Team:ETH Zurich/blog
From 2014.igem.org
{{{1}}}
Blog
Title ?
Digest and miniprep
Digest and miniprep
Miniprep
Preparation of DNA from iGEM kit
Sequencing, Transformation, antibiotic stock preparation
Miniprep
Template:Team:ETH Zurich/labblog/20140510humsum
Week 7: Human practice planning, Plasmid assembly running, Logo design
Wednesday, July 9th
- We have a logo !
- In the lab, we did some :
- Plasmid preparation
- Sequencing
- Digests
- Purification of backbone fragments needed for GA
- On the modeling side, we tried to estimate integrase parameters from the paper from Bonnet et al. [9], more particularly with the figure S4 :
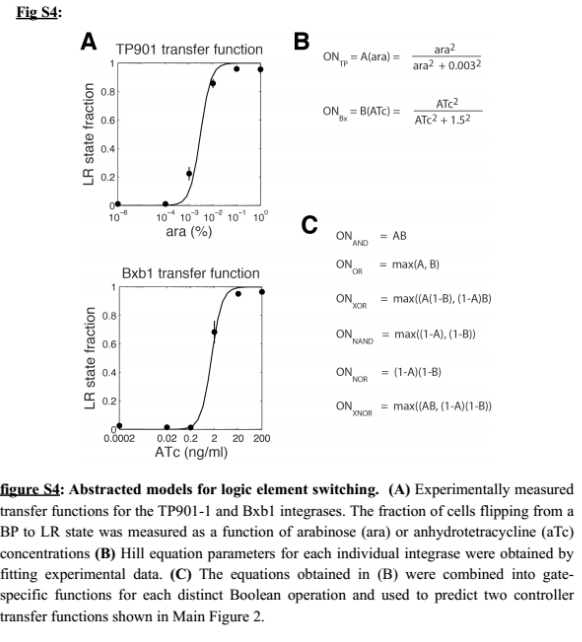
We are trying several strategies : minimization of the error function and Markov Chain Monte Carlo.
- We have a precise plan for our human practice project. We would like to study the emergence of complexity in many different fields and investigate how people deal with it. We will do this with interviews of experts in different fields, and with a survey for a wider outreach. We will also reach younger people by organizing talks in schools. We will finally give our own advice on the question by writing an essay on the subject and linking it to our experience with Mosaicoli.
Digest and miniprep
Tuesday, July 8th
Optimizing Restriction Endonuclease Reactions
Digest of 291 and 561
291: (2x)
1μL AscI
1μL FseI
0.5μL PvuI
0.5μL AseI
20 μL DNA 291
5μL Cut Smart Buffer
fill up with 22 μL H2O to 50μL
dephosphorylation of 181 and 271, cleanup with PCR clean-up system, gel electrophoresis
Gel electrophoresis
0.5 g Agarose in 50 mL TAE (0.5x), heat in microwave to dissolve
let solution cool down to approximately 50 °C, add 5 μL peqGREEN, mix
pour solution in tray, use appropriate comb
add 10μL loading dye (6x) to samples
fill samples and ladder (10 μL) in wells
run gel at 135 V for 50min
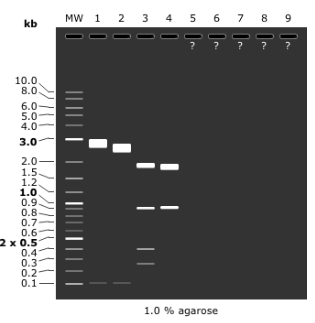
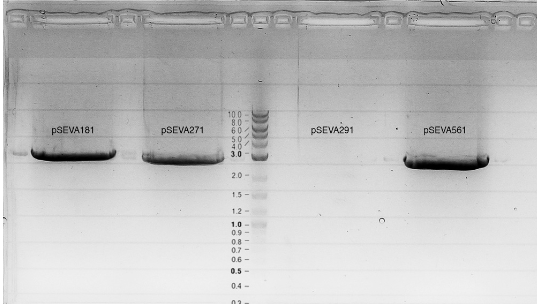
Miniprep of 35 and 36
Digest and miniprep
Monday, July 7th
Optimizing Restriction Endonuclease Reactions
Digest of 291 and 561
291: (2x)
1μL AscI
1μL FseI
0.5μL PvuI
0.5μL AseI
20 μL DNA 291
5μL Cut Smart Buffer
fill up with 22 μL H2O to 50μL
Miniprep of 35 and 36
Miniprep
Sunday, July 6th
Miniprep with bacteria cultures containing 271, 291, 181 and 30
Addition of 200 μL Resuspension solution to culture pellet, vortex, addition of 200 μL Lysis solution
After ca. 5 min addition of 350 μL Neutralization solution, centrifuge samples for 10 min at 4000 rpm
Preparation of columns: addition of 500 μL Preparation solution, centrifugation for 1 min at 12 000 rcf, discard flow-through
Give supernatant of centrifuged samples onto columns, spin for 1 min at 12 000 rcf, discard flow-through
Addition of 500 μL Optional Wash Solution, spin for 1 min at 12 000 rcf, discard flow-through
Addition of 750 μL Wash Solution, spin for 1 min at 12 000 rcf, discard flow-through
Dry columns by centrifuging them for 2 min at 12 000 rcf, subsequently place columns in new collection tubes
Elute DNA with 100 μL Elution solution
According to Sigma-Aldrich Plasmid Miniprep Kit protocol
Preparation of DNA from iGEM kit
Friday, July 4th
E coli transformation
Addition of 10 μL H2O to appropriate well, wait for 5 min, transfer into sterile tube
Transformation of E. coli with pSEVA281 A-C2 (sfGFP), pSEVA271 C-A9 (mCherry) and piG0030:
Addition of 1 μL DNA to 75 μL competent cells (thawed on ice), put samples on ice for approximately 20 min
Heat shock: 90 s at 42 °C
Addition of SOC to samples, let cells recover for ca. 1h at 37 °C, 220 rpm
Plate 100 μL of bacteria suspension on LB-agar-plates containing the appropriate antibiotics (pSEVA281 A-C2, pSEVA271 C-A9: Kanamycin (50 μg/L), piG0030: Chloramphenicol (34 μg/L))
Let bacteria grow overnight at 37 °C
Sequencing, Transformation, antibiotic stock preparation
Thursday, July 3rd
Sequencing results
Biobricks of the plasmids piG0001, piG0015 and piG002 show the correct sequences
E coli Transformation
Transformation of E. coli with pSEVA181, pSEVA271, pSEVA291, pSEVA561.
Addition of 1 μL DNA to 75 μL competent cells (thawed on ice), put samples on ice for approximately 20 min
Heat shock: 90 s at 42 °C
Addition of SOC to samples, let cells recover for ca. 1h at 37 °C, 220 rpm
Plate 100 μL of bacteria suspension on LB-agar-plates containing the appropriate antibiotics (pSEVA181: Ampicillin (200 μg/L), pSEVA271 and pSEVA291: Kanamycin (50 μg/L), pSEVA561: Tetracycline (20 μg/L))
Let bacteria grow overnight at 37 °C
Antibiotic stock preparation
Preparation of antibiotics stock:
2 g Ampicillin were dissolved in 10 mL H2O and sterile filtered
0.5 g Kanamycin were dissolved in 10 mL H2O and sterile filtered
0.34 g Chloramphenicol were dissolved in 10 mL Ethanol
Week 6: Project planning, Gibson assemblies planning, First Matlab simulations
Wednesday, July 2nd
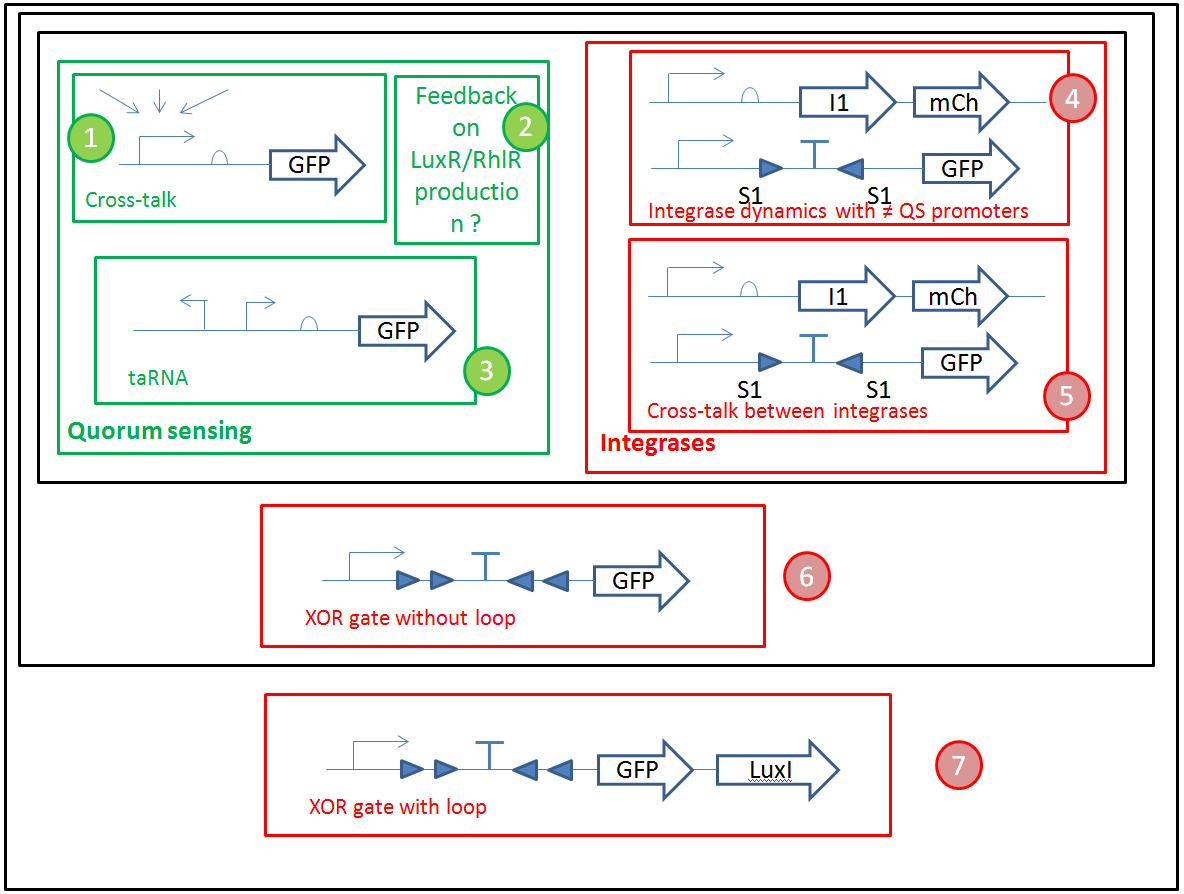
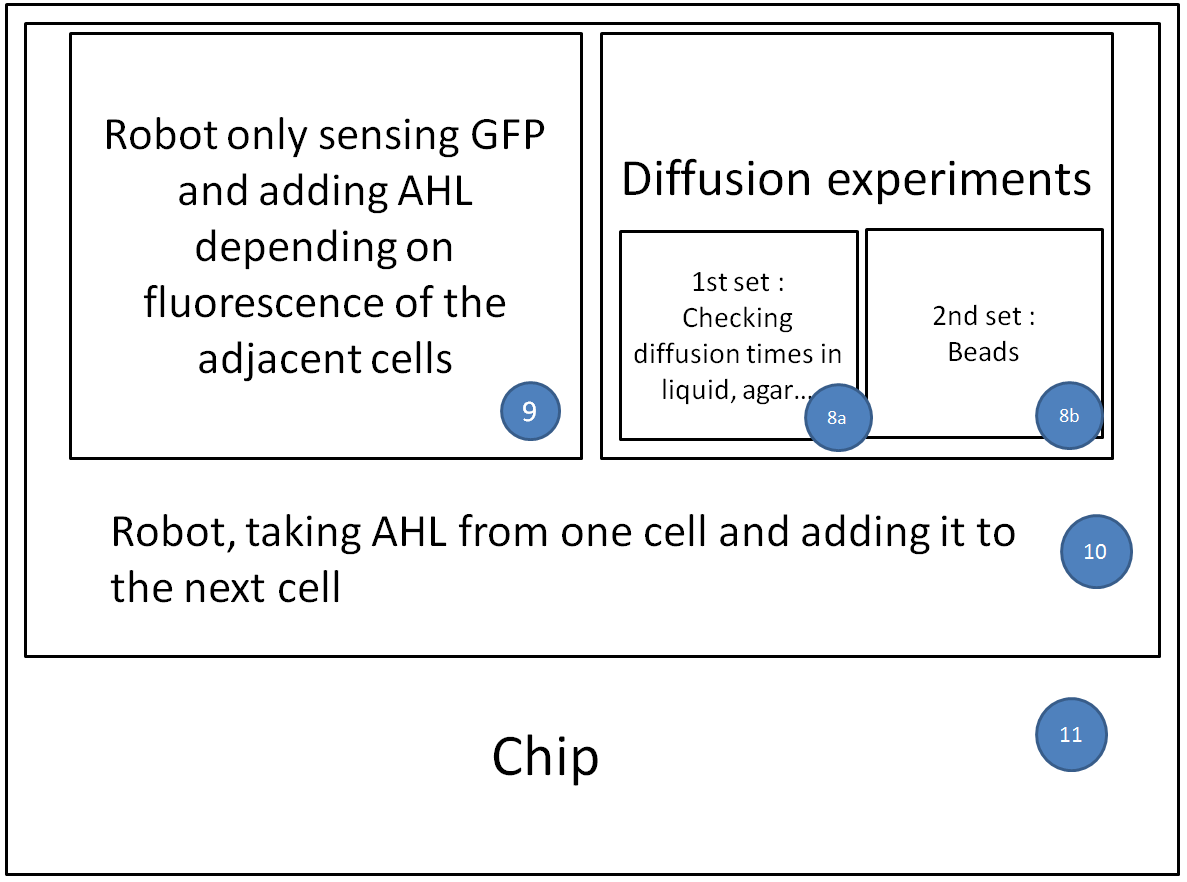
- We have a clear organization of the different experiments we would like to perform.
We want to optimize quorum sensing to have non-leaky non-cross-talking constructs. We want to prevent cross-talk between integrases and check their dynamics with different quorum sensing promoters. Then we will test the XOR gate without production of LuxI, to check how it works without the loop, and we will finally test our final construct with the loop, hoping that the delay between GFP production and the possible second switching due to AHL production is long enough to have enough fluorescence.
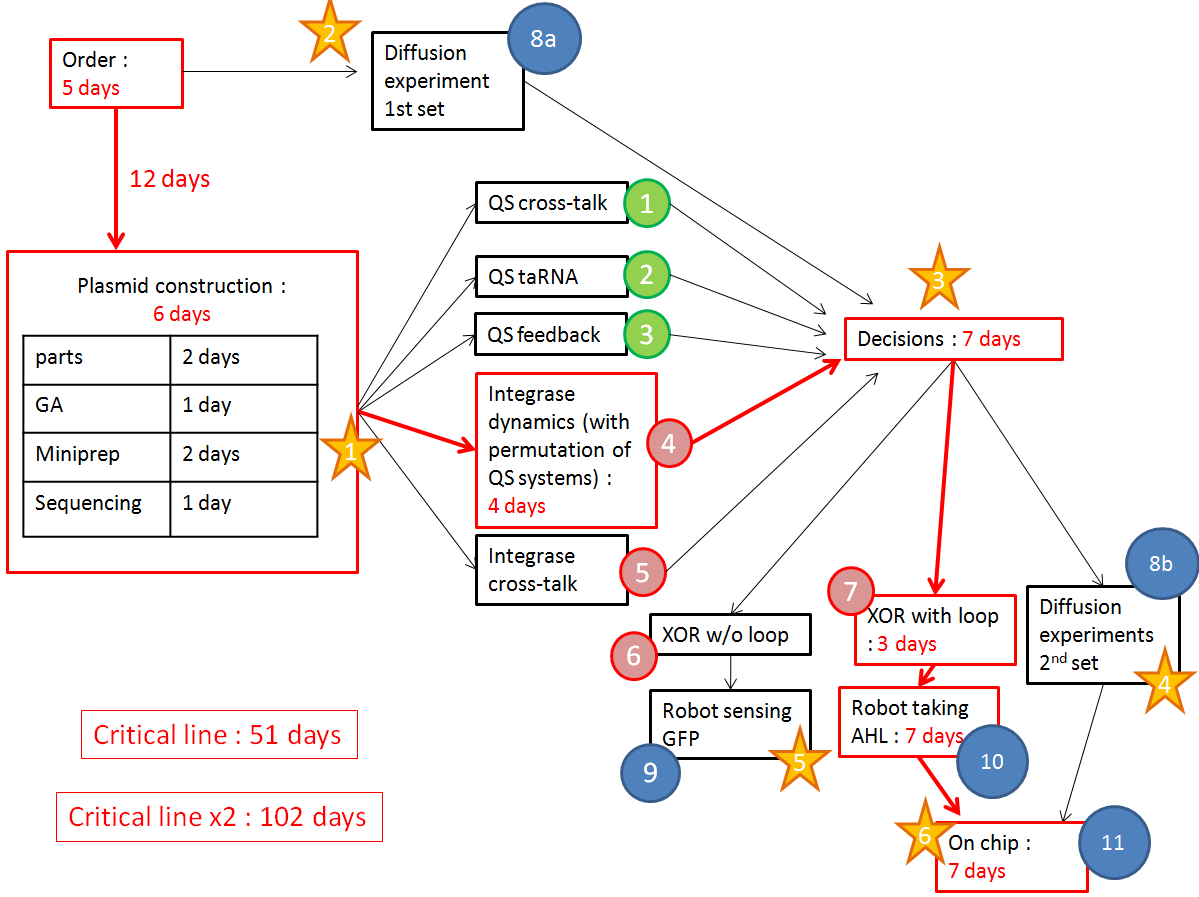
The red line is our critical timeline. Numbers of days are very optimistic,therefore we multiply the whole timeline by 2. The stars stand for modeling inputs.
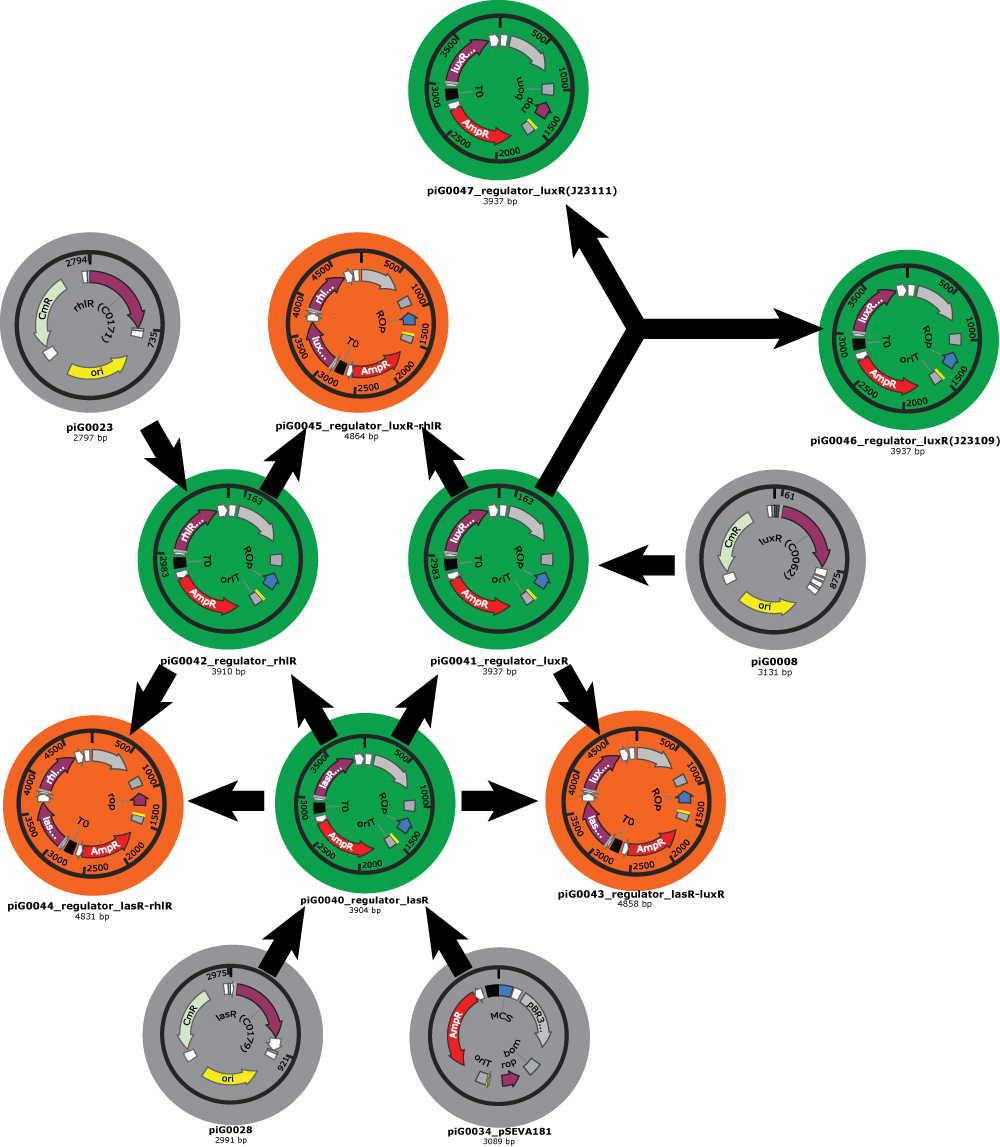
- We have also clear plans for the Gibson assemblies to perform. As an example, here is the plan for regulator plasmids :
- We have a facebook page !
- We simulated quorum sensing without leakiness on Matlab, with parameters from the literature.
BSSE Openhouse Day
Saturday, May 10th
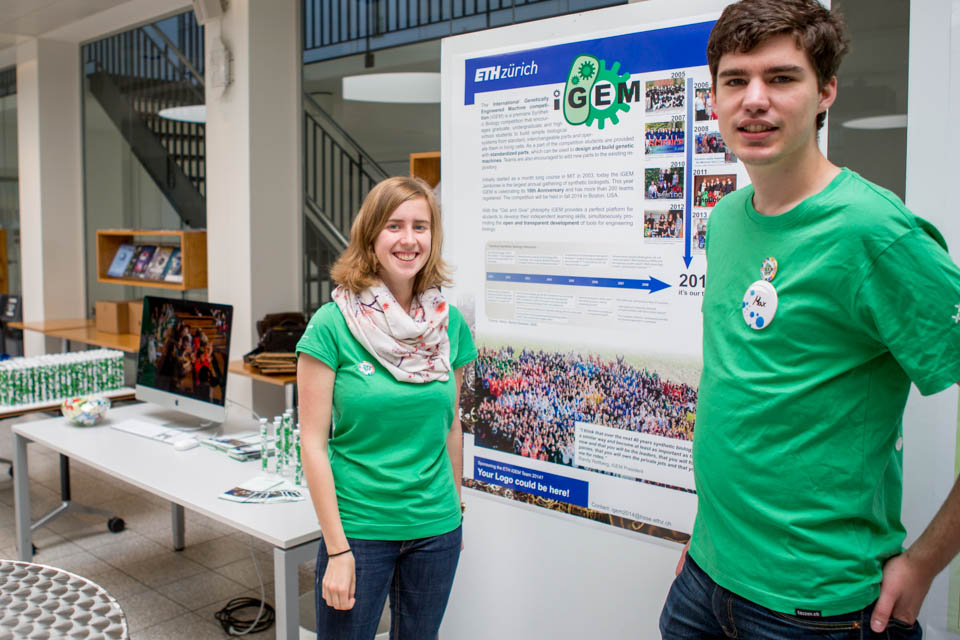
Sharing our iGEM and synthetic biology interest with the public
On May 10th the public in Basel had the unique chance to get an insight into many different scientific laboratories and the work done there. It was the joint open house day of D-BSSE of ETH (Department of Biosystems Science and Engineering) and the Biozentrum of the University of Basel. The many different labs opened their doors to the public and many scientists were present to give interested people some details about their daily work. So did the ETH iGEM team 2014. The team was present with a poster showing the history of iGEM, the previous ETH iGEM teams with their projects and general information about synthetic biology. Additionally there was a slideshow giving a best-of photo collection of last years jamboree. The goal of this day was to inform the public about synthetic biology in general and specifically about the spirit and the many different projects of iGEM. Many people showed strong interest in truly student driven projects and are curious to follow our team wiki for the next months.
 "
"