Team:ETH Zurich/lab/sequences
From 2014.igem.org
(→Combined Sensor and Producer Constructs) |
(→Sensor Constructs) |
||
| Line 143: | Line 143: | ||
[https://static.igem.org/mediawiki/2014/d/da/ETH2014_piG0065_sensor_pluxRRR12y-sfGFP.txt piG0065] | [https://static.igem.org/mediawiki/2014/d/da/ETH2014_piG0065_sensor_pluxRRR12y-sfGFP.txt piG0065] | ||
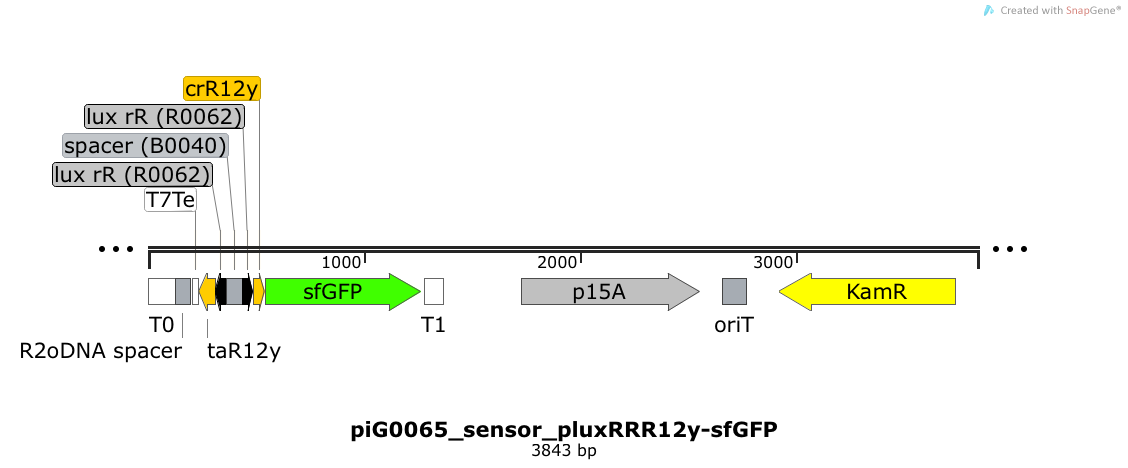
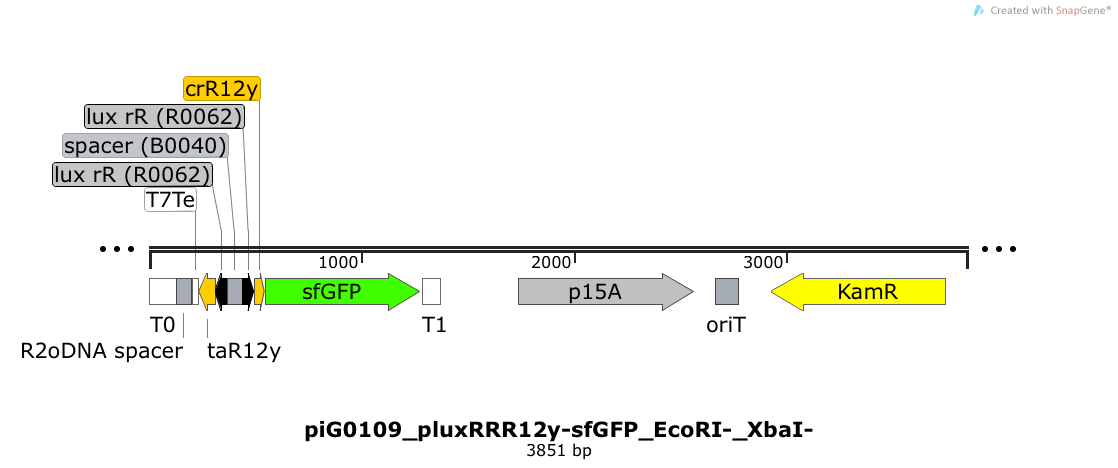
| - | Expression of sfGFP is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a [https://2014.igem.org/Team:ETH_Zurich/expresults#Riboregulators riboregulator] that decreases leakiness of pLuxR. The p15A is present at a copy number of approximately 15 to 25<sup>[[Team:ETH_Zurich/project/references#refChang|[35]]]</sup>. | + | Expression of sfGFP is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a [https://2014.igem.org/Team:ETH_Zurich/expresults#Riboregulators riboregulator] that decreases leakiness of pLuxR. A biologically neutral spacer sequence was designed using the web application R2oDNA<sup>[[Team:ETH_Zurich/project/references#refR2oDNA|[34]]]</sup>. The p15A is present at a copy number of approximately 15 to 25<sup>[[Team:ETH_Zurich/project/references#refChang|[35]]]</sup>. |
[[File:ETH2014 piG0065 sensor pluxRRR12y-sfGFP Map.png|link=https://static.igem.org/mediawiki/2014/d/da/ETH2014_piG0065_sensor_pluxRRR12y-sfGFP.txt]] | [[File:ETH2014 piG0065 sensor pluxRRR12y-sfGFP Map.png|link=https://static.igem.org/mediawiki/2014/d/da/ETH2014_piG0065_sensor_pluxRRR12y-sfGFP.txt]] | ||
| Line 149: | Line 149: | ||
[https://static.igem.org/mediawiki/2014/2/24/ETH2014_piG0066_sensor_prhlRRR12-sfGFP.txt piG0066] | [https://static.igem.org/mediawiki/2014/2/24/ETH2014_piG0066_sensor_prhlRRR12-sfGFP.txt piG0066] | ||
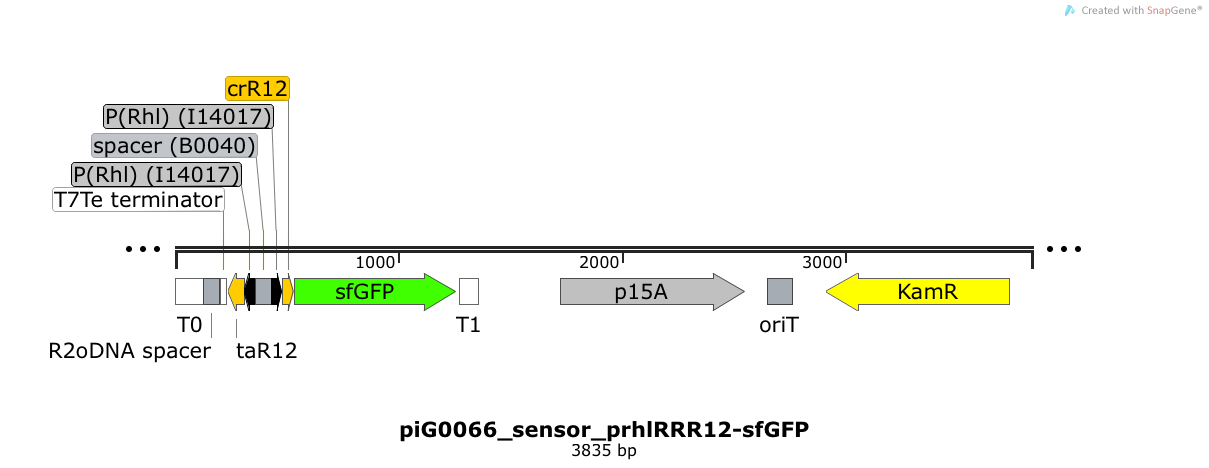
| - | Expression of sfGFP is induced when RhlR bound to 4C-HSL bind to pRhlR. The cis-repressive element (crR12) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a [https://2014.igem.org/Team:ETH_Zurich/expresults#Riboregulators riboregulator] that decreases leakiness of pLuxR. The p15A is present at a copy number of approximately 15 to 25<sup>[[Team:ETH_Zurich/project/references#refChang|[35]]]</sup>. | + | Expression of sfGFP is induced when RhlR bound to 4C-HSL bind to pRhlR. The cis-repressive element (crR12) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a [https://2014.igem.org/Team:ETH_Zurich/expresults#Riboregulators riboregulator] that decreases leakiness of pLuxR. A biologically neutral spacer sequence was designed using the web application R2oDNA<sup>[[Team:ETH_Zurich/project/references#refR2oDNA|[34]]]</sup>. The p15A is present at a copy number of approximately 15 to 25<sup>[[Team:ETH_Zurich/project/references#refChang|[35]]]</sup>. |
[[File:ETH2014 piG0066 sensor prhlRRR12-sfGFP Map.png|link=https://static.igem.org/mediawiki/2014/2/24/ETH2014_piG0066_sensor_prhlRRR12-sfGFP.txt]] | [[File:ETH2014 piG0066 sensor prhlRRR12-sfGFP Map.png|link=https://static.igem.org/mediawiki/2014/2/24/ETH2014_piG0066_sensor_prhlRRR12-sfGFP.txt]] | ||
| Line 155: | Line 155: | ||
[https://static.igem.org/mediawiki/2014/9/95/ETH2014_piG0109_pluxRRR12y-sfGFP_EcoRI-_XbaI-.txt piG0109] | [https://static.igem.org/mediawiki/2014/9/95/ETH2014_piG0109_pluxRRR12y-sfGFP_EcoRI-_XbaI-.txt piG0109] | ||
| - | Expression of sfGFP is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a [https://2014.igem.org/Team:ETH_Zurich/expresults#Riboregulators riboregulator] that decreases leakiness of pLuxR. The p15A is present at a copy number of approximately 15 to 25<sup>[[Team:ETH_Zurich/project/references#refChang|[35]]]</sup>. PiG0109 is a derivate of piG0065 where the restriction sites EcoRI and XbaI have been removed. Thus, the two constructs slightly differ in the sequence of the 3'-end of the trans-activating element and in the sequence of the 5'-end of the cis-repressive element. | + | Expression of sfGFP is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a [https://2014.igem.org/Team:ETH_Zurich/expresults#Riboregulators riboregulator] that decreases leakiness of pLuxR. A biologically neutral spacer sequence was designed using the web application R2oDNA<sup>[[Team:ETH_Zurich/project/references#refR2oDNA|[34]]]</sup>. The p15A is present at a copy number of approximately 15 to 25<sup>[[Team:ETH_Zurich/project/references#refChang|[35]]]</sup>. PiG0109 is a derivate of piG0065 where the restriction sites EcoRI and XbaI have been removed. Thus, the two constructs slightly differ in the sequence of the 3'-end of the trans-activating element and in the sequence of the 5'-end of the cis-repressive element. |
[[File:ETH2014 piG0109 pluxRRR12y-sfGFP EcoRI- XbaI- Map.png|link=https://static.igem.org/mediawiki/2014/9/95/ETH2014_piG0109_pluxRRR12y-sfGFP_EcoRI-_XbaI-.txt]] | [[File:ETH2014 piG0109 pluxRRR12y-sfGFP EcoRI- XbaI- Map.png|link=https://static.igem.org/mediawiki/2014/9/95/ETH2014_piG0109_pluxRRR12y-sfGFP_EcoRI-_XbaI-.txt]] | ||
| Line 161: | Line 161: | ||
[https://static.igem.org/mediawiki/2014/f/f9/ETH2014_piG0110_prhlRRR12-sfGFP_EcoRI-_XbaI-.txt piG0110] | [https://static.igem.org/mediawiki/2014/f/f9/ETH2014_piG0110_prhlRRR12-sfGFP_EcoRI-_XbaI-.txt piG0110] | ||
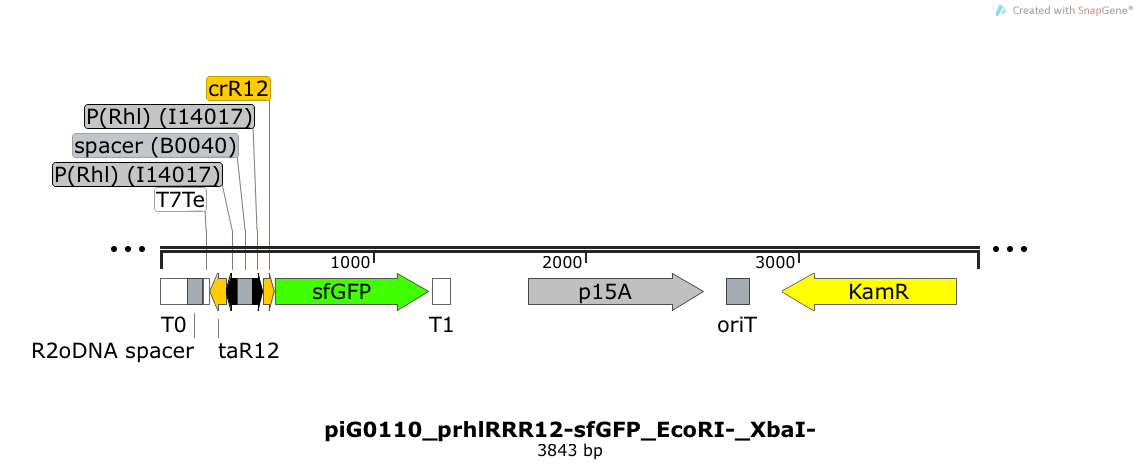
| - | Expression of sfGFP is induced when RhlR bound to 4C-HSL bind to pRhlR. The cis-repressive element (crR12) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a [https://2014.igem.org/Team:ETH_Zurich/expresults#Riboregulators riboregulator] that decreases leakiness of pLuxR. The p15A is present at a copy number of approximately 15 to 25<sup>[[Team:ETH_Zurich/project/references#refChang|[35]]]</sup>. PiG0110 is a derivate of piG0066 where the restriction sites EcoRI and XbaI have been removed. Thus, the two constructs slightly differ in the sequence of the 3'-end of the trans-activating element and in the sequence of the 5'-end of the cis-repressive element. | + | Expression of sfGFP is induced when RhlR bound to 4C-HSL bind to pRhlR. The cis-repressive element (crR12) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a [https://2014.igem.org/Team:ETH_Zurich/expresults#Riboregulators riboregulator] that decreases leakiness of pLuxR. A biologically neutral spacer sequence was designed using the web application R2oDNA<sup>[[Team:ETH_Zurich/project/references#refR2oDNA|[34]]]</sup>. The p15A is present at a copy number of approximately 15 to 25<sup>[[Team:ETH_Zurich/project/references#refChang|[35]]]</sup>. PiG0110 is a derivate of piG0066 where the restriction sites EcoRI and XbaI have been removed. Thus, the two constructs slightly differ in the sequence of the 3'-end of the trans-activating element and in the sequence of the 5'-end of the cis-repressive element. |
[[File:ETH2014 piG0110 prhlRRR12-sfGFP EcoRI- XbaI- Map.png|link=https://static.igem.org/mediawiki/2014/f/f9/ETH2014_piG0110_prhlRRR12-sfGFP_EcoRI-_XbaI-.txt]] | [[File:ETH2014 piG0110 prhlRRR12-sfGFP EcoRI- XbaI- Map.png|link=https://static.igem.org/mediawiki/2014/f/f9/ETH2014_piG0110_prhlRRR12-sfGFP_EcoRI-_XbaI-.txt]] | ||
Revision as of 21:16, 17 October 2014
Sequences
The plasmid sequences can be accessed by clicking on the plasmid name (e.g. piG0040) or the plasmid picture.
Construct types:
Combined Sensor and Producer Constructs
Regulator Constructs
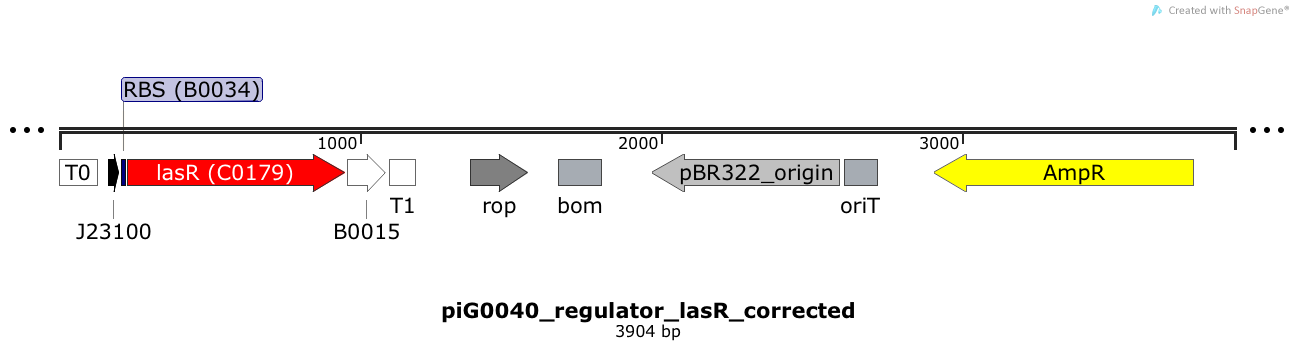
LasR is expressed under the strong constitutive promoter BBa_J23100 while the expression level is influenced by the RBS BBa_B0034. LasR bound to 3OC12-HSL induces the expression of genes under the control of pLasR. The plasmid pBR322 and its derivatives have a copy number of 15 to 20[15].
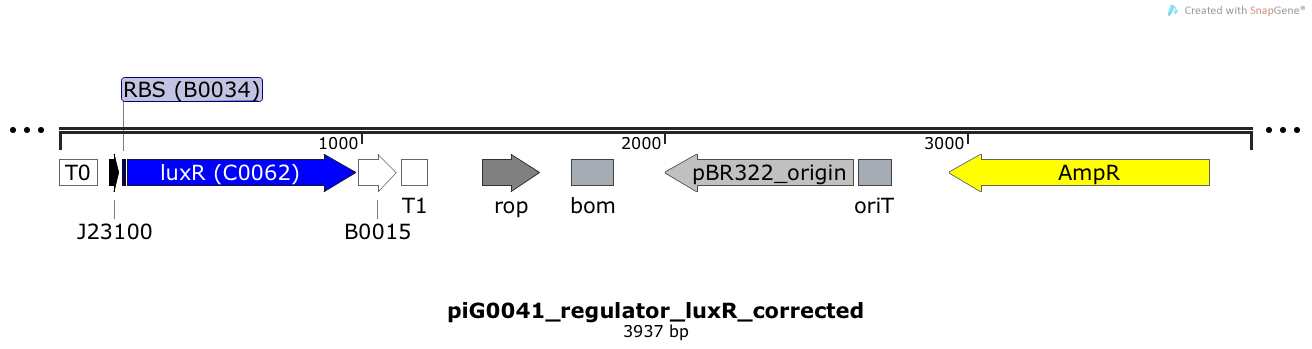
LuxR is expressed under the strong constitutive promoter BBa_J23100 while the expression level is influenced by the RBS BBa_B0034. LuxR bound to 3OC6-HSL induces the expression of genes under the control of pLuxR. The plasmid pBR322 and its derivatives have a copy number of 15 to 20[15].
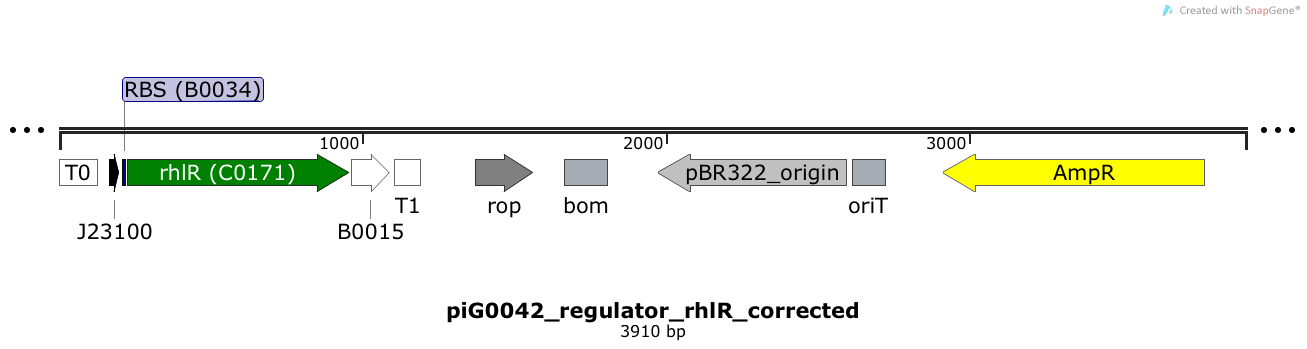
RhlR is expressed under the strong constitutive promoter BBa_J23100 while the expression level is influenced by the RBS BBa_B0034. RhlR bound to C4-HSL induces the expression of genes under the control of pRhlR. The plasmid pBR322 and its derivatives have a copy number of 15 to 20[15].
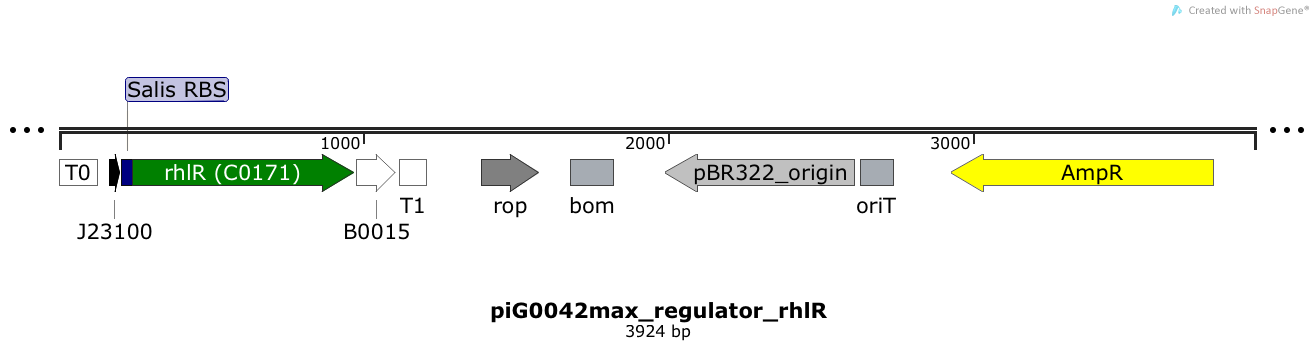
RhlR is expressed under the strong constitutive promoter BBa_J23100 while the expression level is influenced by an RBS optimised for RhlR (RBS calculator). RhlR bound to C4-HSL induces the expression of genes under the control of pRhlR. The plasmid pBR322 and its derivatives have a copy number of 15 to 20[15].
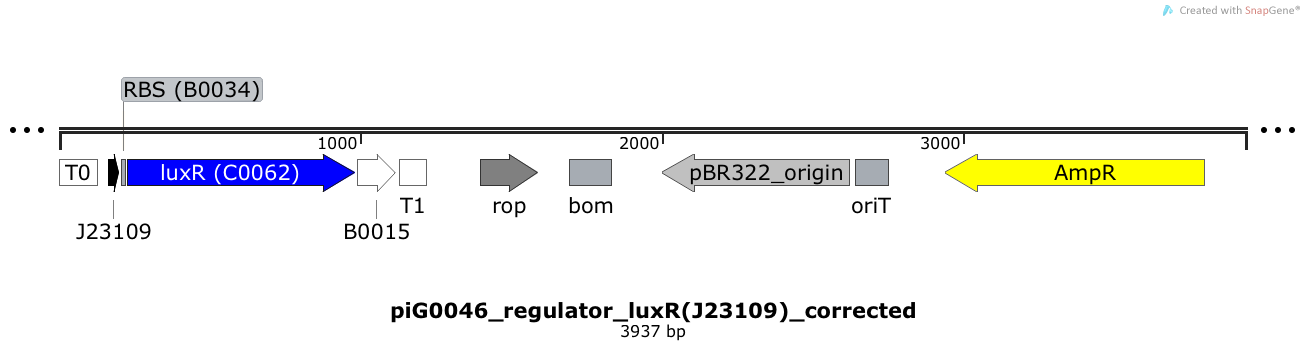
LuxR is expressed under the weak constitutive promoter BBa_J23109 while the expression level is influenced by the RBS BBa_B0034. LuxR bound to 3OC6-HSL induces the expression of genes under the control of pLuxR. The plasmid pBR322 and its derivatives have a copy number of 15 to 20[15].
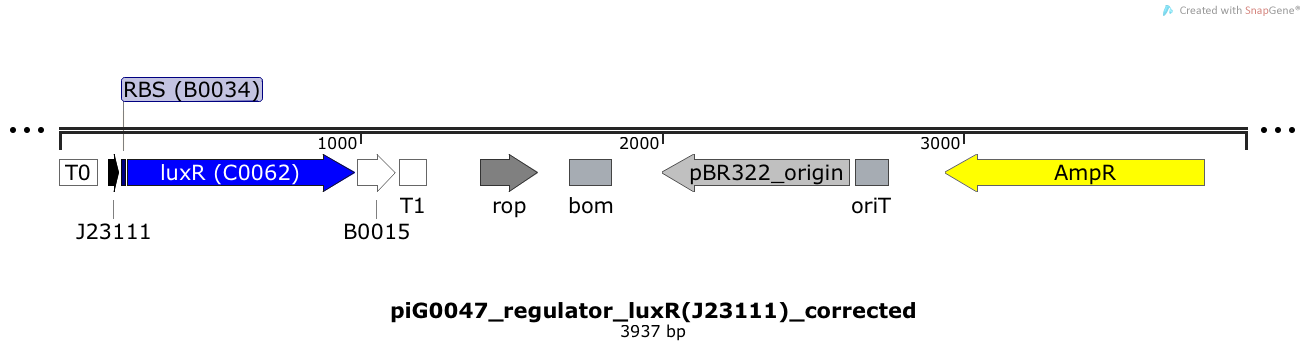
LuxR is expressed under the medium strong constitutive promoter BBa_J23111 while the expression level is influenced by the RBS BBa_B0034. LuxR bound to 3OC6-HSL induces the expression of genes under the control of pLuxR. The plasmid pBR322 and its derivatives have a copy number of 15 to 20[15].
Producer Constructs
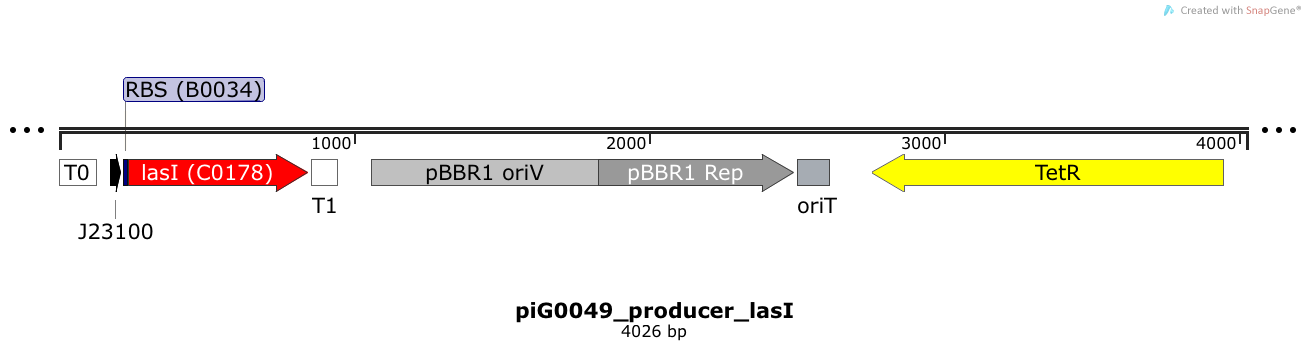
LasI is expressed under the strong constitutive promoter BBa_J23100 while the expression level is influenced by the RBS BBa_B0034. LasI produces the quorum sensing molecule 3OC12-HSL. The pBBR1 origin is present at a copy number of approximately 5, however, the origin is poorly characterized [16].
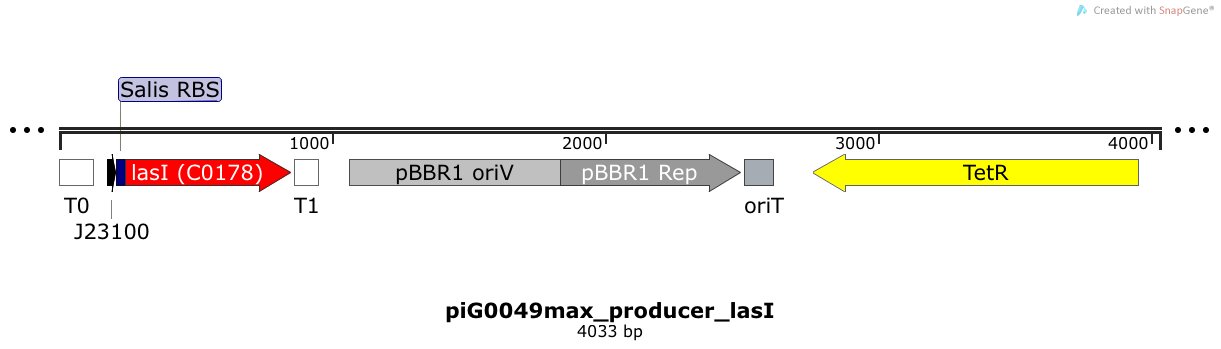
LasI is expressed under the strong constitutive promoter BBa_J23100 while the expression level is increased by an RBS optimised for LasI (RBS calculator). LasI produces the quorum sensing molecule 3OC12-HSL. The pBBR1 origin is present at a copy number of approximately 5, however, the origin is poorly characterized [16].
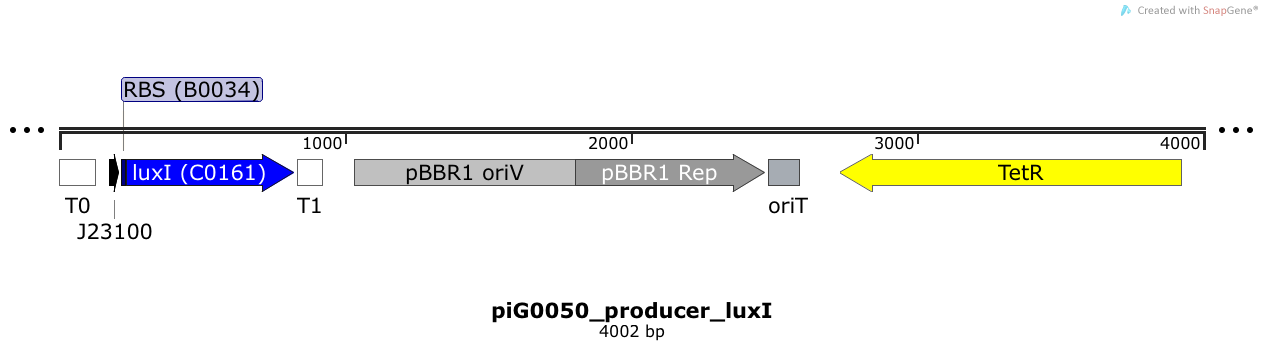
LuxI is expressed under the strong constitutive promoter BBa_J23100 while the expression level is influenced by the RBS BBa_B0034. LuxI produces the quorum sensing molecule 3OC6-HSL. The pBBR1 origin is present at a copy number of approximately 5, however, the origin is poorly characterized[16].
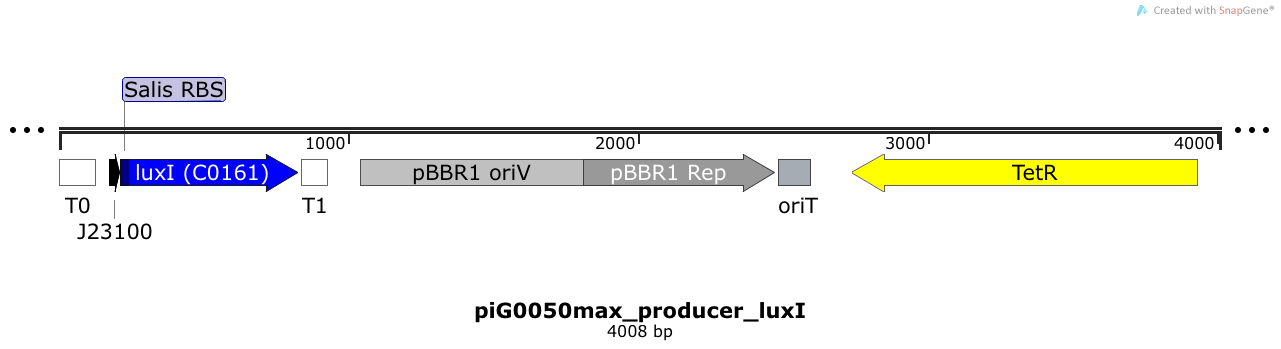
LuxI is expressed under the strong constitutive promoter BBa_J23100 while the expression level is increased by an RBS optimised for LuxI (RBS calculator). LuxI produces the quorum sensing molecule 3OC6-HSL. The pBBR1 origin is present at a copy number of approximately 5, however, the origin is poorly characterized[16].
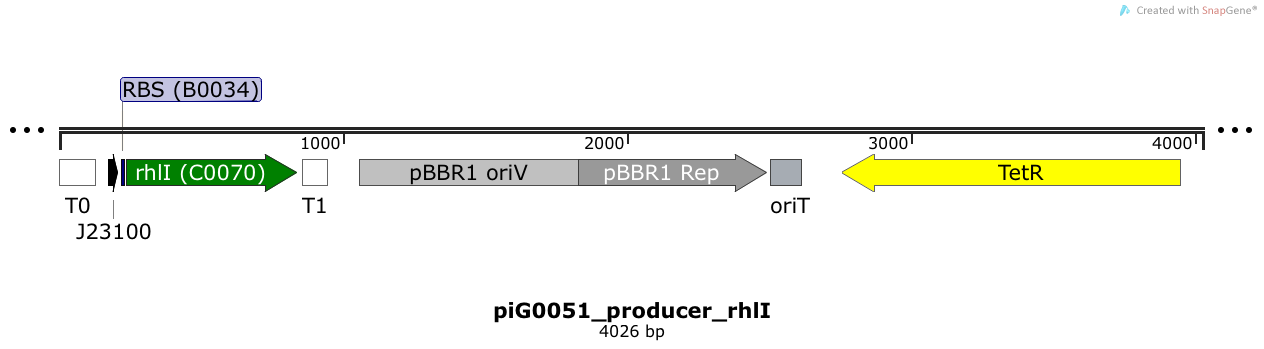
RhlI is expressed under the strong constitutive promoter BBa_J23100 while the expression level is influenced by the RBS BBa_B0034. RhlI produces the quorum sensing molecule C4-HSL. The pBBR1 origin is present at a copy number of approximately 5, however, the origin is poorly characterized[16].
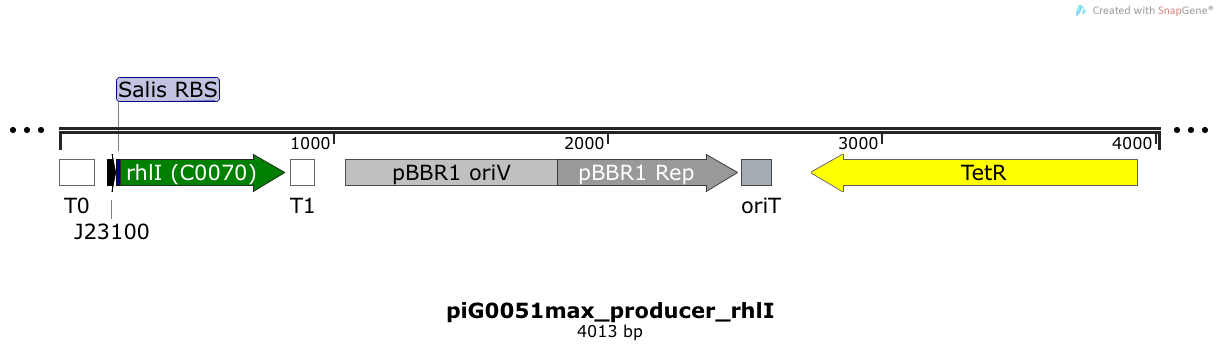
RhlI is expressed under the strong constitutive promoter BBa_J23100 while the expression level is increased by an RBS optimised for RhlI (RBS calculator). RhlI produces the quorum sensing molecule C4-HSL. The pBBR1 origin is present at a copy number of approximately 5, however, the origin is poorly characterized[16].
Sensor Constructs
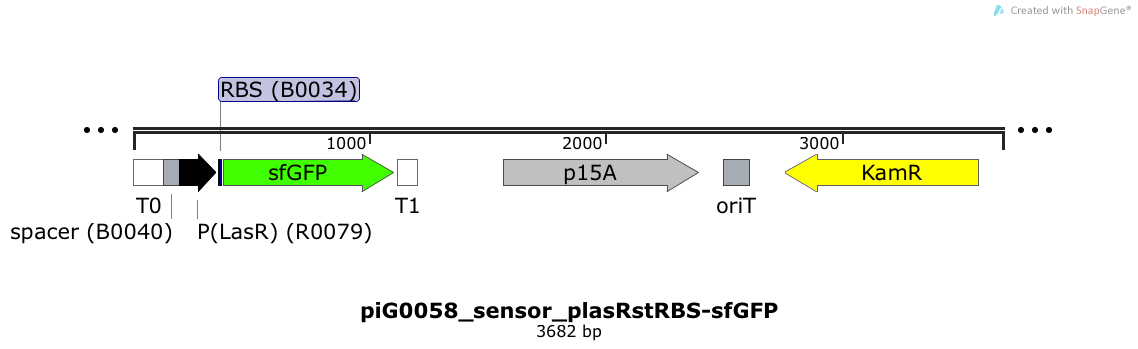
Expression of sfGFP is induced when LasR bound to 3OC12-HSL bind to pLasR. The p15A is present at a copy number of approximately 15 to 25[35].
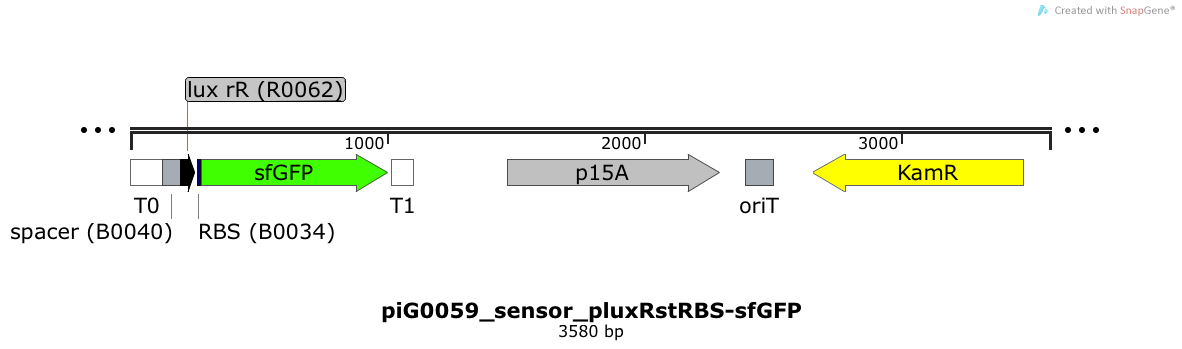
Expression of sfGFP is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The p15A is present at a copy number of approximately 15 to 25[35].
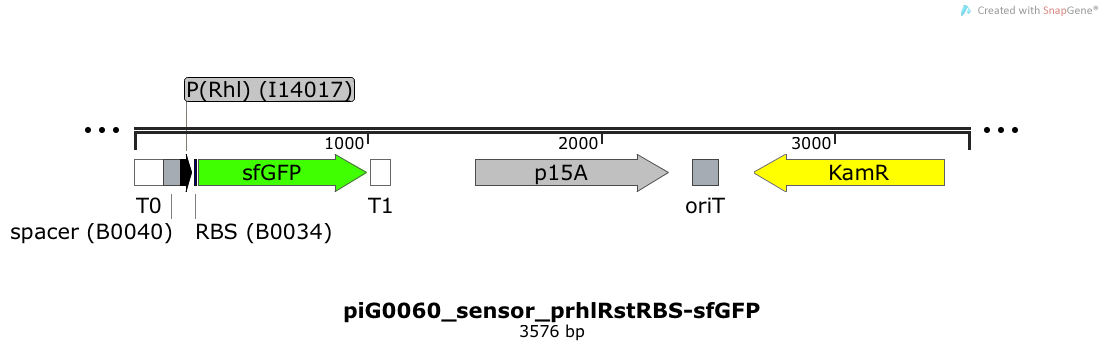
Expression of sfGFP is induced when RhlR bound to 4C-HSL bind to pRhlR. The p15A is present at a copy number of approximately 15 to 25[35].
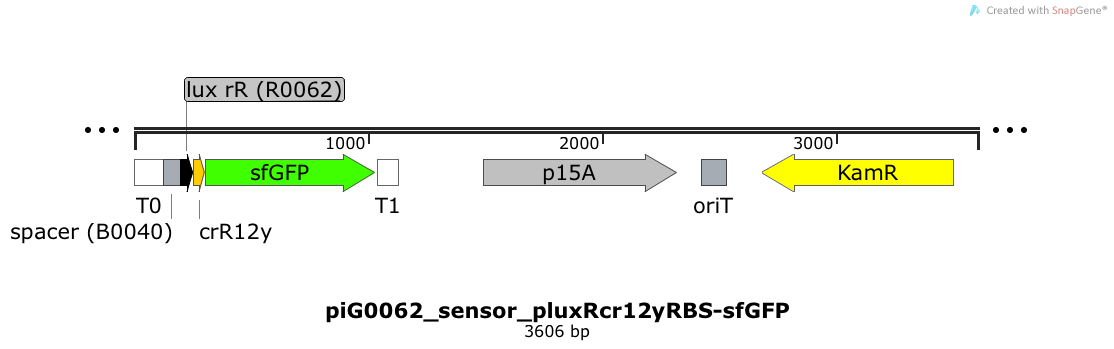
Expression of sfGFP is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The p15A is present at a copy number of approximately 15 to 25[35].
Expression of sfGFP is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a riboregulator that decreases leakiness of pLuxR. A biologically neutral spacer sequence was designed using the web application R2oDNA[34]. The p15A is present at a copy number of approximately 15 to 25[35].
Expression of sfGFP is induced when RhlR bound to 4C-HSL bind to pRhlR. The cis-repressive element (crR12) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a riboregulator that decreases leakiness of pLuxR. A biologically neutral spacer sequence was designed using the web application R2oDNA[34]. The p15A is present at a copy number of approximately 15 to 25[35].
Expression of sfGFP is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a riboregulator that decreases leakiness of pLuxR. A biologically neutral spacer sequence was designed using the web application R2oDNA[34]. The p15A is present at a copy number of approximately 15 to 25[35]. PiG0109 is a derivate of piG0065 where the restriction sites EcoRI and XbaI have been removed. Thus, the two constructs slightly differ in the sequence of the 3'-end of the trans-activating element and in the sequence of the 5'-end of the cis-repressive element.
Expression of sfGFP is induced when RhlR bound to 4C-HSL bind to pRhlR. The cis-repressive element (crR12) inhibits the translation of sfGFP, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a riboregulator that decreases leakiness of pLuxR. A biologically neutral spacer sequence was designed using the web application R2oDNA[34]. The p15A is present at a copy number of approximately 15 to 25[35]. PiG0110 is a derivate of piG0066 where the restriction sites EcoRI and XbaI have been removed. Thus, the two constructs slightly differ in the sequence of the 3'-end of the trans-activating element and in the sequence of the 5'-end of the cis-repressive element.
BUFFER Gate Construct
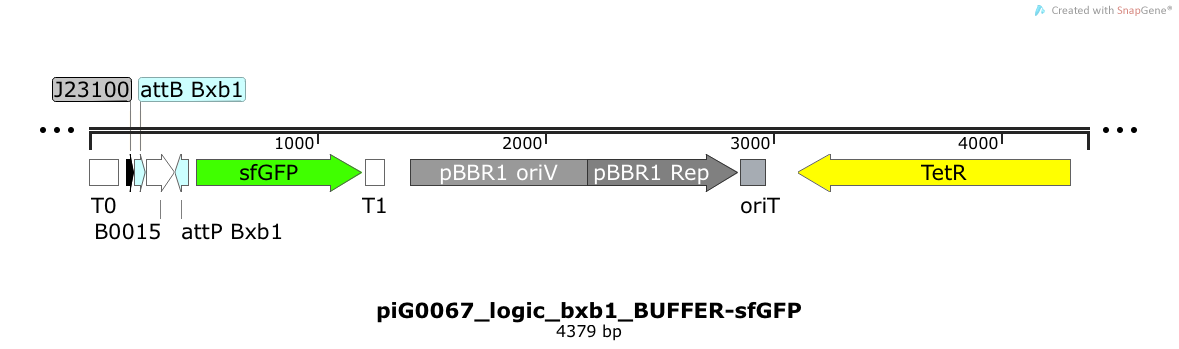
The directional terminator BBa_B0015 blocks transcription of sfGFP under the strong constitutive promoter BBa_J23100. Only if the integrase Bxb1 flips B0015 between the attP and the attB sites, transcription of sfGFP is possible. The pBBR1 origin is present at a copy number of approximately 5, however, the origin is poorly characterized[16]
Signal Propagation Construct
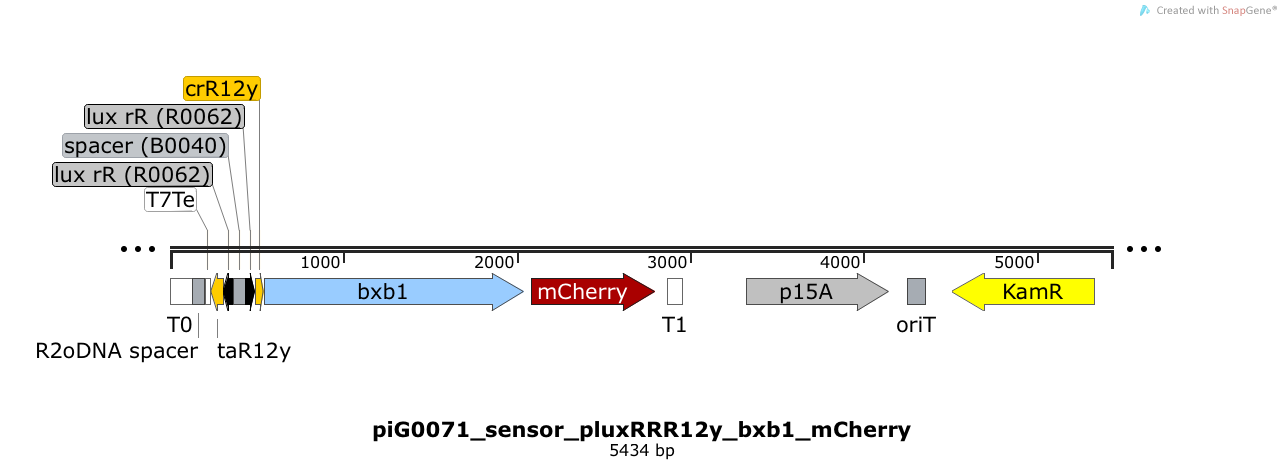
Expression of the integrase Bxb1 and the fluorophore mCherry is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of Bxb1 and mCherry, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a riboregulator that decreases leakiness of pLuxR. The p15A is present at a copy number of approximately 15 to 25[35].
Combined Sensor and Producer Constructs
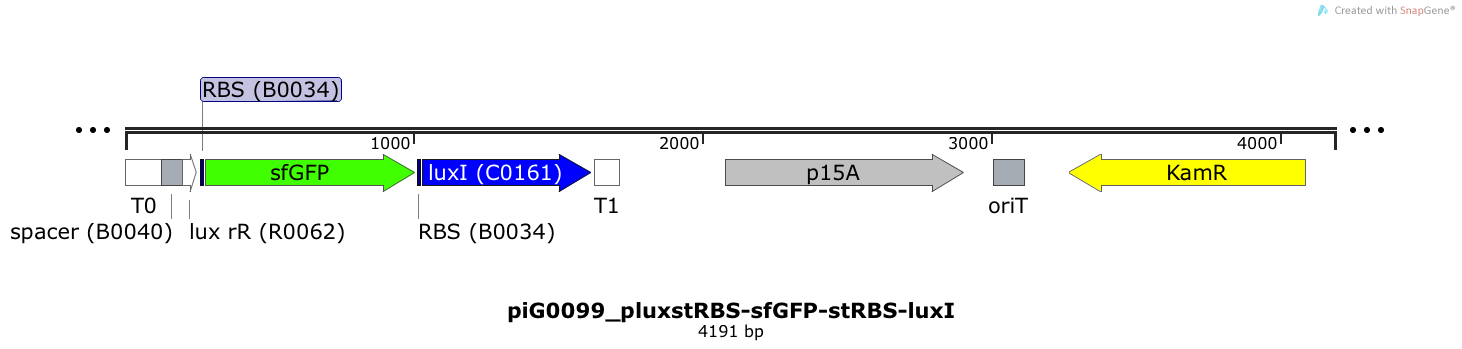
Expression of sfGFP and LuxI is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of the succeeding gene, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a riboregulator that decreases leakiness of pLuxR. The expression level of LuxI is increased by an RBS optimised for LuxI (RBS calculator).The p15A is present at a copy number of approximately 15 to 25[35].
Expression of sfGFP and LuxI is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The cis-repressive element (crR12y) inhibits the translation of the succeeding gene, since the RBS is blocked by secondary structures of the mRNA. The transcript of the trans-activating element (taR12y) binds to the transcript of the cis-repressive element, hence the RBS is not blocked anymore. The two elements build a riboregulator that decreases leakiness of pLuxR. The expression level of LuxI is influenced by the RBS BBa_B0034. The p15A is present at a copy number of approximately 15 to 25[35].
Expression of sfGFP and LuxI is induced when LuxR bound to 3OC6-HSL bind to pLuxR. The expression level of sfGFP and LuxI is influenced by the RBS BBa_B0034. The p15A is present at a copy number of approximately 15 to 25[35].
 "
"