|
July
1st
- transformation efficiency kit is broken
→ run an agarose gel with the kit plasmids to check for nuclease activity/ bands
- inventory of the -80°C freezer was done
2nd
- overnight cultures (LB) of K731520 and K1319042 for chips were inoculated
3rd
- ODs of the overnight culture of
- K731520: 2.3
- K1319042: 2.5
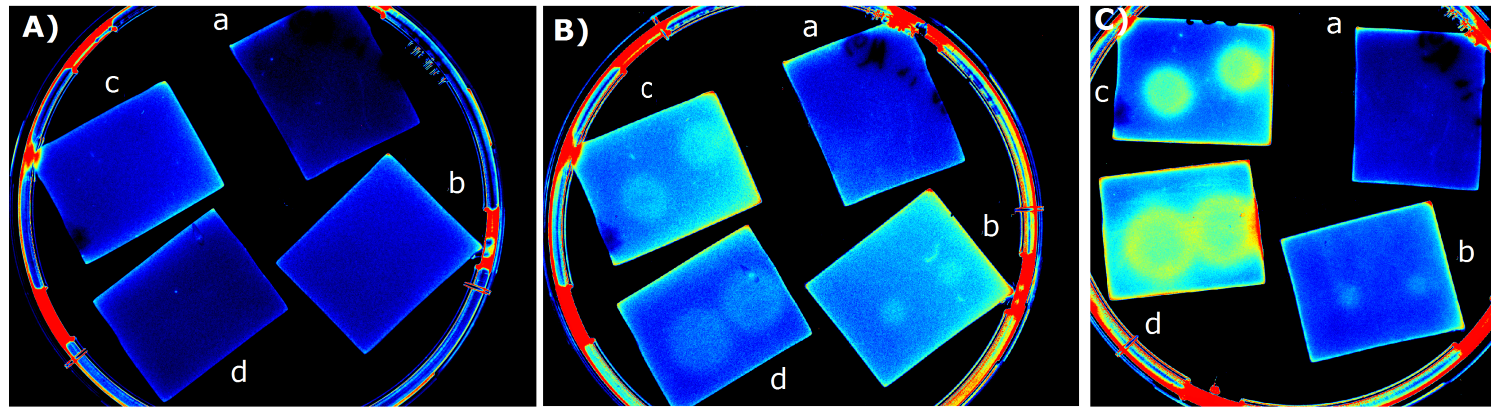
- made chips with K1319042 and K731520 in NA and M9 medium. Images were taken every 30 min with the GelDoc
- print K1319042 Chip on LB plate
- overnight culture of E0030 for plasmid prep
4th
- plasmid prep of E0030: 159.5 ng/µl (concentration)
- print of K1319042 Chip on LB plate made 1 and 7 colonies
7th
- 13 Biobricks were transformed
- overnight culture of K103006 for a plasmid prep was inoculated
- colony PCR master mix was prepared
8th
- overnight culture of K1319042 for chips was inoculated
9th
- made chips with K1319042 in M9, M9 + casamino acids, Hartman medium (HM). Images were taken every 30 min with the Geldoc
- made OmpA-linker_iLOV 3A-Assambley
- colony PCR
- transformation for backbones
10th
- master plates of the strains for backbone isolation
14th
- built again J23115.E0240 for interlab study
15th
- made 500 ml LB and LB plates
- made the transformation of J23115.E0240
- overnight culture of K1319042 for chips
16th
- did plasmid preps
- transformation of different reporter strains
- NEB
- pSEVE641_BsFbFP
- pSEVA234_LasR
- pSEVE641_BsFbFP pSEVA234_LasR
- pSEVE641_BsFbFP pSB1C3_C0179
- BL21
- pSEVE641_BsFbFP pSEVA234_LasR
- pSEVE641_BsFbFP pSB1C3_C0179
- made chips with K1319042 in Hartman, Hartman + 20 % glycerol, M9 medium images were taken every 30 min with the Geldoc
22nd
- made 100x stocks for Hartmans minimal medium.
- made about 25 HM+C plates without glucose.
- K1319042 was plated on a HM+C plate
23rd
- made 25 new HM+C plates with 1 % agar and 4 g/L glucose
- added 170 µL of the glucose stock (500 g/L) to the glucose-free HM+C-plates
- 1 L of sterile HM+glucose was prepared
- K1319042 was plated on HM+C+Glucose and HM+C+Glucose drops, 3x 5 mL HM+C+Glucose precultures were inoculated (17:00)
24th
- K1319042 did not grow, neither on the plates, nor on in the liquid media. The most probable cause is that the E. coli is missing some vitamins
- Based on the M9 recipes by the Knight and Endy labs on OpenWetWare, we made a 20 mL supplement stock solution that can be added to 1x HM media
| Component | supplement for 1x HM [g/L] | final 1x concentration
|
| Casamino acids | 2 | 2 g/L
|
| Thiamine hydrochloride | 0.3 | 300 mg/L
|
| MgSO4/MgSO4*7H2O | 0.242 / 0.494 | 2 mmol/L
|
| CaCl2 | 0.011 | 0.1 mmol/L
|
The powders for 1 L of 1x HM were dissolved in 20 mL of a 10 % w/v Casamino acid stock solution and filter-sterilized (.22 µm PES). To make agar chips, we can add 1 mL to the hot agar mixture, together with the three 500 µL 100x stocks for the HM.
We made 250 mL of HM+C+Glucose+Supplements plates with 1.5 % agar.
Then we plated the following combinations:
| Construct | HM+C+Glucose+Supplements | LB+C
|
| K1319042 | - | +
|
| J23101.E0240 | + | +
|
We also prepared a 150 mL HM+C+Glucose+Supplements culture with K1319042 to hopefully make agar chips on Friday, July 25th.
Six of last weeks J23115.E0240 clones were plated on LB+C, and 5 mL LB+C precultures were inoculated for plasmid preparation.
25th
The K1319042 strain did not grow on the HM plate, while another strain with the J23101.E0240 construct did. Both have the pSB1C3 backbone. To confirm the plasmids and inserts, we set up a colony PCR:
| ID | Template | Product Length | Result
|
| 1 | J23101.E0240 #5 | 1233 |
|
| 2 | J23101.E0240 #6 | 1233 |
|
| 3 | J23101.E0240 #5 plasmid | 1233 |
|
| 4 | K1319042 LB colony | 2053 |
|
| 5 | K1319042 plasmid | 2053 |
|
| 6 | K731520 GFP plasmid | 2437 |
|
| 7 | water | none |
|
Reaction volume per tube was 15 µL. GoTaq Green Mastermix and the VF2 and VR primers were used. You can find the durations and temperatures the table below:
| parameter | duration | temp [°C]
|
| denature | 10:00 | 95
|
| denature | 00:30 | 95
|
| anneal | 00:30 | 49
|
| elongate | 02:36 | 72
|
| elongate | 05:00 | 72
|
| store | forever | 8
|
We mini-prepped the 6 overnight cultures of J23115.E0240 clones. He used 3 mL instead of 1.5 mL culture medium and eluted twice with 25 µL nuclease-free water. Everything else was according to the protocol of the illustra plasmidPrep Mini-Spin Kit.
The resulting DNA concentrations are listed in this table:
| clone # | concentration [ng/µl]
|
| 1 | 73.5
|
| 2 | 94.5
|
| 3 | 100.5
|
| 4 | 53
|
| 5 | 82
|
| 6 | 138
|
In the evening, we plated some strains on the different media to investigate why the K3139042 did not grow on the HM+suppl. plate. (Note: In the morning, we re-inoculated the HM+C+suppl. shake flask with cells from the LB-plate and by afternoon the culture did grow to a high cell density.)
| Construct | Host strain | LB+C | HM+C+Glucose | HM+C+Glucose+supplements
|
| J23101.E0240 | NEB10β | ++ | - | ++
|
| K1319042 | DH5alpha | ++ | - | (+)
|
| K131026 | NEB10β | ++ | - | -
|
| K131026 | DH5alpha | ++ | - | +
|
We inoculated J23101.E0240 in LB and HM+C+Glucose+supplements.
29th
- inoculate 2x 150 ml cultures HM+Glucose+C
- prepared 200 ml 1.5 % agar
- pre-cool centrifuge
- centrifuge 1x 50, 1x 100 and 1x 150 ml
- make 3 kinds of chips - OOx1, ODx2, ODx3
- pre cultures of K1319042 and K131026 in LB
30th
- made chips with K1319042 and K131026 in HM medium images were taken every 30 min with the Geldoc
- print chips on LB and LB + Cam , count 2-10 colonies
31th
- transformation of K1319042 and K131026 in DH5α, BL21 and NEB
|
 "
"