|
|
| (15 intermediate revisions not shown) |
| Line 2: |
Line 2: |
| | <html> | | <html> |
| | <div id="main-content"> | | <div id="main-content"> |
| - | <h>Lactobacillus Casei</h> | + | <h>Probiotic-<i>Lactobacillus casei</i></h> |
| - | | + | <h1>Summary</h1> |
| | + | <div class='abstract'> |
| | + | <ul> |
| | + | <li>Recombinant <i>Lactobacilli</i> are developed to serve as a bio-safely organism, in the hope to express heterologous proteins in oral therapeutic applications.</li> |
| | + | <li>In our project, caries nemesis "HOPE", we use <i>Lactobacillus casei</i> as our final host of the recombinant plasmid, which contains the promoters and the desired genes like YebF and Orf19. Orf19 and YebF are signal peptide and lysing protein which respectively derive from M102 phage and <i>E. coli</i> MG1655. Also, low oxygen and acid environment were set as the initial condition to turn on the gene promoters of our recombinant plasmid. We therefore gain an access to do conditioned expression of killing and anti-biofilm genes in <i>L. casei</i>.</li> |
| | + | <li>However, the isolation of plasmid DNA from G+ lactobacilli is complicated by resilience of the peptidoglican layer.</li> |
| | + | <li>For this reason, it is a relatively feasible and reliable way that we use <i>E. coli</i> as our testing bacterium of “HOPE” but the functional promoters and genes of the shuttle vector will be eventually transformed into our probiotic <i>L. casei</i>, which is safe to use as a food-grade protein expression and secreting system.</li> |
| | + | </ul> |
| | + | </div> |
| | + | <div class='cont-panel'> |
| | + | <div href='#lc-1'><p>Get started.</p></div> |
| | + | <div href='#lc-2'><p>How to do it?</p></div> |
| | + | <div href='#lc-3'><p>Expected results~</p></div> |
| | + | <div href='#lc-4'><p>Reference</p></div> |
| | + | <div style="display:inline-block; width: 640px; height: 0.1px; border: none; margin: 0px"></div> |
| | + | </div> |
| | + | <div class='article indent'> |
| | + | <h1 id='lc-1'>Before we get started:</h1> |
| | + | <div style='float: right;padding-left: 20px;padding-top: 10px;'><img src='/wiki/images/c/c6/NYMU14_lcasei_p1.jpg' style='width: 350px;'><p style='font-size: 14px;'>The Gram positive <b><i>L. casei.</i></b>, LB23</p></div> |
| | + | <p>The purpose of our study attempts to construct an <i>E. coli-Lactobacillus</i> shuttle vector plasmid with an expression cassette consisting of a strong promoter, a signal sequence, and a heterologous model protein gene. We put the plasmid in<i >E. coli</i>, our testing bacterium to conform the feasibility of our study and finally the plasmid will be inserted into <i>L. casei</i> in "HOPE".</p> |
| | + | <p>Generally, lactic acid bacteria (LAB) are regarded as safe and useful commensal bacteria, which are known as probiotics and starters for food fermentation. In recent years there has been an upsurge of interest in the genetic manipulation of lactic acid bacteria (LAB).</p> |
| | + | <p>Many studies on <i>Lactobacillus casei</i> subsp. have been carried out using the plasmid-free strain BL23, ATCC 393 (pLZ15-)[4]. BL23 and its relatives have been used as a host for genetic modification and applied for live delivery agents. For example, the single chain variable fragment of an antibody that binds to a major adhesion molecule of <i>Streptococcus mutants</i> was expressed on the cell surface and showed protective effects against colonization with the pathogen [5]. Thus, it is clear that the development of a highly efficient protein-secreting system in this strain would be important and beneficial.</p> |
| | + | <p>Moreover, <i>L. casei</i> might be a better live delivery agent because the optimal growth temperature of <i>L. casei</i> is the same as the body temperature of mammals. Thus, BL23 is no doubt a suitable bacterial host for our food-grade protein secreting system in “HOPE”.</p> |
| | + | |
| | + | <h1 id='lc-2'>So how did we do it?</h1> |
| | + | <h3>Bacterial Strains and Growth Conditions</h3> |
| | + | <p><i>L. casei</i> BL23 and recombinant strains were grown in Mann-Rogosa-Sharp (MRS) medium at 37℃. </p> |
| | + | <p>Erythromycin (5 μg/ml) was added to MRS for the selection of recombinants <i>L. casei</i> strains. For pH control, 50 mM carbonate buffer was supplemented into the medium at certain ratios (NaHCO3:Na2CO3). The commonly used cloning host of <i>E. coli</i> DH5α (TaKaRa, Shiga, Japan) was grown in LB with or without 100 μg/ml ampicillin. <i>L casei</i>, BL 23 were maintained in a 10% glycerol stock collection at -80 °C and were subcultured in MRS medium (Oxoid) when required.</p> |
| | + | <p>Bacterial strains used in this study:</p> |
| | + | <pre> |
| | + | Strain code | Strain | Source |
| | + | BL23 | <i>L. casei ATCC 393</i> (pLZ15-) | [Dr. Ying-Chieh Tsai] of NYMU, Taiwan |
| | + | </pre> |
| | + | |
| | + | <h3>Construction of Plasmids and Transformation of <bi><i>L. casei</i></ib></h3> |
| | + | <p>The sequences of all PCR primers used in this study are listed in the following Table 2. PCR products and plasmids were digested with restriction endonucleases followed by ligation and transformation to <i> E. coli DH5α</i>. In the cloning of <i> E. coli DH5α</i> we used a high copy number plasmid named <i>E. coli</i>, pUC19 After harvesting <i>L. casei</i> plasmid by digesting pUC19, we use a E-coli-Lactobacillus shuttle vector, pLP402, to perform our desired features. And the transfer process was carried out by electrophoresis. </p> |
| | + | <p>PCR primers used in this study:</p> |
| | + | <pre> |
| | + | Primers Sequences Restriction sites |
| | + | IGM289 cccaagcttagatctgattacaaaggctttaagcagg HindIII, BglII |
| | + | IGM290 gggctcgaggcccggttgttcgcggccgcttttgttaagaattttatttcataacattagcgg XhoI, NotI |
| | + | IGM291 cccgtcgacgcccggttgttcgcggccgcttcggttatactattcttgcttgata SalI, NotI |
| | + | IGM292 ggggaattcctgcagggatccaaacttgattgcataatctttcttcc BamHI, PstI, EcoRI |
| | + | IGM350 acatattttatgtttggagggtattggatg |
| | + | IGM351 catccaataccctccaaacataaaatatgt |
| | + | IGM468 ccccggatccgagcaggggccagtacagccg BamHI |
| | + | IGM478 ccccctcgagttagcttttcattttgatcatcatgta XhoI |
| | + | IGM479 gcgaaatccaagcaaaggcgagcaggggccagtacagccg |
| | + | IGM480 cggctgtactggcccctgctcgcctttgcttggatttcgc |
| | + | IGM482 cggctgtactggcccctgctcggatccgagtttgtgtccgcctttgcttggatttcgc BamHI |
| | + | </pre> |
| | + | |
| | + | <h1 id='lc-3'>Results~</h1> |
| | + | <p>Construction-flow diagram of expression vectors. MCS: multiple cloning site; Rep: replication protein; Emr: erythromycin resistance; Ampr: ampicillin resistance</p> |
| | + | <img src='/wiki/images/7/76/NYMU14_lcasei_res1.jpg' style='width: 650px;'> |
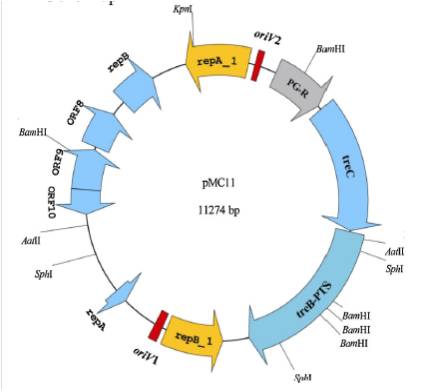
| | + | <p>Gene map of plasmid pMC11 and putative replicons, ORFs.RepA_1 and RepB_1 are replication initiation proteins; RepA and RepB are putative replication proteins. For trehalose operon, PG-R is trehalose operon repressor, TreC is trehalose-6-phosphate hydrolase and TreB-PTS is PTS system trehalose-specific IIBCA components.</p> |
| | + | <img src='/wiki/images/2/27/NYMU14_lcasei_res2.jpg'> |
| | + | <p>C, D Replication origins fragment of pMC11. Repeats of oriV1 (iterons contain 4.5 direct repeats) |
| | + | and oriV2 (iterons contain 3.5 direct repeats) and invert repeat are diagrammed. Promoter elements and ribosome binding sites (RBS) of repA_1 and repB_1 genes are indicated.</p> |
| | + | <img src='/wiki/images/2/21/NYMU14_lcasei_res3.jpg'> |
| | + | |
| | + | </div> <!-- article --> |
| | + | <h1 id='lc-4'>Reference</h1> |
| | + | <ol> |
| | + | <li>Steidler, L., W. Hans, L. Schotte, S. Neirynck, F. Obermeier, W. Falk, W. Fiers, and E. Remaut. 2000. Treatment of murine colitis by Lactococcus lactis secreting interleukin-10. Science 289: 1352-1355.</li> |
| | + | <li>Braat, H., P. Rottiers, D. W. Hommes, N. Huyghebaert, E. Remaut, J. P. Remon, et al. 2006. A phase I trial with transgenic bacteria expressing interleukin-10 in Crohn’s disease. Clin. Gastroenterol. Hepatol. 4: 754-759.</li> |
| | + | <li>Pusch, O., D. Boden, S. Hannify, F. Lee, L. D. Tucker, M. R. Boyd, J. M. Wells, and B. Ramratnam. 2005. Bioengineering lactic acid bacteria to secrete the HIV-1 virucide cyanovirin. J. Acquir. Immune Defic. Syndr. 40: 512-220.</li> |
| | + | <li>Acedo-Félix, E. and G. Pérez-Martínez. 2003. Significant differences between Lactobacillus casei subsp. casei ATCC 393T and a commonly used plasmid-cured derivative revealed by a polyphasic study. Int. J. Syst. Evol. Microbiol. 53: 67-75.</li> |
| | + | <li>Krüger, C., Y. Hu, Q. Pan, H. Marcotte, A. Hultberg, D. Delwar, et al. 2002. In situ delivery of passive immunity by lactobacilli producing single-chain antibodies. Nat. Biotechnol. 20: 702-706.</li> |
| | + | <li>Kajikawa, Akinobu1, Eiko Ichikawa2, and Shizunobu Igimi2 2010. Development of a Highly Efficient Protein-Secreting System in Recombinant Lactobacillus casei. J. Microbiol. Biotechnol. (2010), 20(2), 375–382</li> |
| | + | <li>Zhengjun Chen & Jinzhong Lin & Chengjie Ma & Shumiao Zhao & Qunxin She & Yunxiang Liang Characterization of pMC11, a plasmid with dual origins of replication isolated from Lactobacillus casei MCJ and construction of shuttle vectors with each replicon. Appl Microbiol Biotechnol (2014) 98:5977–5989</li> |
| | + | <li>Evelia Acedo-Fe´ lix3 and Gaspar Pe´ rez-Martı´nez. Significant differences between Lactobacillus casei subsp. casei ATCC 393T and a commonly used plasmid-cured derivative revealed by a polyphasic study. International Journal of Systematic and Evolutionary Microbiology (2003), 53, 67–75</li> |
| | + | </ol> |
| | </div> | | </div> |
| | </html> | | </html> |
| | {{:Team:NYMU-Taipei/NYMU14_Footer}} | | {{:Team:NYMU-Taipei/NYMU14_Footer}} |
Probiotic-Lactobacillus casei
Summary
- Recombinant Lactobacilli are developed to serve as a bio-safely organism, in the hope to express heterologous proteins in oral therapeutic applications.
- In our project, caries nemesis "HOPE", we use Lactobacillus casei as our final host of the recombinant plasmid, which contains the promoters and the desired genes like YebF and Orf19. Orf19 and YebF are signal peptide and lysing protein which respectively derive from M102 phage and E. coli MG1655. Also, low oxygen and acid environment were set as the initial condition to turn on the gene promoters of our recombinant plasmid. We therefore gain an access to do conditioned expression of killing and anti-biofilm genes in L. casei.
- However, the isolation of plasmid DNA from G+ lactobacilli is complicated by resilience of the peptidoglican layer.
- For this reason, it is a relatively feasible and reliable way that we use E. coli as our testing bacterium of “HOPE” but the functional promoters and genes of the shuttle vector will be eventually transformed into our probiotic L. casei, which is safe to use as a food-grade protein expression and secreting system.
Before we get started:
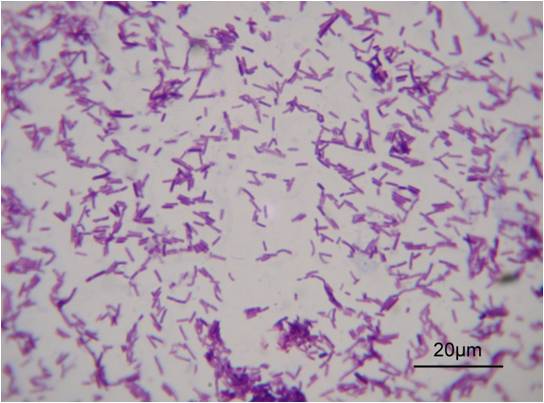
The Gram positive L. casei., LB23
The purpose of our study attempts to construct an E. coli-Lactobacillus shuttle vector plasmid with an expression cassette consisting of a strong promoter, a signal sequence, and a heterologous model protein gene. We put the plasmid inE. coli, our testing bacterium to conform the feasibility of our study and finally the plasmid will be inserted into L. casei in "HOPE".
Generally, lactic acid bacteria (LAB) are regarded as safe and useful commensal bacteria, which are known as probiotics and starters for food fermentation. In recent years there has been an upsurge of interest in the genetic manipulation of lactic acid bacteria (LAB).
Many studies on Lactobacillus casei subsp. have been carried out using the plasmid-free strain BL23, ATCC 393 (pLZ15-)[4]. BL23 and its relatives have been used as a host for genetic modification and applied for live delivery agents. For example, the single chain variable fragment of an antibody that binds to a major adhesion molecule of Streptococcus mutants was expressed on the cell surface and showed protective effects against colonization with the pathogen [5]. Thus, it is clear that the development of a highly efficient protein-secreting system in this strain would be important and beneficial.
Moreover, L. casei might be a better live delivery agent because the optimal growth temperature of L. casei is the same as the body temperature of mammals. Thus, BL23 is no doubt a suitable bacterial host for our food-grade protein secreting system in “HOPE”.
So how did we do it?
Bacterial Strains and Growth Conditions
L. casei BL23 and recombinant strains were grown in Mann-Rogosa-Sharp (MRS) medium at 37℃.
Erythromycin (5 μg/ml) was added to MRS for the selection of recombinants L. casei strains. For pH control, 50 mM carbonate buffer was supplemented into the medium at certain ratios (NaHCO3:Na2CO3). The commonly used cloning host of E. coli DH5α (TaKaRa, Shiga, Japan) was grown in LB with or without 100 μg/ml ampicillin. L casei, BL 23 were maintained in a 10% glycerol stock collection at -80 °C and were subcultured in MRS medium (Oxoid) when required.
Bacterial strains used in this study:
Strain code | Strain | Source
BL23 | L. casei ATCC 393 (pLZ15-) | [Dr. Ying-Chieh Tsai] of NYMU, Taiwan
Construction of Plasmids and Transformation of L. casei
The sequences of all PCR primers used in this study are listed in the following Table 2. PCR products and plasmids were digested with restriction endonucleases followed by ligation and transformation to E. coli DH5α. In the cloning of E. coli DH5α we used a high copy number plasmid named E. coli, pUC19 After harvesting L. casei plasmid by digesting pUC19, we use a E-coli-Lactobacillus shuttle vector, pLP402, to perform our desired features. And the transfer process was carried out by electrophoresis.
PCR primers used in this study:
Primers Sequences Restriction sites
IGM289 cccaagcttagatctgattacaaaggctttaagcagg HindIII, BglII
IGM290 gggctcgaggcccggttgttcgcggccgcttttgttaagaattttatttcataacattagcgg XhoI, NotI
IGM291 cccgtcgacgcccggttgttcgcggccgcttcggttatactattcttgcttgata SalI, NotI
IGM292 ggggaattcctgcagggatccaaacttgattgcataatctttcttcc BamHI, PstI, EcoRI
IGM350 acatattttatgtttggagggtattggatg
IGM351 catccaataccctccaaacataaaatatgt
IGM468 ccccggatccgagcaggggccagtacagccg BamHI
IGM478 ccccctcgagttagcttttcattttgatcatcatgta XhoI
IGM479 gcgaaatccaagcaaaggcgagcaggggccagtacagccg
IGM480 cggctgtactggcccctgctcgcctttgcttggatttcgc
IGM482 cggctgtactggcccctgctcggatccgagtttgtgtccgcctttgcttggatttcgc BamHI
Results~
Construction-flow diagram of expression vectors. MCS: multiple cloning site; Rep: replication protein; Emr: erythromycin resistance; Ampr: ampicillin resistance

Gene map of plasmid pMC11 and putative replicons, ORFs.RepA_1 and RepB_1 are replication initiation proteins; RepA and RepB are putative replication proteins. For trehalose operon, PG-R is trehalose operon repressor, TreC is trehalose-6-phosphate hydrolase and TreB-PTS is PTS system trehalose-specific IIBCA components.
C, D Replication origins fragment of pMC11. Repeats of oriV1 (iterons contain 4.5 direct repeats)
and oriV2 (iterons contain 3.5 direct repeats) and invert repeat are diagrammed. Promoter elements and ribosome binding sites (RBS) of repA_1 and repB_1 genes are indicated.

Reference
- Steidler, L., W. Hans, L. Schotte, S. Neirynck, F. Obermeier, W. Falk, W. Fiers, and E. Remaut. 2000. Treatment of murine colitis by Lactococcus lactis secreting interleukin-10. Science 289: 1352-1355.
- Braat, H., P. Rottiers, D. W. Hommes, N. Huyghebaert, E. Remaut, J. P. Remon, et al. 2006. A phase I trial with transgenic bacteria expressing interleukin-10 in Crohn’s disease. Clin. Gastroenterol. Hepatol. 4: 754-759.
- Pusch, O., D. Boden, S. Hannify, F. Lee, L. D. Tucker, M. R. Boyd, J. M. Wells, and B. Ramratnam. 2005. Bioengineering lactic acid bacteria to secrete the HIV-1 virucide cyanovirin. J. Acquir. Immune Defic. Syndr. 40: 512-220.
- Acedo-Félix, E. and G. Pérez-Martínez. 2003. Significant differences between Lactobacillus casei subsp. casei ATCC 393T and a commonly used plasmid-cured derivative revealed by a polyphasic study. Int. J. Syst. Evol. Microbiol. 53: 67-75.
- Krüger, C., Y. Hu, Q. Pan, H. Marcotte, A. Hultberg, D. Delwar, et al. 2002. In situ delivery of passive immunity by lactobacilli producing single-chain antibodies. Nat. Biotechnol. 20: 702-706.
- Kajikawa, Akinobu1, Eiko Ichikawa2, and Shizunobu Igimi2 2010. Development of a Highly Efficient Protein-Secreting System in Recombinant Lactobacillus casei. J. Microbiol. Biotechnol. (2010), 20(2), 375–382
- Zhengjun Chen & Jinzhong Lin & Chengjie Ma & Shumiao Zhao & Qunxin She & Yunxiang Liang Characterization of pMC11, a plasmid with dual origins of replication isolated from Lactobacillus casei MCJ and construction of shuttle vectors with each replicon. Appl Microbiol Biotechnol (2014) 98:5977–5989
- Evelia Acedo-Fe´ lix3 and Gaspar Pe´ rez-Martı´nez. Significant differences between Lactobacillus casei subsp. casei ATCC 393T and a commonly used plasmid-cured derivative revealed by a polyphasic study. International Journal of Systematic and Evolutionary Microbiology (2003), 53, 67–75




 "
"