Team:Austin Texas/kit
From 2014.igem.org
(Difference between revisions)
| Line 73: | Line 73: | ||
<!-- WIKI CONTENT BEGINS --> | <!-- WIKI CONTENT BEGINS --> | ||
| - | + | ==Kit Introduction== | |
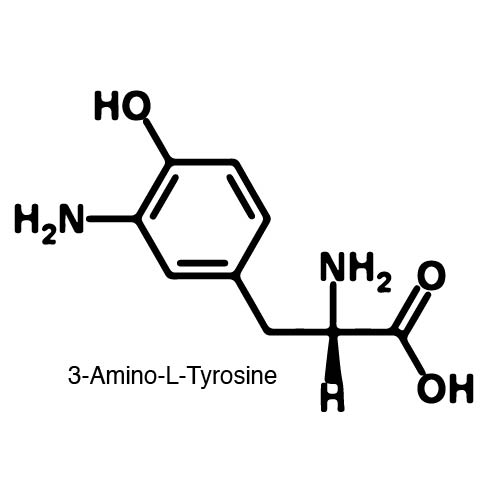
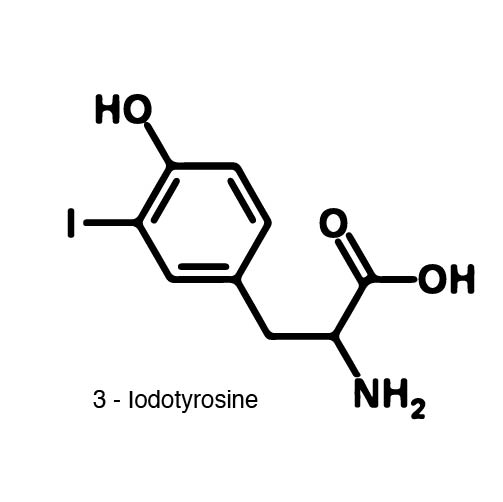
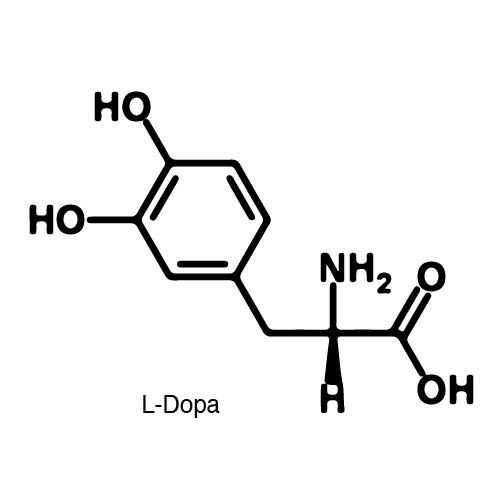
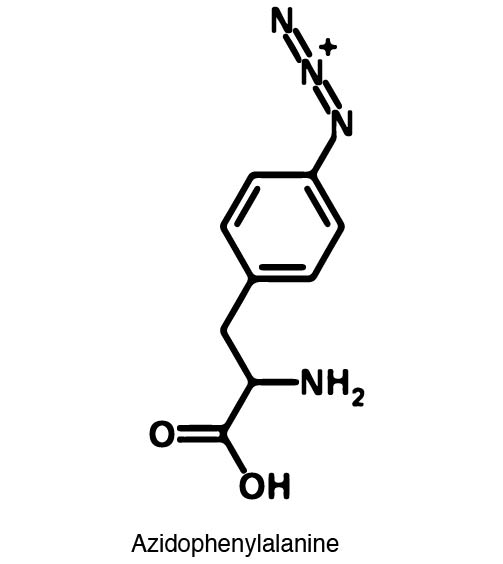
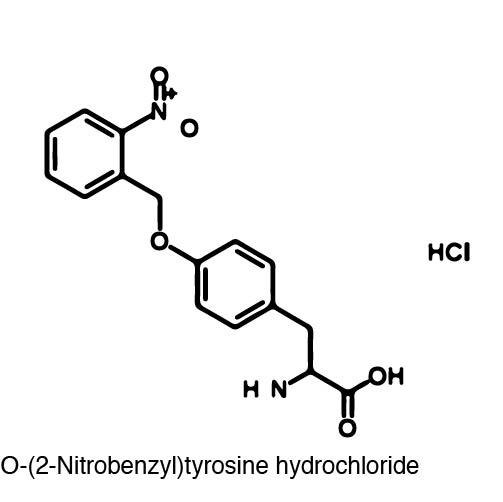
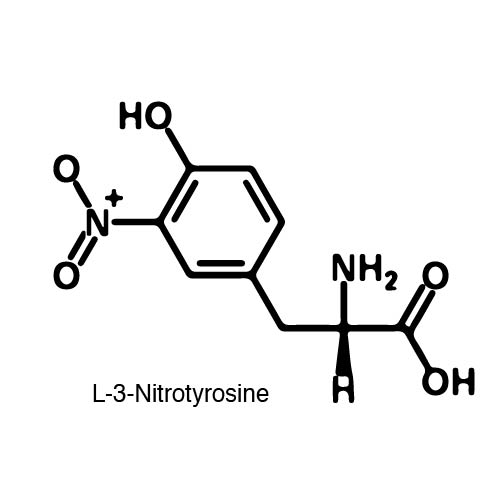
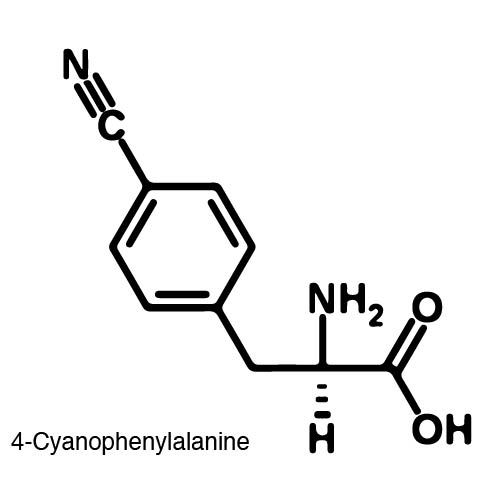
In recent years the ability to expand the genetic code has been made possible by re-coding the Amber stop codon UAG. As the library of synthetase/tRNA pairs continue to grow for non-canonical incorporation, the characterization of each remains largely vague. Measuring the efficiency of a synthetase can be time consuming and costly when considering all that is necessary for mass spec. The University of Texas iGEM Team has developed a method that allows for in vivo, qualitative, quantitative, and affordable efficiency characterization of synthetase/tRNA pairs. | In recent years the ability to expand the genetic code has been made possible by re-coding the Amber stop codon UAG. As the library of synthetase/tRNA pairs continue to grow for non-canonical incorporation, the characterization of each remains largely vague. Measuring the efficiency of a synthetase can be time consuming and costly when considering all that is necessary for mass spec. The University of Texas iGEM Team has developed a method that allows for in vivo, qualitative, quantitative, and affordable efficiency characterization of synthetase/tRNA pairs. | ||
| - | + | ==Background== | |
The genetic code is a composition of 20 highly conserved amino acids that are essential to all organisms on Earth. While specific, the genetic code is degenerate which conveniently adds flexibility to the code. By re-coding one of the redundancies, a codon can signal for the incorporation of an ncAA rather than the original canonical. The Amber codon is the least abundant stop codon of amber, ochre, and opal and thus perfect for re-coding purposes due to minimization of resulting disruptions in the cell. | The genetic code is a composition of 20 highly conserved amino acids that are essential to all organisms on Earth. While specific, the genetic code is degenerate which conveniently adds flexibility to the code. By re-coding one of the redundancies, a codon can signal for the incorporation of an ncAA rather than the original canonical. The Amber codon is the least abundant stop codon of amber, ochre, and opal and thus perfect for re-coding purposes due to minimization of resulting disruptions in the cell. | ||
| Line 84: | Line 84: | ||
Complications arise when the genetic code is re-coded. Release Factor RF1 is responsible for terminating translation when the ribosome reaches the Amber stop codon. To avoid termination at UAG, a strain of E. Coli was engineered to remove the RF1 gene. During translation, the ncAA synthetase charges an ncAA onto the tRNA. Rather than translation termination, the tRNA enters the ribosome for ncAA incorporation and translation continues. Another complication occurs in the RFO strain of E. Coli due to the presence of Amber codons on the F plasmid. The F plasmid contains essential genes to the cell. The functionality of the proteins encoded would be compromised in a genetically expanded cell. To prevent this lethality, a strain of E. Coli with the RF1 knockout was engineered to have alternative stop codons on the F plasmid known as Amberless E. Coli. | Complications arise when the genetic code is re-coded. Release Factor RF1 is responsible for terminating translation when the ribosome reaches the Amber stop codon. To avoid termination at UAG, a strain of E. Coli was engineered to remove the RF1 gene. During translation, the ncAA synthetase charges an ncAA onto the tRNA. Rather than translation termination, the tRNA enters the ribosome for ncAA incorporation and translation continues. Another complication occurs in the RFO strain of E. Coli due to the presence of Amber codons on the F plasmid. The F plasmid contains essential genes to the cell. The functionality of the proteins encoded would be compromised in a genetically expanded cell. To prevent this lethality, a strain of E. Coli with the RF1 knockout was engineered to have alternative stop codons on the F plasmid known as Amberless E. Coli. | ||
| - | + | ==Experimental Design and Method== | |
In order to re-code UAG, a synthetase/tRNA pair much be modified to effectively grab onto an ncAA. Various methods of directed evolution are typically used to modify a synthase such that it can grab onto and charge a non-canonical. The library of ncAA synthetases available have a ranging levels of reported efficiency and are not well characterized. The This year the UT iGEM Team created a test kit designed to characterize the efficiency of an ncAA synthetase/tRNA system. | In order to re-code UAG, a synthetase/tRNA pair much be modified to effectively grab onto an ncAA. Various methods of directed evolution are typically used to modify a synthase such that it can grab onto and charge a non-canonical. The library of ncAA synthetases available have a ranging levels of reported efficiency and are not well characterized. The This year the UT iGEM Team created a test kit designed to characterize the efficiency of an ncAA synthetase/tRNA system. | ||
Revision as of 20:52, 11 October 2014
|
 "
"