Team:ETH Zurich/labblog/20140615mod
From 2014.igem.org
(→Sunday, June 15th) |
|||
| Line 1: | Line 1: | ||
| - | <html><article class=" | + | <html><article class="modeling" date="20140615"></html> |
== Modularization of the system == | == Modularization of the system == | ||
Revision as of 23:38, 7 October 2014
Modularization of the system
Sunday, June 15th
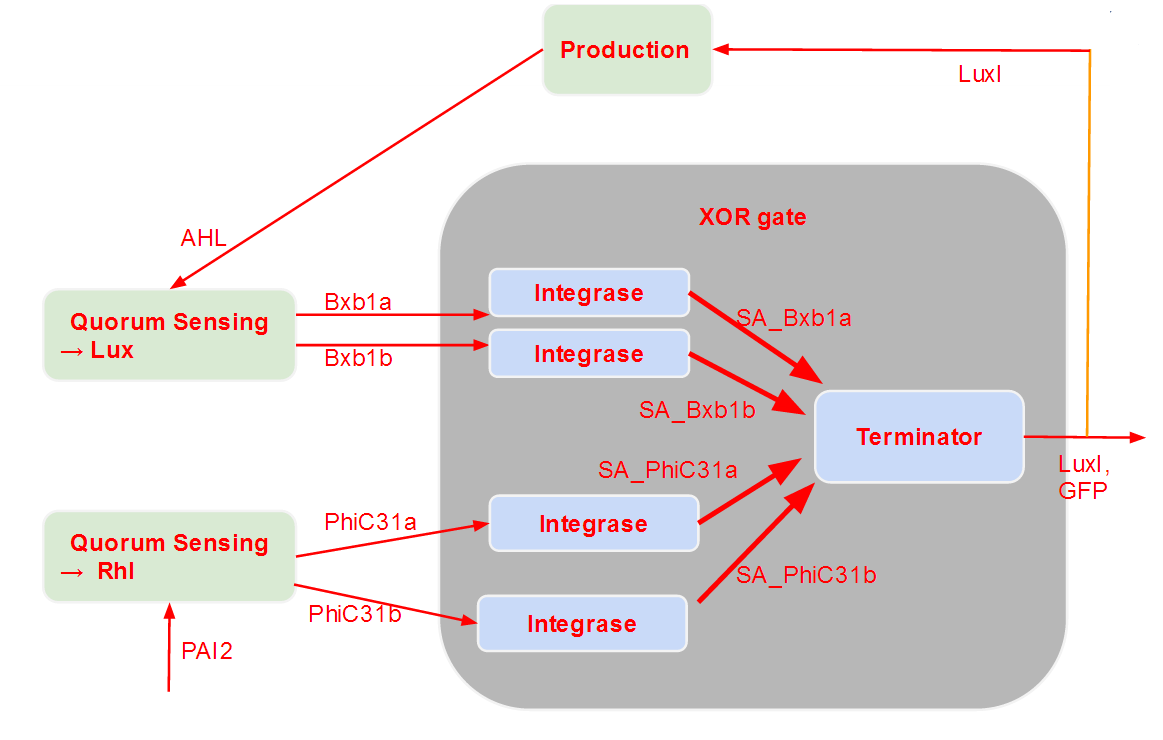
The idea of our system is to associate a quorum sensing part, which should be able to react to a certain concentration of input, with a integrase-base logic part, which should be able to derive a computation given the transformed inputs.
From a modeling point of view, it seemed obvious that we had to consider these two entities separately. That is why we chose to have a modularized approach, where we regrouped, on one hand, the equations corresponding to quorum sensing and, on the other hand, the ones corresponding to the integrase logic phenomena.
As the integrase module is not that well known, we further decomposed it into two successive parts. The first one only models the path integrases go through in order to bind to DNA. The second one models the switch behavior of the integrases (after they are bound to DNA) and describe the XOR logic of our system.
As we see, these modules are successive. After the logic module, we added a production module, which is responsible for the production of our outputs (GFP and an Homoserine Lactone that depends on the type of cell modeled). As one of our output is a messenger molecule, it has an influence on the quorum sensing module, which detects those molecules. Therefore, we have a feedback loop. This feedback loop comes from our choice to put only two different strains in our grid.
 "
"