Team:Oxford/Parts
From 2014.igem.org
(diff) ← Older revision | Latest revision (diff) | Newer revision → (diff)

Oxford iGEM has submitted three parts to the registry, which are described on the relevant registry pages.
Part no. 1 (BBa_K1446001) is the gene coding for DcmA with an N-terminal pdu-microcompartment tag.

Part no. 2 (BBa_K1446002) is a superfolder GFP-gene with the N-terminal pdu-microcompartment tag. We have included a ribosome binding site upstream of the coding region as well as a his-tag at the C-terminus. This part has been used for fluorescent image analysis to show that the targeting mechanism of the N-terminal microcompartment tag works as expected.

Part no. 3 (BBa_K1446003) is the dcmR gene with a tetR promoter. We have used this part to get fluorescence data for our model described here and showed that mCherry expression increased at higher ATC (anhydrous tetracycline) concentration, as expected.

Parts no. 1 and 2 belong to our bioremediation subproject and part 3 to our biosensor subproject.
Oxford iGEM have also constructed further parts but were not fully in a BioBrick compatible plasmid thus were not sent before the deadline but will be on the registry in the near future
The following two constructs are based on the intergenic region shown below from the native DCM-degrading bacterium (Methylobacterium extorquens DM4):
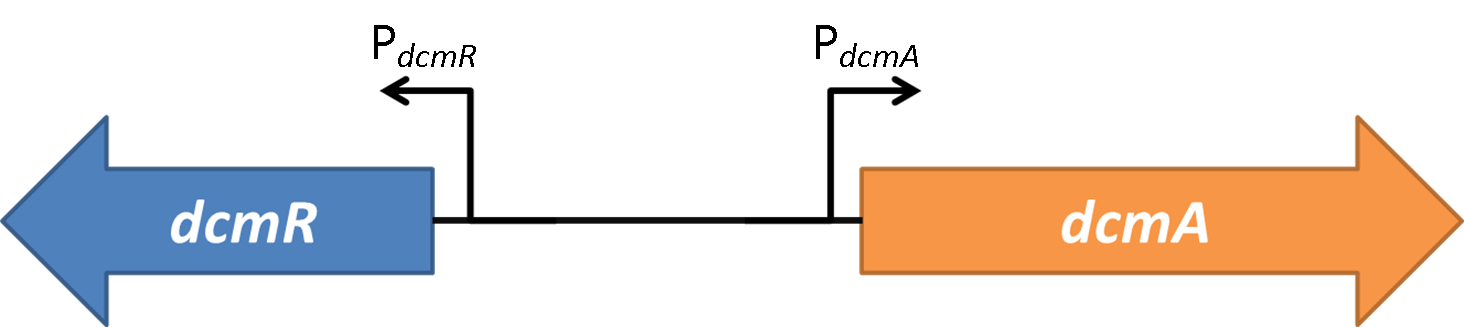
PdcmA-sfGFP

This construct involves the bidirectional promoter that has been characterised as a new tool for molecular biologists. In this construct sfGFP is placed under control in the PdcmA direction. Click here to see characterisation on this part in plasmid PJ404.
PdcmR-sfGFP
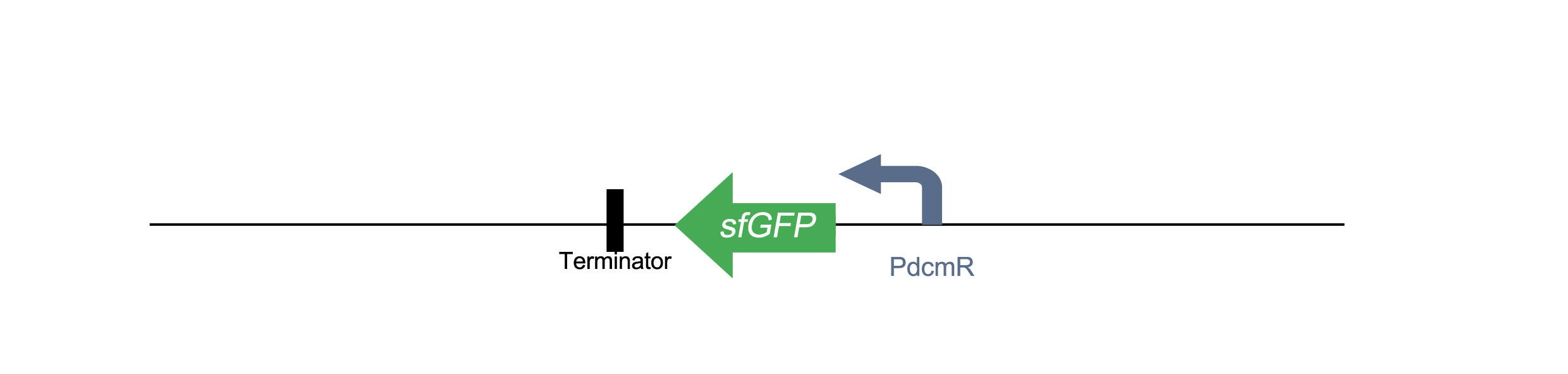
Similar to the construct above, this involves the same bidirectional promoter with sfGFP in placed in the opposite direction under control of PdcmR. Click here to see characterisation on this part in plasmid PJ404.
dcmA-sfGFP

For the analysis of dcmA-expression we have used dcmA-fused to sfGFP in the pRSFDuet vector, which has a T7-promoter that is under lac-control.



































Retrieved from "http://2014.igem.org/Team:Oxford/Parts"
 "
"




















