Team:ETH Zurich/blog
From 2014.igem.org
| Line 15: | Line 15: | ||
</html> | </html> | ||
| - | <html><article | + | <html><article id="20140705"></html> |
== Saturday, July 5th == | == Saturday, July 5th == | ||
{{:Team:ETH_Zurich/labblog/20140705}} | {{:Team:ETH_Zurich/labblog/20140705}} | ||
Revision as of 14:22, 9 July 2014
{{{1}}}
Blog
title of article
Preparation of DNA from iGEM kit
Sequencing, Transformation, antibiotic stock preparation
Miniprep
Saturday, July 5th
Construction of the regulator plasmids
Thursday, July 5th
Construction of the lasR regulator plasmid (piG0040)
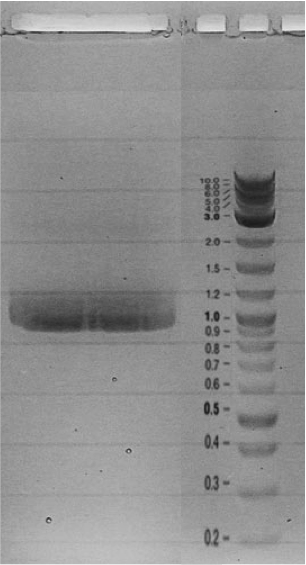
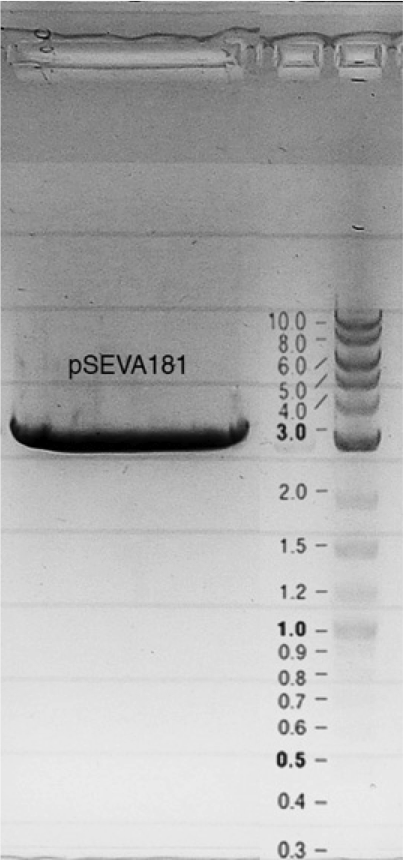
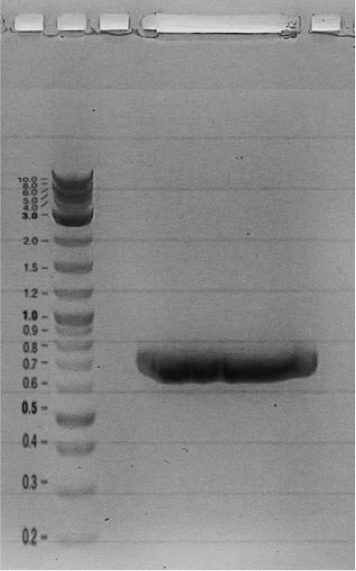
Competent cells were transformed with piG0028 (K553003, lasR) and selected on chloramphenicol-LB-plates. Plasmid DNA was extracted by performing a miniprep. The relevant plasmid sequence was sequenced by Microsynth using the primers oiG0001 and oiG0002. Amplification of lasR by PCR using oiG0003 and oiG0004 resulted in the fragment fiG0004 (1.0 kb). The vector piG0034 (pSEVA 181) was digested with the restriction enzymes HindIII and PacI to fiG0001 (3.0 kb). The two fragments, fiG0001 and fiG0004, were assembled using Gibbson assembly. Thus we used the plasmid backbone of piG0034 and lasR of piG0028 to construct the lasR regulator plasmid piG0040. Competent cells were transformed with piG0040 and selected on ampicillin-LB-plates. Colony PCR using the primers oiG0035 and oiG0036 was conducted to check the size of the inserted fragment (expected bands at 1.0 and 1.2 kb). The sequence was verified by Microsynth using the same primers.
Construction of the luxR regulator plasmid (piG0041)
Competent cells were transformed with piG0008 (F2620, luxR) and selected on chloramphenicol-LB-plates. Plasmid DNA was extracted by performing a miniprep. The relevant plasmid sequence was sequenced by Microsynth using the primers oiG0001 and oiG0002. Amplification of luxR by PCR using oiG0005 and oiG0006 resulted in the fragment fiG0005 (0.8 kb). The backbone of the vector piG0040 (lasR in pSEVA 181) was amplified by PCR using oiG0007 and oiG0008. This fragment, fiG0011 (3.2 kb), and fiG0005 were assembled using Gibbson assembly. Thus we formally replaced lasR by luxR to construct the luxR regulator plasmid piG0041. Competent cells were transformed with piG0041 and selected on ampicillin-LB-plates. Colony PCR using the primers oiG0035 and oiG0036 was conducted to check the size of the inserted fragment (expected bands at 1.1 and 1.2 kb). The sequence was verified by Microsynth using the same primers.
Construction of the rhlR regulator plasmid (piG0042)
Competent cells were transformed with piG0023 (C0171, rhlR) and selected on chloramphenicol-LB-plates. Plasmid DNA was extracted by performing a miniprep. The relevant plasmid sequence was sequenced by Microsynth using the primers oiG0001 and oiG0002. Amplification of rhlR by PCR using oiG0009 and oiG0010 resulted in the fragment fiG0006 (0.8 kb). The backbone of the vector piG0040 (lasR in pSEVA 181) was amplified by PCR using oiG0011 and oiG0012. This fragment, fiG0012 (3.2 kb), and fiG0006 were assembled using Gibbson assembly. Thus we formally replaced lasR by rhlR to construct the rhlR regulator plasmid piG0042. Competent cells were transformed with piG0042 and selected on ampicillin-LB-plates. Colony PCR using the primers oiG0035 and oiG0036 was conducted to check the size of the inserted fragment (expected bands at 1.1 and 1.2 kb). The sequence was verified by Microsynth using the same primers.
Construction of the luxR regulator plasmids with alternative constitutive promoters (piG0046 and piG0047)
The promoter site of the luxR regulator plasmid piG0041 was mutated by QuikChange site-specific mutagenesis to produce promoters of different strength. The primers oiG0031 and oiG0032 were used to establish piG0047, a plasmid with a promoter of intermediate strength (J23111). Analoguos the primers oiG0033 and oiG0034 were used to construct piG0046, a plasmid with a weak promoter (J23109). Competent cells were transformed with piG0046 and piG0047 and selected on ampicillin-LB-plates. The sequence was verified by Microsynth using the primers oig0035 and oiG0036.
Friday, July 4th
Preparation of DNA from iGEM kit
Friday, July 4th
E coli transformation
Addition of 10 μL H2O to appropriate well, wait for 5 min, transfer into sterile tube
Transformation of E. coli with pSEVA281 A-C2 (sfGFP), pSEVA271 C-A9 (mCherry) and piG0030:
Addition of 1 μL DNA to 75 μL competent cells (thawed on ice), put samples on ice for approximately 20 min
Heat shock: 90 s at 42 °C
Addition of SOC to samples, let cells recover for ca. 1h at 37 °C, 220 rpm
Plate 100 μL of bacteria suspension on LB-agar-plates containing the appropriate antibiotics (pSEVA281 A-C2, pSEVA271 C-A9: Kanamycin (50 μg/L), piG0030: Chloramphenicol (34 μg/L))
Let bacteria grow overnight at 37 °C
Thursday, July 3rd
Sequencing, Transformation, antibiotic stock preparation
Thursday, July 3rd
Sequencing results
Biobricks of the plasmids piG0001, piG0015 and piG002 show the correct sequences
E coli Transformation
Transformation of E. coli with pSEVA181, pSEVA271, pSEVA291, pSEVA561.
Addition of 1 μL DNA to 75 μL competent cells (thawed on ice), put samples on ice for approximately 20 min
Heat shock: 90 s at 42 °C
Addition of SOC to samples, let cells recover for ca. 1h at 37 °C, 220 rpm
Plate 100 μL of bacteria suspension on LB-agar-plates containing the appropriate antibiotics (pSEVA181: Ampicillin (200 μg/L), pSEVA271 and pSEVA291: Kanamycin (50 μg/L), pSEVA561: Tetracycline (20 μg/L))
Let bacteria grow overnight at 37 °C
Antibiotic stock preparation
Preparation of antibiotics stock:
2 g Ampicillin were dissolved in 10 mL H2O and sterile filtered
0.5 g Kanamycin were dissolved in 10 mL H2O and sterile filtered
0.34 g Chloramphenicol were dissolved in 10 mL Ethanol
Wednesday, July 2nd
Week 6: Project planning, Gibson assemblies planning, First Matlab simulations
Wednesday, July 2nd
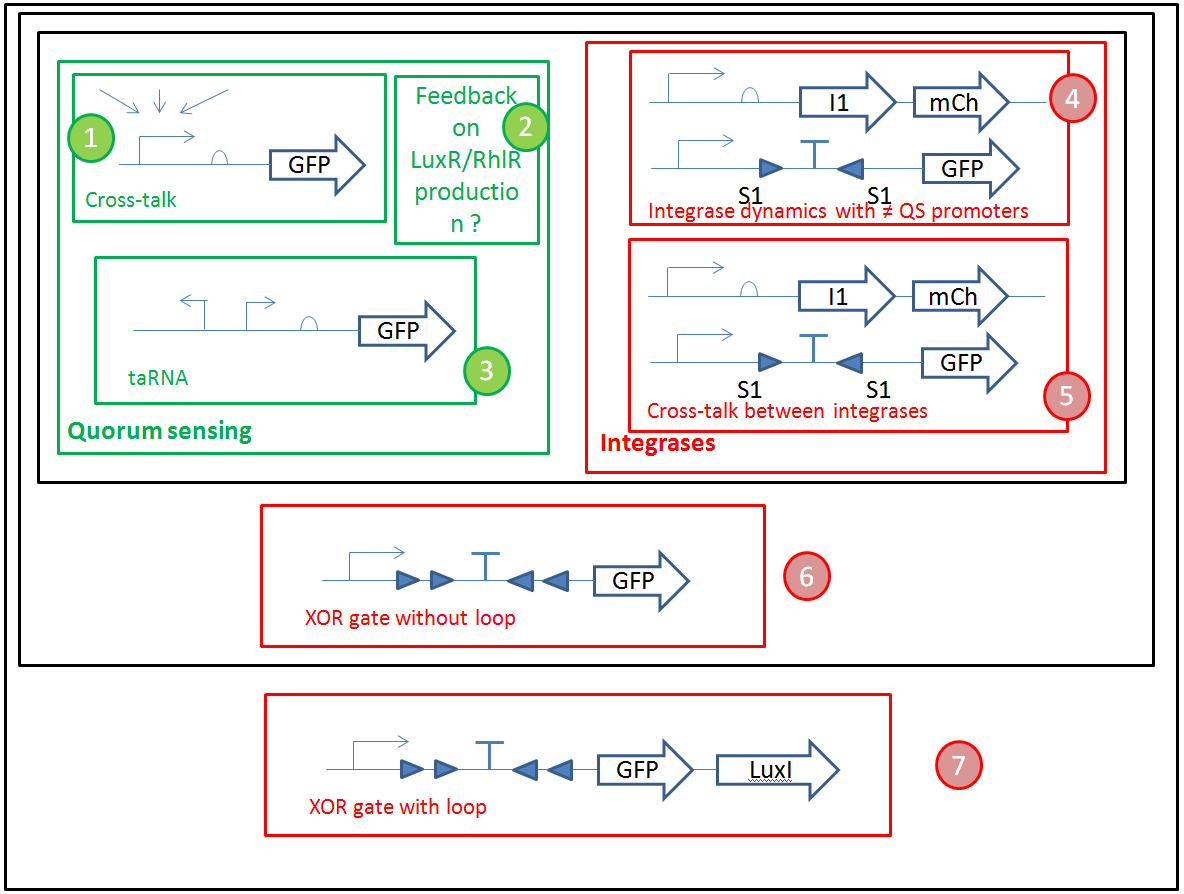
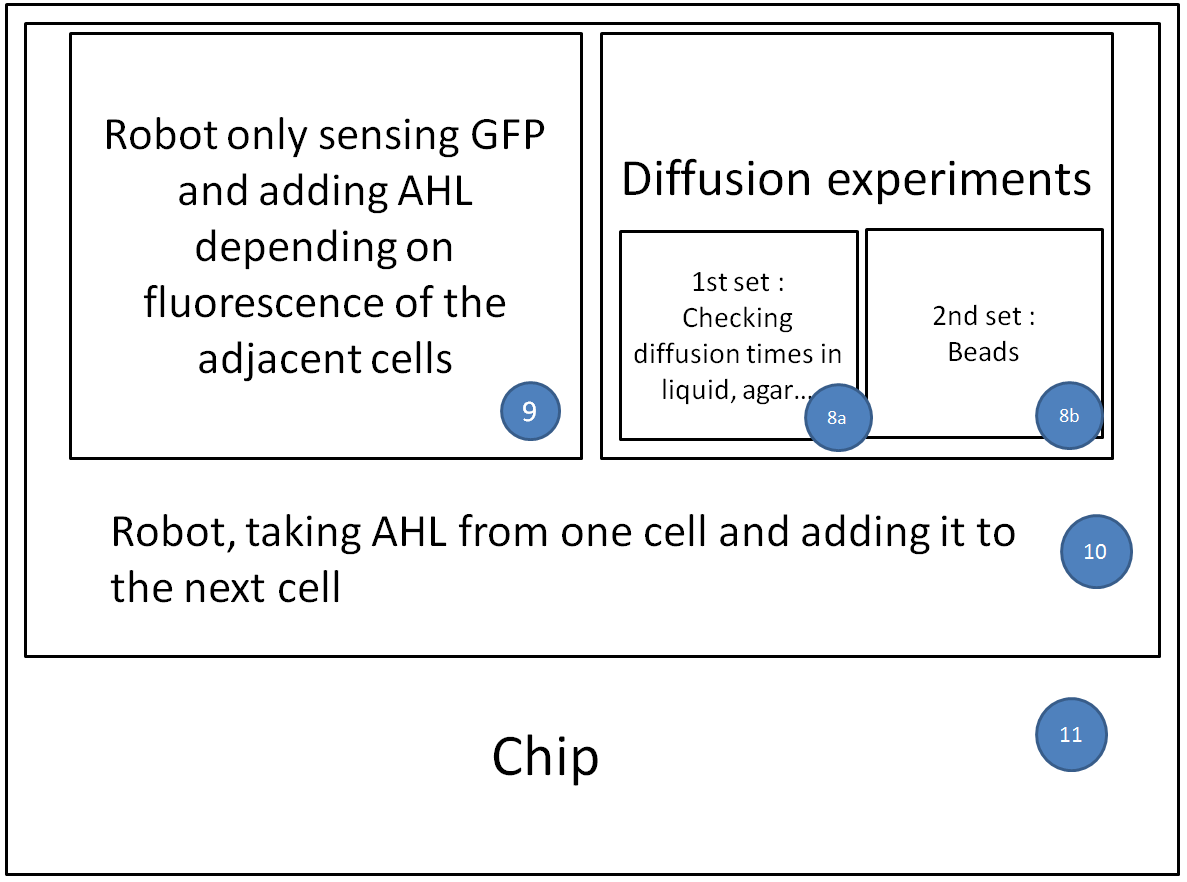
- We have a clear organization of the different experiments we would like to perform.
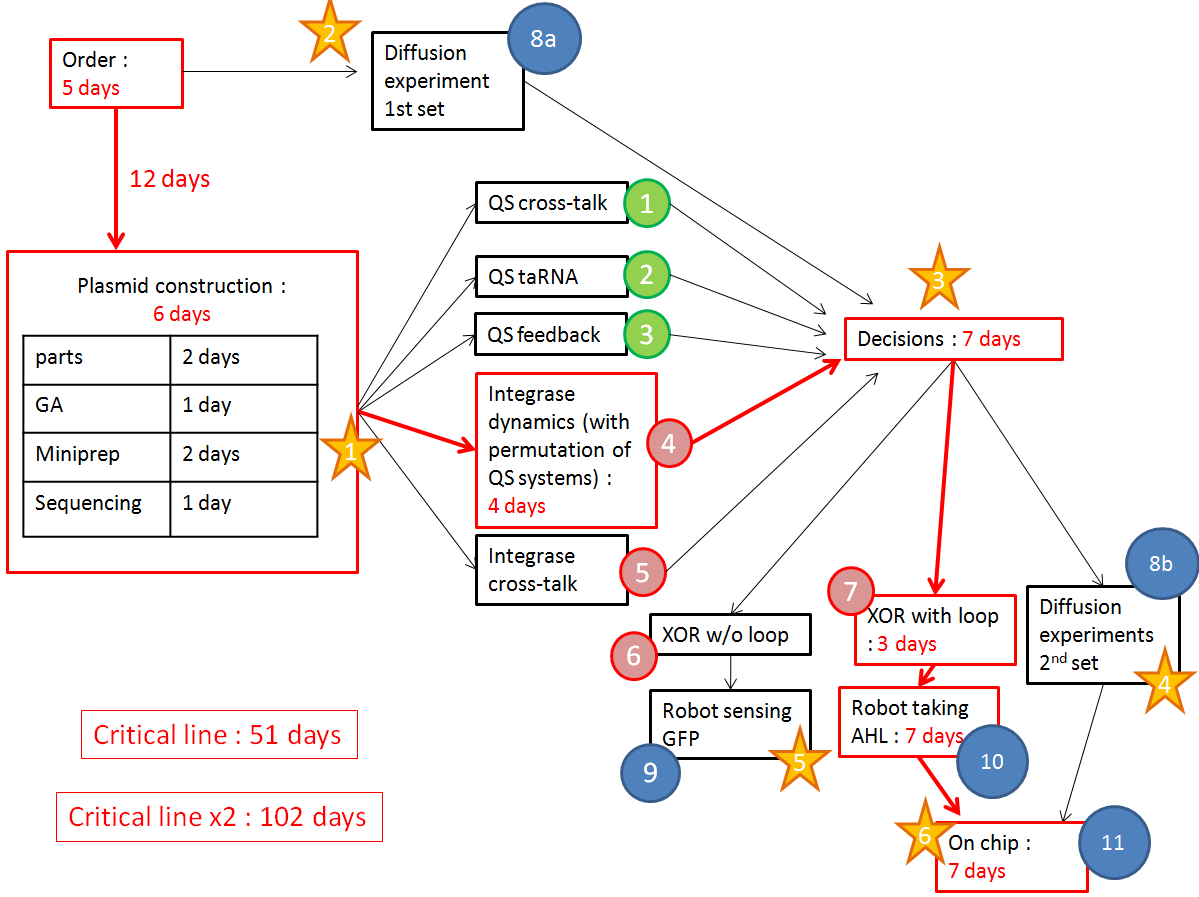
We want to optimize quorum sensing to have non-leaky non-cross-talking constructs. We want to prevent cross-talk between integrases and check their dynamics with different quorum sensing promoters. Then we will test the XOR gate without production of LuxI, to check how it works without the loop, and we will finally test our final construct with the loop, hoping that the delay between GFP production and the possible second switching due to AHL production is long enough to have enough fluorescence.
The red line is our critical timeline. Numbers of days are very optimistic,therefore we multiply the whole timeline by 2. The stars stand for modeling inputs.
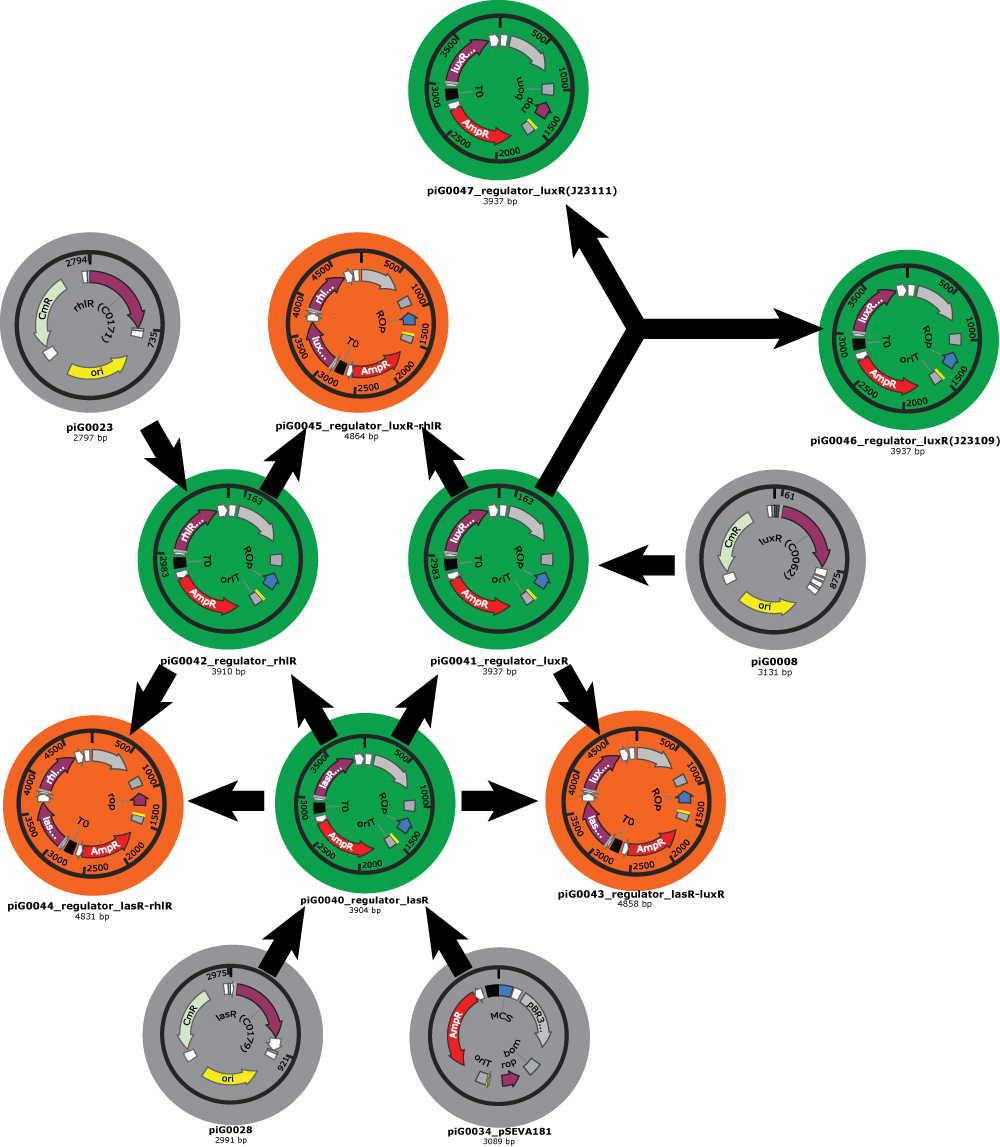
- We have also clear plans for the Gibson assemblies to perform. As an example, here is the plan for regulator plasmids :
- We have a facebook page !
- We simulated quorum sensing without leakiness on Matlab, with parameters from the literature.
 "
"