaperone. <br /> | ||
| + | CsgF: CsgF is a membrane-associated protein, required for CsgB surface exposure, and is crucial for efficient CsgA polymerization. <br /> | ||
| + | CsgG: Once in the periplasm, the lipoprotein CsgG is transported by the LOL (lipoprotein outer-membrane localization) transport system to the outer membrane where it oligomerizes into a pore-like structure that is required for secretion of CsgA and CsgB into the extracellular milieu. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
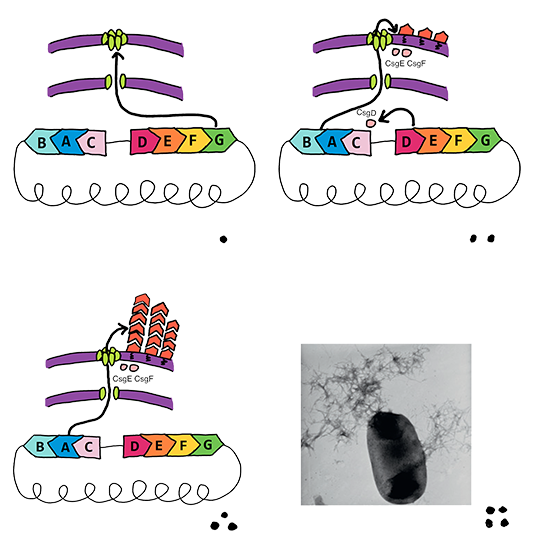
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/1/19/Df.png"><img src="https://static.igem.org/mediawiki/2014/1/19/Df.png" width="557" height="545" border="0" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 1. This figure demonstrates the cooperation of curli-associated genes when bacteria respond to external environment change.</span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33">For the reason that curli fiber mostly acts an emergency mechanism that could enhance bacterial survival in extreme conditions such as low osmolality, low growth temperature (<32°C) and hydroponic, the regulation system attached strictly limited the expression of two curli operons under normal circumstances that bio-sensing processes operating. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
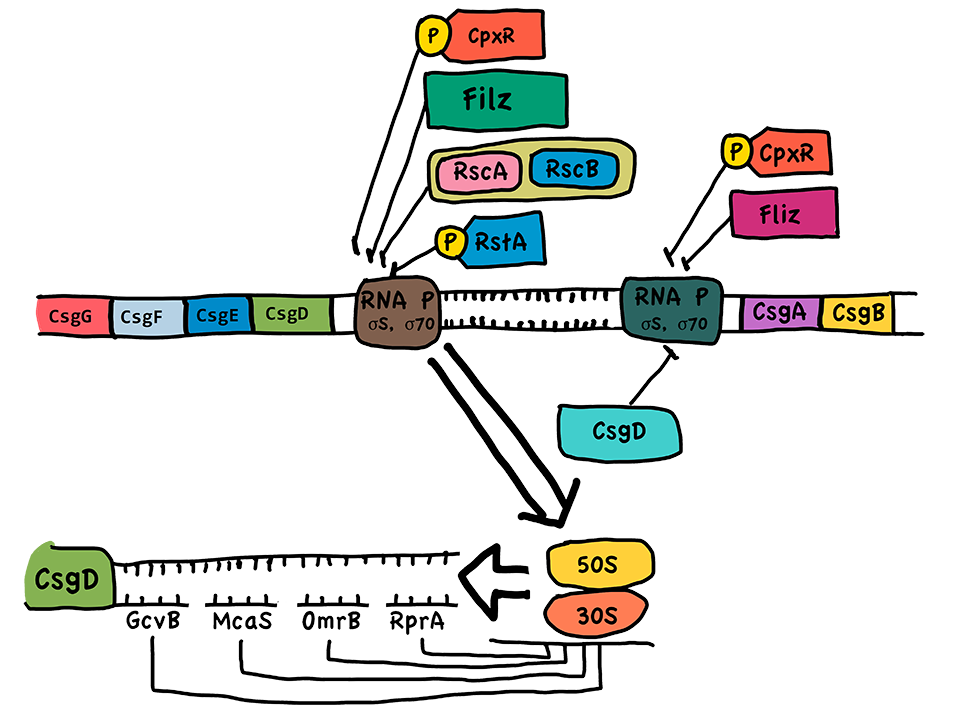
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/5/59/11.png"><img src="https://static.igem.org/mediawiki/2014/5/59/11.png" width="760" height="520" border="0" /></a></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 2. Genes that associated with curli fiber assembly are located on two operons in E coli. K-12 genome. They are under strict control and low level of expression at normal cultivation conditions. </span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33">2. The design idea <br /> | ||
| + | </p> | ||
| + | <p class="STYLE33">According to the synthetic properties of curli fiber, CsgA interacts with molecular chaperones CsgE and maintains it in a soluble state in vivo, preventing CsgA from aggregation without CsgB protein. Thus, controllable expression of CsgB can provide nucleation for CsgA rapid aggression and recruits CsgA monomers that pre-secreted in culture into fibers, which meets the request of controllable expression and immediate respond for biosensor output platform. <br /> | ||
| + | Following this strategy, we devise an inducible expression system of CsgB and a constitutive expression system of the other Csg-genes (excluding CsgD). Referring to the original curli assembly device, CsgA, the major subunit of curli fiber, and CsgC are constructed in a high copy plasmid (pSB1C3) and constitutive expression by a relatively strong promoter, while CsgEFG using a weaker promoter and docked in pSB3K3. <br /> | ||
| + | </p> | ||
| + | <p class="STYLE33">To obtain target genes from Csg operons, method PCR was applied on the genome of DH5alpha strains (The original genes of CsgA and CsgE has SpeI and PstI restriction enzyme cutting sites. They have been substituted by synonym codon according to codon usage bias). Then, construct expression vector of these genes to transform E coli. BL21 and transT-1. (In BL21, CsgA and CsgB genes has been knocked out by λ-red recombinase system to eliminate disturb.) </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/7/7c/完成版.png"><img src="https://static.igem.org/mediawiki/2014/7/7c/完成版.png" width="760" height="380" border="0" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig.3 The concept of bio-electrical transmit. The modified CsgAs are aggregate on the inductive expression of CsgB and bridges the circuit.</span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33">Two promoters, T7 promoter and Pbad promoter, are selected whose inducers, IPTG and L-arabinose respectively, are chosen to simulate the target inducing process. Meanwhile, for CsgA and CsgE, F, G, we also choose three promoters with different strength--- P1 (BBa_J23100), P2 (BBa_J23104) and P3(BBa_J23110) to screen out a better gene expression combination through this method. Thus, six parts are constructed with different levels of CsgA, C expression to work out the best fiber polymerization. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><table border="1" cellspacing="0" cellpadding="0" width="559"> | ||
| + | <tr> | ||
| + | <td width="559" colspan="2" valign="top"><p class="STYLE33">Biobricks for Promoter Strength Trail</p></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td width="253" valign="top"><p class="STYLE33">T7-CsgB-P1-CsgA, C</p></td> | ||
| + | <td width="307" valign="top"><p class="STYLE33">Pbad-CsgB-P1-CsgA, C</p></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td width="253" valign="top"><p class="STYLE33">T7-CsgB-P2-CsgA, C</p></td> | ||
| + | <td width="307" valign="top"><p class="STYLE33">Pbad-CsgB-P1-CsgA, C</p></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td width="253" valign="top"><p class="STYLE33">P2-CsgE, F, G</p></td> | ||
| + | <td width="307" valign="top"><p class="STYLE33">P3-CsgE, F, G</p></td> | ||
| + | </tr> | ||
| + | </table></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Table 1. Six parts with distinguished strength of constitutive promoters were constructed. And combinations of them were tested. </span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33">In addition, we truncate 19 amino acids from CsgB molecule’s C-terminus and get protein CsgBtrunc. It has been tested to be not outer membrane-associated, but secreted away. Soluble CsgBtrunc can also assemble into fibers that bound to the amyloid-specific dyes Congo red, and CsgBtrunc are able to seed soluble CsgA polymerization in vitro. Although it displayed only modest nucleator activity in vivo, considering that our modules could secrete much higher concentration of CsgA, it could acts relatively good nucleating capacity. </p> | ||
| + | <p align="left" class="STYLE33">The establishment of CsgBtrunc modules resembles with CsgB. The difference is that, after the addition of inducers, the curli fiber will grow suspendingly in liquid, not anchored on the outer membrane of E.coli. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><strong><a href="https://static.igem.org/mediawiki/2014/5/52/已完成2.png"><img src="https://static.igem.org/mediawiki/2014/5/52/已完成2.png" width="480" height="360" border="0" /></a></strong></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 4. This figure shows the concept of CsgBtrunc expression and off-cell nucleating on CsgA. <br /> | ||
| + | 3. An alternative expression strategy</span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p align="left" class="STYLE33">We can transform CsgA and CsgB into separate bacteria and put them in same culture. In this mode, one kind of E. coli acts as ‘material background’ that provide constitutively expressed CsgBtrunc, and another acts as ‘inducible nucleator’ which CsgA is under induction. Detection acting at a distance could be accomplished by this method. However, this method was not accomplished and we don’t know its advantages clearly. </p> | ||
| + | <p class="STYLE33"><strong>Part 2: The method of electric conduction</strong> </p> | ||
| + | <p class="STYLE33">Our curli fiber is supposed to adsorb nanogold, which is conductive and can act as wires when linked to each other. However, the wild type CsgA monomer is unable to adsorb nanogold. Thus, some measures should be taken to functionize it. <br /> | ||
| + | According to literature research, we designed three methods to achieve this goal: inserting two histidine tags (one before the first repeat domain and one after the last repeat domain in CsgA), adding a cysteine tag and inserting an amino acid sequence of pentapeptide. The principle of these three methods is the same: adding ligands like sulfhydryl that have good affinity with gold to gain CsgA ability of absorbing nanogold. <br /> | ||
| + | As for the first method, CsgA-His is constitutive expression whereas CsgB is under the control of Pbad promoter. Once we adding the inducer, CsgA would quickly aggregates and thus curli fiber with his tags would generate immediately and absorbing Au-NTA-Ni particle. We want to test the affinity between Meanwhile, the CsgA-His can be used for CsgA protein detection and purification. For the second method that adding a cysteine tag, the integral mechanism of generating the curli fibre is same as above, but the sulfhydryl of cysteine tags can link to the nanogold particles itself because the monodisperse spherical gold nanoparticles are arranged to chained structure due to the effect of these molecules. Besides, the citrate layer on the metal surface is replaced by cysteine leading to a formation of organic double layer structure, which enhances the absorption. As for the third one, the composition of the pentapeptide is Leu-Pro-Phe-Phe-Asp-OH (LPFFD-OH), the existence of N-donors containing side-chains will help a lot in the attachment or binding to the surface of gold nanoparticles, though the detailed mechanisms of the binding of the pentapeptide was not fully resolved. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><img width="760" height="560" src="https://static.igem.org/mediawiki/2014/a/ac/已完成.png" /></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 5. The modified CsgA that obtains the ability of absorbing nanogold will firm conductive wires between electrodes on the respond of inducer.</span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33">In all three methods, the primary concept is to produce conductible curli fiber and accomplish the bio-signal transformation. We have already known the modified curli fiber were produced by a certain kind of inducer (arabinose and IPTG were used in our experiments) and then attached to AuNPs. Since we assumed that the intensity of the induction could be reflected by the quantity of the AuNPs-linked curli, the concentration of the inducer could thus determine the resistance and consequent current of the whole circuits. In this way, we transformed the bio-signal into the electric signal. Furthermore, as the system is highly modularized, we can substitute the promoter of other inducers so that we could extend this system to detect sorts of subsidence apart from arabinose we used. </p> | ||
| + | <p class="STYLE33"><strong>Part 3: the upgrade of idea-logic gate.</strong> </p> | ||
| + | <p class="STYLE33">As mentioned, CsgA will aggregate into curli fibers with the seduction by seducer (like arabinose). In order to make the whole system a contenting device to characterize the concentration of seducer, a NOT gate is added to sabotage the nanowire. Thus, the current will decline and the change of concentration can still follow a kind of function, which can be determined by experiments. <br /> | ||
| + | AuNPs-based nanowire may be digested by protease or tear into pieces under the influence of physical forces. Because of the endurance of CsgA to proteases, we design a forces-destroy pathway. <br /> | ||
| + | CheY and CheZ are antagonist in a system regulating the motion and the resulting chemotaxis of E.coli: Phosphorylated CheY motivates the rolling of the bacterium, and it is damped by CheZ through prompting the dephosphorylation. <br /> | ||
| + | We construct two plasmids. One is BBa_K1361000, which has the stronger promoter P1, and the other has the weaker promoter P2. CI is a repressor gene that can restrain the expression of CheZ, and Plam acts as the promoter to start the gene of expression of cheZ. If the concentration of the inducer increases, the Pbad promotes the gene expression of CsgB as well as the repressor CI. So the effect of inhibition of CheZ increases as the curli fiber grows, because the repressor CI can bind to the Plam and inhibit the expression of CheZ. Thus, CheZ gene is depressed. If the concentration of the inducer decreases, the concentration of the repressor decreases. So the promoter Plam could start the expression of CheZ in some degree. The CheZ gene expresses and it can and, afterwards, motivate the motion of flagellum, that naturally destroyed curli fiber. So the intensity of the electric current will descend. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/7/7a/9.png"><img src="https://static.igem.org/mediawiki/2014/7/7a/9.png" width="796" height="579" border="0" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 6. The mechanic of CheZ-Curli interaction is demonstrated in the figure, hypothesizing that the increasing motility of cells will destruct the curli fiber. </span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33">Because our conductive biofilm build a bridge between bacteria and electricity, our E.coli is expected to operate machine. As the concentration of inducer increases, the circuit can be turned on, and it can switch almost any electrical devices and machines with or without the help of signal amplifier. It, which may extend the application of transfibre. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/7/7f/已完成_(2).png"><img src="https://static.igem.org/mediawiki/2014/7/7f/%E5%B7%B2%E5%AE%8C%E6%88%90_%282%29.png" width="442" height="327" border="0" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 7. The schematic diagram of coordination between bio-device and machines.</span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33"><strong>Part 4: The detection device</strong> </p> | ||
| + | <p class="STYLE33">Our living cell system can be used to create environmentally switchable conductive biofilms. We hypothesized that IPTG-or-L-arabinose-inducible production of CsgB monomers by our cells would generate extracellular amyloid fibrils that organize AuNPs into chains, and form a conductive biofilm network. <br /> | ||
| + | Engineered biofilms were grown on interdigitated electrodes deposited on glass slides, with our E.coli cultured in the presence of AuNPs and in the presence or absence of IPTG or L-arabinose inducer. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bordercolor="#FFED97" bgcolor="#FFED97"><span class="STYLE33"><img width="760" height="200" src="https://static.igem.org/mediawiki/2014/a/a9/7676.png" /></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 8. The picture shows the making process from test tube to our detection petri dish.<br /> | ||
| + | We designed the petri dish by ourselves. For the convenience of depositing electrodes, we composed a detachable culture dish. Firstly, we cut the test tube into a 3cm-long section without bottom. Secondly, we cut another test tube with longer diameter into a 4cm-long section with a bottom. The agar is utilized to attach the 3cm-long section to a glass slide to make the petri dish, and the 4cm-long section acts as a cover to ensure sterility. </span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/8/80/仪器.JPG"><img src="https://static.igem.org/mediawiki/2014/8/80/仪器.JPG" width="708" height="497" border="0" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 9. The real object of our self-made petri dish.</span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33">The fabrication of electrodes resembles the method of Synthesis and patterning of tunable multiscale materials with engineered cells. Interdigitated electrodes for measuring biofilm conductance were created by sputtering gold through shadowing masks onto glass slides. </p> | ||
| + | <p class="STYLE33">Our circuit is designed by Yuanchi Ma at College of Precision Instrument& Opto-Electronics Engineering. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/9/99/DSC00001.png"><img src="https://static.igem.org/mediawiki/2014/9/99/DSC00001.png" width="560" height="373" border="0" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"> </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33"><strong>Part 5: Stabilized matrix sensor for MMP-7</strong> </p> | ||
| + | <p class="STYLE33">The qualitative or quantitative detection of proteases can be achieved by various methods such as mass spectrum, two-dimensional gel electrophoresis and antigen-antibody complex reaction. However, they are usually time-consumed, demanding and the data is difficult to be stored and processed. Our project can transfer bio-signal to electrical signal, and as a result, we try to make it a convenient device to detect proteases without those aforementioned drawbacks. </p> | ||
| + | <p class="STYLE33">MMP-7 is the abbreviation of matrix metalloproteinase-7, which belongs to the family of MMPs (a family of extracellular zinc proteases that have long been associated with tumor invasion and metastasis and thus may serve as important cancer biomarkers). It has be proved that a helix peptide (Arg-Pro-Leu-Ala-Leu-Trp-Arg-Ser) is the substrate of MMP-7. </p> | ||
| + | <p class="STYLE33">The substrate peptide of MMP-7 and binding-site peptide of nano-gold particles (AuNPs) are modified with CsgA. These two modules added to the C-terminal of the wild type CsgA will not seriously interrupt the normal aggregation of CsgA. Consequently, we get redesigned curli fibers with the capacity to detect the MMP-7: When no MMP-7 exists in the system with applied voltage, the most electrons flows through the nanowire composed by AuNPs binding to the modified CsgAs; however, the resistance of the fibers will skyrocket for the detrimental enzymolysis of the substrate peptide by MMP-7, which result in the wreck of nanowire. Based on this blueprint, the existence of MMP-7 can be detected and the concentration can be calculated by the concentration function of current. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a name="_GoBack" id="_GoBack"><a href="https://static.igem.org/mediawiki/2014/b/b1/MMP-7插图,已完成.png"><img src="https://static.igem.org/mediawiki/2014/b/b1/MMP-7插图,已完成.png" width="790" height="359" border="0" /></a></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 10. Detection of MMP-7</span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33">Reference: <br /> | ||
| + | [1] Dobson, C.M. (2005) Structural biology: prying into prions. Nature 435,747–749 <br /> | ||
| + | [2] Collinson, S.K. et al. (1999) Structural predictions of AgfA, the insoluble fimbrial subunit of Salmonella thin aggregative fimbriae. J. Mol. Biol. 290, 741–756 <br /> | ||
| + | [3] Shewmaker, F. et al. (2009) The functional curli amyloid is not based on in-register parallel beta-sheet structure. J. Biol. Chem. 284, 25065–25076 <br /> | ||
| + | [4] Wang, X. et al. (2007) In vitro polymerization of a functional scherichia coli amyloid protein. J. Biol. Chem. 282, 3713–3719 <br /> | ||
| + | [5] Barnhart, M.M. and Chapman, M.R. (2006) Curli biogenesis and function. Annu. Rev. Microbiol. 60, 131–147 <br /> | ||
| + | [6] Wang, X. and Chapman, M.R. (2008) Curli provide the template for understanding controlled amyloid propagation. Prion 2, 57–60 <br /> | ||
| + | [7] Stathopoulos, C. et al. (2000) Secretion of virulence determinants by the general secretory pathway in Gram-negative pathogens: an evolving story. Microbes Infect. 2, 1061–1072 <br /> | ||
| + | [8] Taylor, J.D. et al. (2011) Atomic resolution insights into curli fiber biogenesis. Structure 19, 1307–1316 <br /> | ||
| + | [9] Nenninger, A.A. et al. (2011) CsgE is a curli secretion specificity factor that prevents amyloid fibre aggregation. Mol. Microbiol. 81, 486–499 <br /> | ||
| + | [10] Narita, S. et al. (2004) Lipoprotein trafficking in Escherichia coli. Arch. Microbiol. 182, 1–6 <br /> | ||
| + | [11] Epstein, E.A. et al. (2009) Spatial clustering of the curlin secretion lipoprotein requires curli fiber assembly. J. Bacteriol. 191, 608–615 <br /> | ||
| + | [12] Robinson, L.S. et al. (2006) Secretion of curli fibre subunits is mediated by the outer membrane-localized CsgG protein. Mol. Microbiol. 59, 870–881. <br /> | ||
| + | [13] Wang, X. et al. (2010) Gatekeeper residues in the major curlin subunit modulate bacterial amyloid fiber biogenesis. Proc. Natl. Acad. Sci.U.S.A. 107, 163–168 <br /> | ||
| + | [14] Majzik, A. et al. (2010). Functionalization of gold nanoparticles with amino acid, β-amyloid peptides and fragment. Colloids and Surfaces B: Biointerfaces, 81(1), 235-241. <br /> | ||
| + | [15] Liu G, Wang J, Wunschel DS, Lin Y (2006) Electrochemical proteolytic beacon for detection of matrix metalloproteinase activities. J Am Chem Soc 128(38):12382–12383. doi:10.1021/ja0626638 <br /> | ||
| + | [16] Hammer N D, Schmidt J C, Chapman M R. The curli nucleator protein, CsgB, contains an amyloidogenic domain that directs CsgA polymerization[J]. Proceedings of the National Academy of Sciences, 2007, 104(30): 12494-12499. <br /> | ||
| + | [17] Chen A Y, Deng Z, Billings A N, et al. Synthesis and patterning of tunable multiscale materials with engineered cells[J]. Nature materials, 2014. </p> | ||
| + | <span class="STYLE33">[18] Scheibel T, Parthasarathy R, Sawicki G, et al. Conducting nanowires built by controlled self-assembly of amyloid fibers and selective metal deposition[J]. Proceedings of the National Academy of Sciences, 2003, 100</span></td> | ||
| + | </tr> | ||
| + | </table></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#C67B14"><p class="STYLE62"><strong><a name="part4" id="part4"></a>④Result</strong></p></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><p class="STYLE79">Part 1. Controllable Expression of Curli fiber</p> | ||
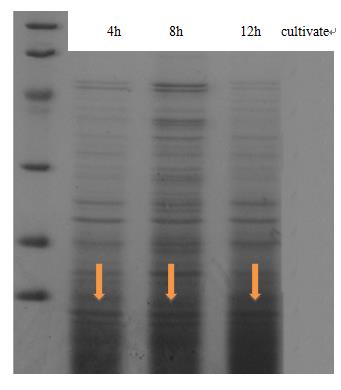
| + | <p class="STYLE33">Expression and secrete of CsgA, the major subunit of curli fiber, was carried out in <em>E coli.</em> trans-T1 transformed with CsgA, CsgB expression cassette (BBa_K1361007 docked into pSB1C3) and curli-secrete device (BBa_K1361005 docked into pSB3K3). We tracked cell growth and total quantities of secrete protein in culture. Samples were taken from culture and Bradford protein assay was adopted after centrifuging and sterilization filtered through 0.22μm filter membrane. Meanwhile, another sample taken at 2-hour interval was wall-broken and examined by SDS-PAGE, which confirm the expression of CsgA-his.</p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr bgcolor="#FFED97"> | ||
| + | <td class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/4/45/234520141017025038.jpg"><img src="https://static.igem.org/mediawiki/2014/4/45/234520141017025038.jpg" width="341" height="390" border="0" /></a></td> | ||
| + | </tr> | ||
| + | <tr bgcolor="#FFED97"> | ||
| + | <td class="STYLE33">Fig 1. The result of SDS-PAGE shows that CsgA-his has been constitutive expressed in cell. Inducating that BBa_K1361007 works as anticipation.</td> | ||
| + | </tr> | ||
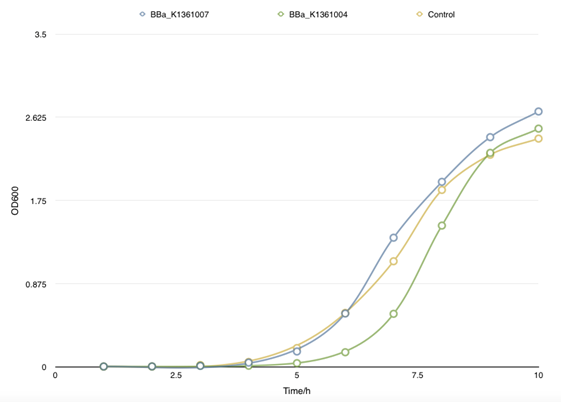
| + | </table> <p class="STYLE33">In order to find the proper promoter strength to control the concentration of CsgA monomer, Cell growth curve was drew to compare the CsgA secrete and self- aggregation level when adapted with a strong promoter BBa_J23100 and a weaker one BBa_J23104. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/4/44/Project2_clip_image002.png"><img src="https://static.igem.org/mediawiki/2014/4/44/Project2_clip_image002.png" alt="Screen Shot 2014-10-16 at 11" width="561" height="402" border="0" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 2. Cell growth curve of BBa_K1361007 and BBa_K1361004 that a relatively weaker promoter was used to control the CsgA secrete level. </span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/5/5a/Project2_clip_image004.png"><img src="https://static.igem.org/mediawiki/2014/5/5a/Project2_clip_image004.png" alt="说明: Screen Shot 2014-10-16 at 4" width="564" height="336" border="0" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
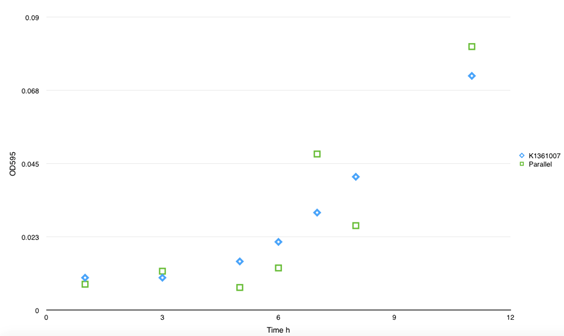
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 3. This figure shows the absorbency of secrete protein in culture between BBa_K1361007 and parallel BBa_K1361004. No significant divergence was found. And no self- aggregation was observed without inducer.</span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <p class="STYLE33"> </p> | ||
| + | <p class="STYLE33">However, there is no obvious divergence of the concentration of matrix soluble protein between control samples (with empty plasmids) and cells that had been transformed with both BBa_K1361007/BBa_K1361004 (Curli Fiber generator under the control of Pbad promoter with CsgA modified by His tag) and BBa_K1361005 (the outer membrane secrete device for curli fiber). Coupled with the failure of CsgA-his purification from cell culture, suggesting that the outer membrane channel protein CsgG and its assist proteins CsgE, F we using wasn’t functioning and may get mutation.</p> | ||
| + | <p class="STYLE33"><strong>Part 2. Congo Red (CR) Assay to Determine the Degree of Curli Fiber Polymerization</strong></p> | ||
| + | <p class="STYLE33">To examine our controlling force on curli fiber inductively aggregation, we measured the curli fiber polymerization properties after L-arabinose inducing by CR assay. The experiments showed that adding of the inducer seemed not greatly affect the curli fiber assembly. Regardless of induction, there would be decay at A480 within 10min after transferring cells into static condition.</p> | ||
| + | <p class="STYLE33">Speculating the non-function of outer membrane secrete system, we restructured the out membrane channel CsgEFG and send it to iGEM Headquarter (BBa_K1361005), while using it to process further characterization. Result of which will be demonstrated in our presentation. </p> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/8/8a/Project2_clip_image006.png"><img src="https://static.igem.org/mediawiki/2014/8/8a/Project2_clip_image006.png" width="720" height="382" hspace="12" border="0" align="left" /></a></span></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><span class="STYLE33">Fig 4. The polymerization rate of curli fiber was determined by CR assay.? After 1μM L- arabinose inducing at 0min, culture medium was subpackaged into glass tubes and cultivates under 37C, static. Samples were taken to filter and suspended in 700μl 1XPBS; 300μl of cell suspension was used for OD600 cell number normalization, and the remaining 400μl combined with 5X Congo Red (CR) for a final concentration of 20μg/ml CR, and incubated for 5min at RT. The cells and curli with bound CR were spun down at 15,000-x g for 5min, and 300μl supernatant was removed for CR quantification. Concentration of CR in supernatant was quantified by absorbance at 480nm. The amount of CR bound by cells and curli were quantified by subtracting the Ab480nm of supernatant from Ab480nm of 20μg/ml CR. </span></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <table width="100%" border="0" cellpadding="0" cellspacing="0"> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><p class="STYLE33"><a href="https://static.igem.org/mediawiki/2014/8/85/Project2_clip_image008.png"><img src="https://static.igem.org/mediawiki/2014/8/85/Project2_clip_image008.png" width="560" height="380" border="0" /></a></p></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td height="115" bgcolor="#FFED97"><p class="STYLE33">Fig 5. The adding of 1μM L- arabinose inducer at 0min didn’t significantly affect the CsgA aggregation in matrix judging by OD480. Which indicates the failure of outer membrane secrete device. </p> | ||
| + | <p class="STYLE33"> </p></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <table width="100%" border="0" cellspacing="0" cellpadding="0"> | ||
| + | <tr> | ||
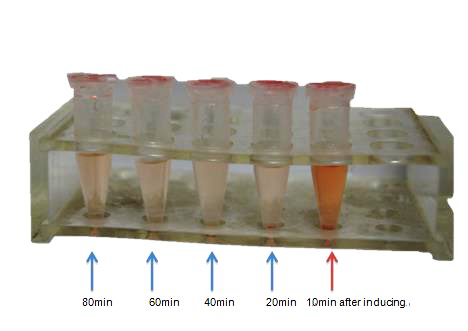
| + | <td bgcolor="#FFED97"><a href="https://static.igem.org/mediawiki/2014/1/15/2345截图20141017025120.png"><img src="https://static.igem.org/mediawiki/2014/1/15/2345截图20141017025120.png" width="468" height="331" border="0" /></a></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97" class="STYLE33">Fig 6. This pic shows the color change of CR solution after 1μM L- arabinose inducing. In the fist 20min, distinct color change was observed. Nevertheless, this could be the expression of curli fiber from <em>E coli</em> K-12’s own genome.</td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <br /></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#C67B14" class="STYLE62"><strong><a name="part5" id="part5"></a>⑤Future Work</strong></td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td bgcolor="#FFED97"><p class="STYLE33">We believe the enlightening idea of the transformation from bio-signal to electrical signal will exert far-reaching influence on the future society for the convenience and the help of highly efficient computing and analyzing capacity of computers. Our project is born ignited attributed to the belief, and as a result, we have devoted ourselves to the theories and experiments constructing to the final project. However, some work is expected to be done to perfect the project in the future due to the limited time we have during the competition.</p> | ||
| + | <p class="STYLE33">Firstly, no ideal method have been found to reverse the electroconductivity when the concentration of the seducer is lower than the threshold value. Although we have paid great effort on searching information on the problem and assumed that enzymolysis and the motivation of bacterial flagella can destroy the nanowire composed by CsgA, no further experiments can be carried on for the lack of time and grounded references.</p> | ||
| + | <p class="STYLE33">Secondly, the linker used in the experiment of the detection of MMP-7 is obtained semiempirically. Thus, this linker will potentially and probably effect the protein folding of CsgA and the substrate polypeptide, exert negative influence on the aggregation of CsgA and the enzymolysis of the polypeptide by MMP-7. Consequently, an optimal linker may be found through future theoretical calculation and experiments to lead to better results.</p> | ||
| + | <p class="STYLE33">Finally, the application of the project will be considered in the future. This project is endowed with innovative ideas and constituted of various sub-modules, but the experiments only prove that the system can be a great practical device to detect the existence of protease(MMP-7). We believe the ideas embedded in the project can ignite our ideas of the novel application of biosensors related to pharmacy, detection of organics, medicine treatment, etc.</p></td> | ||
| + | </tr> | ||
| + | </table></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <br /> | ||
| + | <table width="1230" border="5" cellpadding="0" cellspacing="0" bordercolor="#993300" bgcolor="#FFFFFF"> | ||
| + | <tr> | ||
| + | <td height="59" bgcolor="#993300"><table width="100%" border="15" cellpadding="0" cellspacing="0" bordercolor="#3A2108"> | ||
| + | <tr> | ||
| + | <td width="19%" bgcolor="#3A2108"><a href="http://www.tju.edu.cn/"><img src="https://static.igem.org/mediawiki/2014/0/01/Tianjin-school.png" width="175" height="163" border="0" /></a></td> | ||
| + | <td width="81%" bgcolor="#3A2108"><p><span class="STYLE61">Tianjin University,Tianjin, China<br /> | ||
| + | Email:michaelss@tju.edu.cn</span></p> | ||
| + | <p> </p></td> | ||
| + | </tr> | ||
| - | < | + | </table> |
| - | + | <p class="STYLE18"><br /> | |
| - | + | </p> | |
| - | + | </td> | |
| - | < | + | </tr> |
| - | < | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | </tr> | + | |
</table> | </table> | ||
| - | + | <script language="javascript"> | |
| - | < | + | var speed = 100; |
| - | + | var scrollTop = null; | |
| - | + | var hold = 0; | |
| - | + | var float_banner; | |
| - | + | var pos = null; | |
| - | + | var timer = null; | |
| - | + | var moveHeight = null; | |
| - | + | float_banner = document.getElementById("float_banner"); | |
| - | + | window.onscroll=scroll_ad; | |
| - | + | function scroll_ad(){ | |
| - | + | scrollTop = document.documentElement.scrollTop+document.body.scrollTop; | |
| - | + | pos = scrollTop - float_banner.offsetTop; | |
| - | + | pos = pos/10 | |
| - | + | moveHeight = pos>0?Math.ceil(pos):Math.floor(pos); | |
| - | + | if(moveHeight!=0){ | |
| - | + | float_banner.style.top = float_banner.offsetTop+moveHeight+"px"; | |
| - | + | setTimeout(scroll_ad,speed); | |
| - | + | } | |
| - | + | //alert(scrollTop); | |
| - | + | } | |
| - | + | </script> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</p> | </p> | ||
| + | </body> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
</html> | </html> | ||
Latest revision as of 21:34, 17 October 2014
|
|
Welcome to Team Tianjin! |
 |
|
 "
"